Empirical Benchmarking of LLMs on Molecular Tasks
This is an Empirical paper that systematically evaluates whether large language models (LLMs) can handle molecular property prediction tasks. The primary contribution is a structured benchmarking framework that compares LLMs (GPT-3.5, GPT-4, Llama-2-7b, Llama-2-13b) against conventional ML models (DeBERTa, GCN, GIN) across six standard molecular benchmark datasets from OGB. The study also introduces a collaborative framework where LLM-generated responses augment ML model features.
Why Benchmark LLMs on Molecular Property Prediction
LLMs have demonstrated strong capabilities across many NLP tasks, but their effectiveness on structured scientific data, particularly molecular graphs, remains unclear. Prior work has explored LLMs for chemistry tasks such as reaction prediction, name-to-SMILES translation, and molecule description. However, a systematic evaluation of LLMs on standard molecular property prediction benchmarks (classification and regression) with controlled prompt engineering has been lacking.
The key questions motivating this work:
- Can LLMs effectively predict molecular properties when given SMILES strings and textual descriptions of molecular structure?
- Does encoding geometric structure information as text help LLMs understand molecules?
- Can LLM responses serve as useful augmentations for traditional ML models?
Prompt Engineering for Molecular Prediction
The core methodological contribution is a systematic prompt engineering framework for querying LLMs on molecule tasks. Given a molecule $\mathcal{G} = (S, G, D)$ where $S$ is the SMILES string, $G$ is the geometric structure, and $D$ is a generated text description of atom features and graph structure, the authors design several prompt templates:
Zero-shot prompts (three variants):
- Input-Feature (IF): Asks for general insights about a molecule given its SMILES and description
- Input-Prediction (IP): Asks for a direct prediction in a specified format
- Input-Explanation (IE): Asks for both a prediction and an explanation
Each zero-shot prompt has a variant with descriptions (IFD, IPD, IED) that encodes atom features and graph structure as additional text following the approach of Fatemi et al. (2023).
Few-shot prompts (FS-k): Provide $k$ labeled examples as in-context learning demonstrations before the query. The study uses $k \in {1, 2, 3}$.
The authors also explore three predictive model pipelines:
- Solo: A single model (LLM, LM, or GNN) makes predictions independently
- Duo: An ML model receives both the original features and LLM-generated responses as input
- Trio: A GNN receives SMILES embeddings from an LM plus LLM response embeddings alongside geometric features
The LLM prediction can be formalized as $A = f_{LLM}(Q)$ where $Q$ is the prompt and $A$ is the response. For the ML augmentation pipelines, the LM-based Duo model predicts as:
$$\hat{y} = f_{LM}(S, R)$$
where $R$ is the LLM response, and the GNN-based Trio model predicts as:
$$\hat{y} = f_{GNN}(G, X)$$
where $X$ includes features derived from both SMILES embeddings and LLM response embeddings.
Experimental Setup Across Six OGB Benchmarks
Datasets
The study uses six molecular property prediction datasets from OGB and MoleculeNet:
| Dataset | Molecules | Avg. Nodes | Avg. Edges | Task Type |
|---|---|---|---|---|
| ogbg-molbace | 1,513 | 34.1 | 73.7 | Binary classification (BACE-1 inhibition) |
| ogbg-molbbbp | 2,039 | 24.1 | 51.9 | Binary classification (BBB penetration) |
| ogbg-molhiv | 41,127 | 25.5 | 27.5 | Binary classification (HIV inhibition) |
| ogbg-molesol | 1,128 | 13.3 | 27.4 | Regression (water solubility) |
| ogbg-molfreesolv | 642 | 8.7 | 16.8 | Regression (hydration free energy) |
| ogbg-mollipo | 4,200 | 27.0 | 59.0 | Regression (lipophilicity) |
Classification tasks are evaluated by ROC-AUC (higher is better) and regression tasks by RMSE (lower is better).
Models Compared
- LLMs: GPT-3.5 (primary), GPT-4, Llama-2-7b, Llama-2-13b, all used as black-box APIs with fixed parameters
- Language Model: DeBERTa, fine-tuned on SMILES strings
- GNNs: GCN and GIN, trained on geometric molecular structure
Key Results: LLMs Alone vs. ML Models
The paper presents five main observations:
Observation 1: GPT models outperform Llama models on molecule tasks. On the ogbg-molhiv dataset, GPT-3.5 and GPT-4 consistently outperform Llama-2-7b and Llama-2-13b across all prompt variants. GPT-4 offers marginal improvement over GPT-3.5 at 20x the cost and 10x the latency, so GPT-3.5 is used as the default LLM.
