An LLM-Powered Agent for Autonomous Chemical Experimentation
This is a Method paper that introduces Coscientist, an AI system driven by GPT-4 that autonomously designs, plans, and performs complex chemical experiments. The primary contribution is a modular multi-LLM agent architecture that integrates internet search, documentation retrieval, code execution, and robotic experimentation APIs into a unified system capable of end-to-end experimental chemistry with minimal human intervention.
Bridging LLM Capabilities and Laboratory Automation
Transformer-based large language models had demonstrated strong capabilities in natural language processing, biology, chemistry, and code generation by early 2023. Simultaneously, laboratory automation had progressed with autonomous reaction discovery, automated flow systems, and mobile robotic platforms. However, these two threads remained largely separate: LLMs could reason about chemistry in text, but could not act on that reasoning by controlling physical experiments.
The gap this work addresses is the integration of LLM reasoning with laboratory automation in a closed-loop system. Prior automated chemistry systems relied on traditional optimization algorithms or narrow AI components. The question was whether GPT-4’s general reasoning capabilities could be combined with tool access to produce a system that autonomously designs experiments, writes instrument code, executes reactions, and interprets results, all from natural language prompts.
This work was developed independently and in parallel with other autonomous agent efforts (AutoGPT, BabyAGI, LangChain), with ChemCrow serving as another chemistry-specific example.
A Modular Multi-LLM Architecture with Tool Access
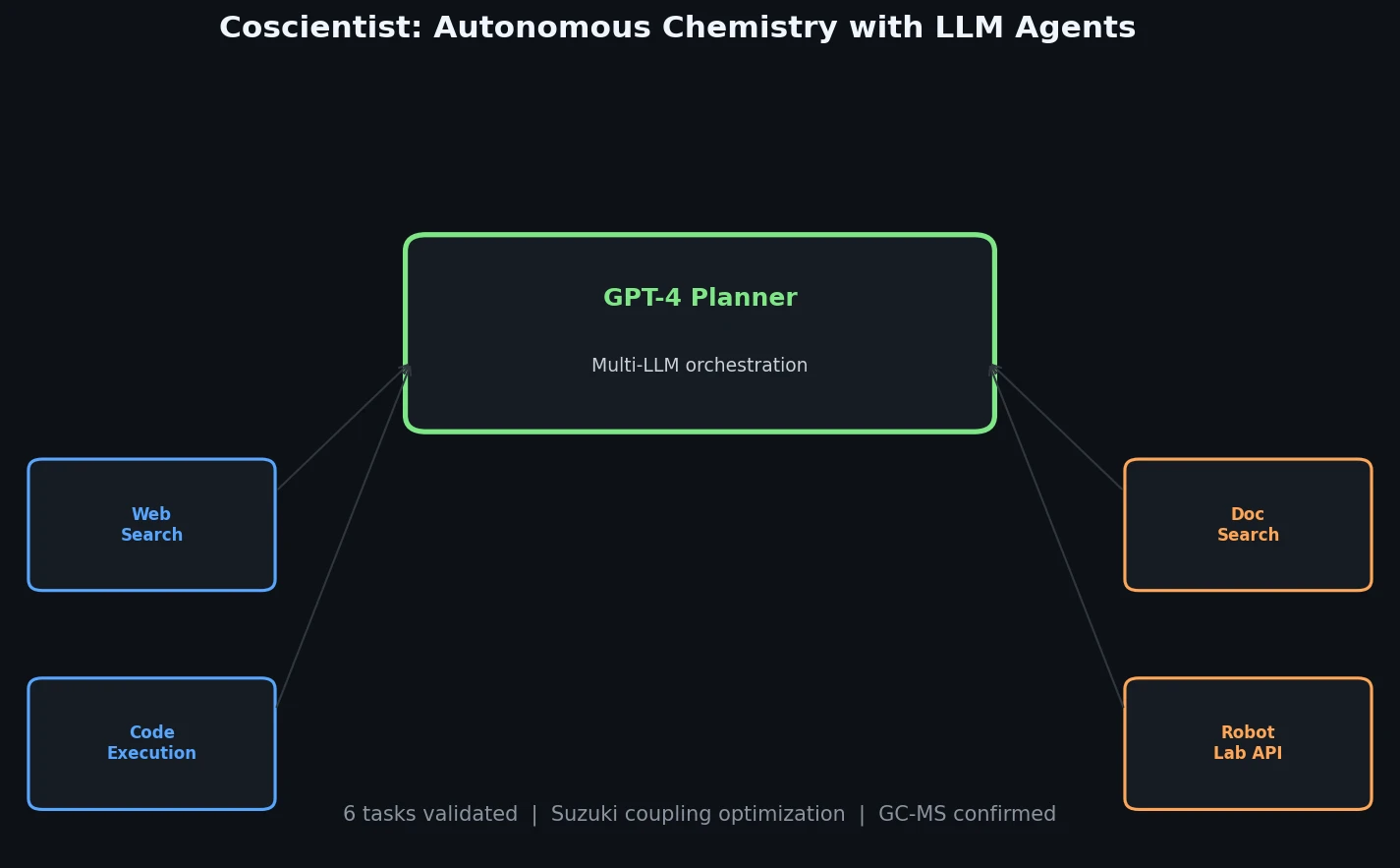
The core innovation is Coscientist’s modular architecture, centered on a “Planner” module (a GPT-4 chat completion instance) that orchestrates four command types:
- GOOGLE: A Web Searcher module (itself an LLM) that transforms prompts into search queries, browses results, and funnels answers back to the Planner.