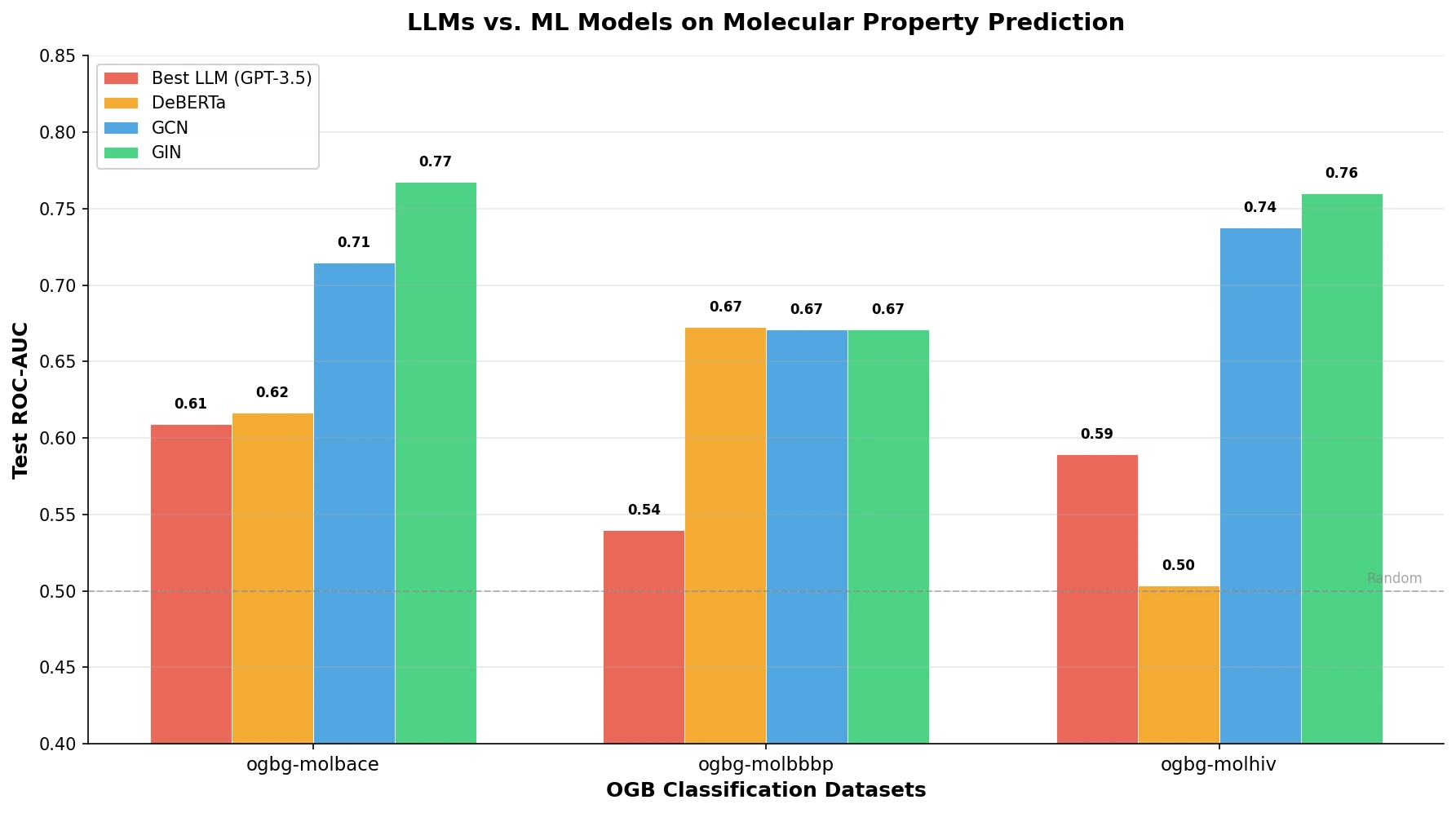
Observation 2: LLMs lag behind ML models across all datasets. Across all six datasets, LLM-based approaches underperform compared to DeBERTa, GCN, and GIN. For example, on ogbg-molhiv, the best LLM achieves 0.5892 ROC-AUC (IP prompt) compared to GIN’s 0.7601. On regression tasks, the gap is even larger: GIN achieves 0.9555 RMSE on ogbg-molesol versus the best LLM’s 1.9963.
Observation 3: Text descriptions of molecular geometry do not help LLMs. Adding structural descriptions (the “D” variants of prompts) generally degrades LLM performance and reduces response consistency. The additional tokens from structure descriptions appear to introduce noise rather than useful geometric information.
Observation 4: Geometric structure is critical for molecular prediction. GNN models that directly process molecular graphs substantially outperform both LLMs and text-based language models, confirming that geometric information is essential for accurate property prediction.