- PYTHON: A Code Execution module running in an isolated Docker container for calculations and data analysis, with no LLM dependency.
- DOCUMENTATION: A Docs Searcher module that retrieves and summarizes technical documentation (e.g., Opentrons Python API, Emerald Cloud Lab Symbolic Lab Language) using ada embeddings and distance-based vector search.
- EXPERIMENT: An Automation module that executes generated code on laboratory hardware or provides synthetic procedures.
The system prompts are engineered in a modular fashion, with the Planner receiving initial user input and command outputs as messages. The Planner can iteratively call commands, fix software errors, and refine its approach. This design allows natural language instructions (e.g., “perform multiple Suzuki reactions”) to be translated into complete experimental protocols.
For documentation retrieval, all sections of the OT-2 API documentation were embedded using OpenAI’s ada model, and relevant sections are retrieved via cosine similarity search. For the Emerald Cloud Lab, the system learned to program in a symbolic lab language (SLL) that was completely unknown to GPT-4 at training time, demonstrating effective in-context learning from supplied documentation.
Six Tasks Demonstrating Autonomous Chemistry Capabilities
The paper evaluates Coscientist across six tasks of increasing complexity.
Task 1: Chemical Synthesis Planning
A benchmark of seven compounds was used to compare synthesis planning across models (GPT-4, GPT-3.5, Claude 1.3, Falcon-40B-Instruct) with and without web search. Outputs were scored on a 1-5 scale:
| Score | Meaning |
|---|---|
| 5 | Very detailed and chemically accurate procedure |
| 4 | Detailed and accurate but without reagent quantities |
| 3 | Correct chemistry but no step-by-step procedure |
| 2 | Extremely vague or unfeasible |
| 1 | Incorrect or failure to follow instructions |
The GPT-4-powered Web Searcher achieved maximum scores for acetaminophen, aspirin, nitroaniline, and phenolphthalein. It was the only approach to achieve acceptable scores (3+) for ibuprofen, which all non-browsing models synthesized incorrectly. These results highlight the importance of grounding LLMs to avoid hallucinations.
Task 2: Documentation Search
The system correctly identified relevant ECL functions from documentation and generated valid SLL code that was successfully executed at ECL, including an HPLC experiment on a caffeine standard sample.
Task 3: Cloud Laboratory Execution
Using prompt-to-function and prompt-to-SLL pipelines, Coscientist generated executable code for the Emerald Cloud Lab. It also searched a catalogue of 1,110 model samples to identify relevant stock solutions from simple search terms.
Task 4: Liquid Handler Control
Using the Opentrons OT-2, Coscientist translated natural language prompts (e.g., “colour every other line with one colour of your choice,” “draw a red cross”) into accurate liquid handling protocols.
Task 5: Integrated Multi-Module Experiment
The most complex demonstration combined web search, code execution, documentation retrieval, and hardware control to design and execute Suzuki-Miyaura and Sonogashira cross-coupling reactions. Coscientist:
- Searched the internet for reaction conditions and stoichiometries
- Selected correct coupling partners (never misassigning phenylboronic acid to Sonogashira)
- Calculated reagent volumes and wrote OT-2 protocols
- Self-corrected when using an incorrect heater-shaker method by consulting documentation
- Successfully produced target products confirmed by GC-MS analysis (biphenyl at 9.53 min for Suzuki, diphenylacetylene at 12.92 min for Sonogashira)
Task 6: Reaction Optimization
Coscientist was tested on two fully mapped reaction datasets:
- Suzuki reaction flow dataset (Perera et al.): varying ligands, reagents/bases, and solvents
- Buchwald-Hartwig C-N coupling dataset (Doyle et al.): varying ligands, additives, and bases
Performance was evaluated using a normalized advantage metric:
$$\text{Normalized Advantage} = \frac{\text{yield}_i - \overline{\text{yield}}}{\text{yield}_{\max} - \overline{\text{yield}}}$$
A value of 1 indicates maximum yield reached, 0 indicates random performance, and negative values indicate worse than random. The normalized maximum advantage (NMA) tracks the best result achieved up to each iteration.