Observation 5: LLMs can augment ML models effectively. When LLM responses are used as additional features for GNN models (Duo and Trio pipelines), several configurations show improvements. For example, on ogbg-molbace, GCN with FS-2 augmentation achieves 0.7903 test ROC-AUC versus baseline GCN’s 0.7147. GIN with SMILES features (Duo pipeline) achieves 0.7837 on ogbg-molhiv versus the baseline GIN’s 0.7601.
Response Consistency
The study also measures response consistency, defined as the fraction of LLM responses conforming to the required output format. Adding descriptions to prompts reduces consistency, and few-shot prompts generally improve consistency over zero-shot variants.
Findings, Limitations, and Future Directions
Key Findings
- LLMs are not competitive with specialized ML models for molecular property prediction when used directly, with GNNs maintaining clear advantages across all six benchmark datasets.
- Converting molecular geometric structure to text descriptions is insufficient for conveying structural information to LLMs, as evidenced by degraded performance and reduced response consistency with description-augmented prompts.
- LLMs show the most promise as augmenters of existing ML models rather than as standalone predictors, with the Duo and Trio pipelines yielding improvements over Solo baselines in many configurations.
- Among LLMs, GPT-3.5 offers the best cost-performance tradeoff for molecule tasks.
Limitations
- The study is limited to black-box API access with fixed LLM parameters. Fine-tuning or parameter-efficient adaptation (e.g., LoRA) was not explored due to computational constraints and API limitations.
- Advanced prompting techniques (Chain-of-Thought, Tree-of-Thought, Graph-of-Thought, RAG) were tested in preliminary experiments but performed worse, which the authors attribute to the difficulty of designing proper reasoning chains for molecular property prediction.
- Only six datasets from OGB/MoleculeNet are evaluated. Other molecular tasks (e.g., reaction prediction, retrosynthesis) are not covered.
- The evaluation uses a single random seed for LLM queries, and the stochastic nature of LLM outputs means results may vary across runs.
Future Directions
The authors identify three promising avenues: (1) developing methods to better incorporate molecular geometric structure into LLM inputs, (2) designing more sophisticated frameworks for integrating LLMs with traditional ML models, and (3) training domain-specialized chemistry LLMs that can reduce hallucinations in chemical reasoning.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Evaluation | ogbg-molbace | 1,513 molecules | Binary classification, BACE-1 inhibition |
| Evaluation | ogbg-molbbbp | 2,039 molecules | Binary classification, BBB penetration |
| Evaluation | ogbg-molhiv | 41,127 molecules | Binary classification, HIV inhibition |
| Evaluation | ogbg-molesol | 1,128 molecules | Regression, water solubility |
| Evaluation | ogbg-molfreesolv | 642 molecules | Regression, hydration free energy |
| Evaluation | ogbg-mollipo | 4,200 molecules | Regression, lipophilicity |
All datasets use standard OGB scaffold splits.
Algorithms
- Zero-shot prompts: IF, IP, IE (and description-augmented variants IFD, IPD, IED)
- Few-shot prompts: FS-1, FS-2, FS-3
- Solo/Duo/Trio integration pipelines for combining LLM outputs with ML models
- DeBERTa fine-tuned on SMILES strings
- GCN and GIN with OGB benchmark implementations
Models
- GPT-3.5 and GPT-4 via OpenAI API with default hyperparameters
- Llama-2-7b and Llama-2-13b via HuggingFace
- DeBERTa (DeBERTaV3)
- GCN and GIN following OGB leaderboard implementations
Evaluation
| Metric | Task | Notes |
|---|---|---|
| ROC-AUC | Classification (molbace, molbbbp, molhiv) | Higher is better |
| RMSE | Regression (molesol, molfreesolv, mollipo) | Lower is better |
| Response consistency | All tasks | Fraction of format-conforming LLM outputs |
Hardware
Hardware details are not specified in the paper. LLM experiments use API calls (OpenAI) and HuggingFace inference. GNN and DeBERTa training uses standard implementations from OGB benchmark leaderboards.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| LLMaMol | Code | Not specified | Official implementation with prompt templates and evaluation pipeline |
Paper Information
Citation: Zhong, Z., Zhou, K., & Mottin, D. (2024). Benchmarking Large Language Models for Molecule Prediction Tasks. arXiv preprint arXiv:2403.05075.
@article{zhong2024benchmarking,
title={Benchmarking Large Language Models for Molecule Prediction Tasks},
author={Zhong, Zhiqiang and Zhou, Kuangyu and Mottin, Davide},
journal={arXiv preprint arXiv:2403.05075},
year={2024},
doi={10.48550/arxiv.2403.05075}
}