Key findings from the optimization experiments:
- GPT-4 with prior information (10 random data points) produced better initial guesses than GPT-4 without prior information
- Both GPT-4 approaches converged to similar NMA values at the limit
- Both GPT-4 approaches outperformed standard Bayesian optimization in NMA and normalized advantage
- GPT-3.5 largely failed due to inability to output correct JSON schemas
- On the Buchwald-Hartwig dataset, GPT-4 performed comparably whether given compound names or SMILES strings, and could reason about electronic properties from SMILES representations
All experiments used a maximum of 20 iterations (5.2% and 6.9% of the total reaction space for the two datasets).
Demonstrated Versatility with Safety Considerations
Coscientist demonstrated that GPT-4, when equipped with appropriate tool access, can autonomously handle the full experimental chemistry workflow from literature search to reaction execution and data interpretation. The system showed chemical reasoning capabilities, including selecting appropriate reagents, providing justifications for choices based on reactivity and selectivity, and using experimental data to guide subsequent iterations.
Several limitations are acknowledged:
- The experimental setup was not yet fully automated (plates were moved manually between instruments), though no human decision-making was involved
- GPT-3.5 consistently underperformed due to inability to follow formatting instructions
- The synthesis planning evaluation scale is inherently subjective
- It is unclear whether GPT-4’s training data contained information from the optimization datasets
- The comparison with Bayesian optimization may reflect different exploration/exploitation balances rather than pure capability differences
The authors raise safety concerns about dual-use potential and note that full code and prompts were withheld pending development of US AI regulations. A simplified implementation was released for reproducibility purposes.
Future directions include extending the system with reaction databases (Reaxys, SciFinder), implementing advanced prompting strategies (ReAct, Chain of Thought, Tree of Thoughts), and developing automated quality control for cloud laboratory experiments.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Synthesis benchmark | 7 compound set | 7 compounds | Acetaminophen, aspirin, ibuprofen, nitroaniline, etc. |
| Optimization | Perera et al. Suzuki flow dataset | Fully mapped condition space | Varying ligands, bases, solvents |
| Optimization | Doyle Buchwald-Hartwig dataset | Fully mapped condition space | Varying ligands, additives, bases |
| Reagent selection | SMILES compound database | Not specified | Used for computational experiments |
Algorithms
- Planner: GPT-4 chat completion with modular system prompts
- Web Searcher: GPT-4 or GPT-3.5-turbo for query generation and result parsing
- Documentation embedding: OpenAI ada model with distance-based vector search
- Code execution: Isolated Docker container (no LLM dependency)
- Baseline: Bayesian optimization with varying initial sample sizes (1-10)
Models
- GPT-4 (primary)
- GPT-3.5-turbo (baseline)
- Claude 1.3 (baseline for synthesis planning)
- Falcon-40B-Instruct (baseline for synthesis planning)
- OpenAI ada (for documentation embedding)
Evaluation
| Metric | Context | Notes |
|---|---|---|
| Synthesis score (1-5) | 7-compound benchmark | Subjective expert grading |
| Normalized advantage | Optimization tasks | Measures improvement over random |
| NMA | Optimization tasks | Maximum advantage achieved through iteration N |
| GC-MS confirmation | Cross-coupling reactions | Product formation verified experimentally |
Hardware
- Opentrons OT-2 liquid handler with heater-shaker module
- UV-Vis plate reader
- Emerald Cloud Lab (cloud-based automation)
- Computational requirements not specified (relies on OpenAI API calls)
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| gomesgroup/coscientist | Code | Apache-2.0 with Commons Clause | Simplified implementation; full code withheld for safety |
Paper Information
Citation: Boiko, D. A., MacKnight, R., Kline, B. & Gomes, G. (2023). Autonomous chemical research with large language models. Nature, 624(7992), 570-578. https://doi.org/10.1038/s41586-023-06792-0
@article{boiko2023autonomous,
title={Autonomous chemical research with large language models},
author={Boiko, Daniil A. and MacKnight, Robert and Kline, Ben and Gomes, Gabriel dos Passos},
journal={Nature},
volume={624},
number={7992},
pages={570--578},
year={2023},
publisher={Springer Nature},
doi={10.1038/s41586-023-06792-0}
}