Key Contribution
MARCEL provides a benchmark for conformer ensemble learning. It demonstrates that explicitly modeling full conformer distributions improves property prediction across drug-like molecules and organometallic catalysts.
Overview
The Molecular Representation and Conformer Ensemble Learning (MARCEL) dataset provides 722K+ conformations across 76K+ molecules spanning four diverse chemical domains: drug-like molecules (Drugs-75K), organophosphorus ligands (Kraken), chiral catalysts (EE), and organometallic complexes (BDE). MARCEL evaluates conformer ensemble methods across both pharmaceutical and catalysis applications.
Dataset Examples
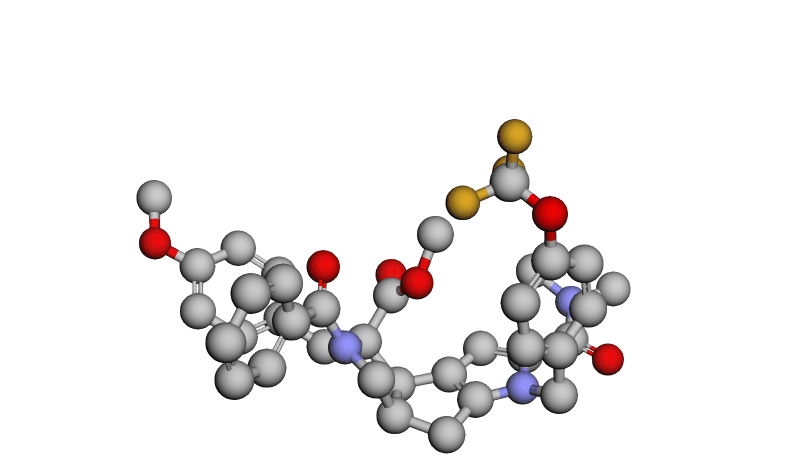
COC(=O)[C@@]1(Cc2ccc(OC)cc2)[C@H]2c3cc(C(=O)N(C)C)n(Cc4ccc(OC(F)(F)F)cc4)c3C[C@H]2CN1C(=O)c1ccccc1; IUPAC: methyl (2R,3R,6R)-4-benzoyl-10-(dimethylcarbamoyl)-3-[(4-methoxyphenyl)methyl]-9-[[4-(trifluoromethoxy)phenyl]methyl]-4,9-diazatricyclo[6.3.0.02,6]undeca-1(8),10-diene-3-carboxylate)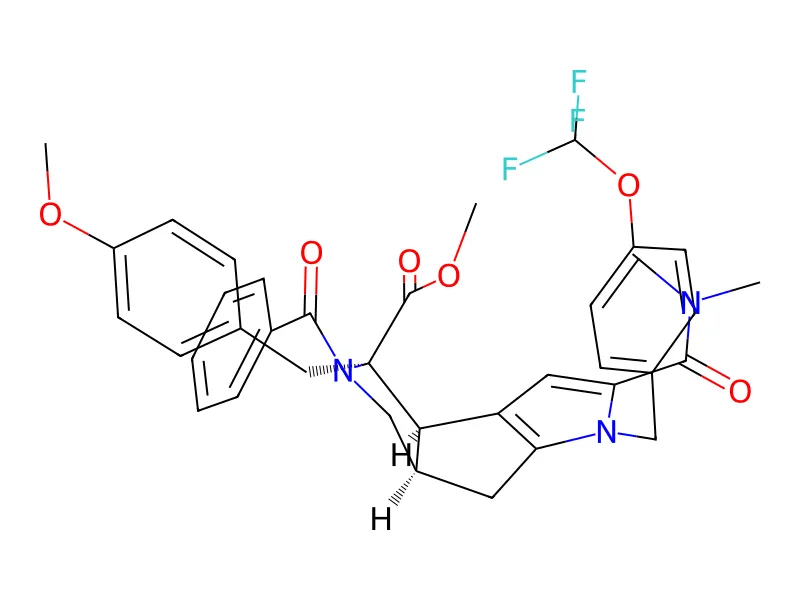
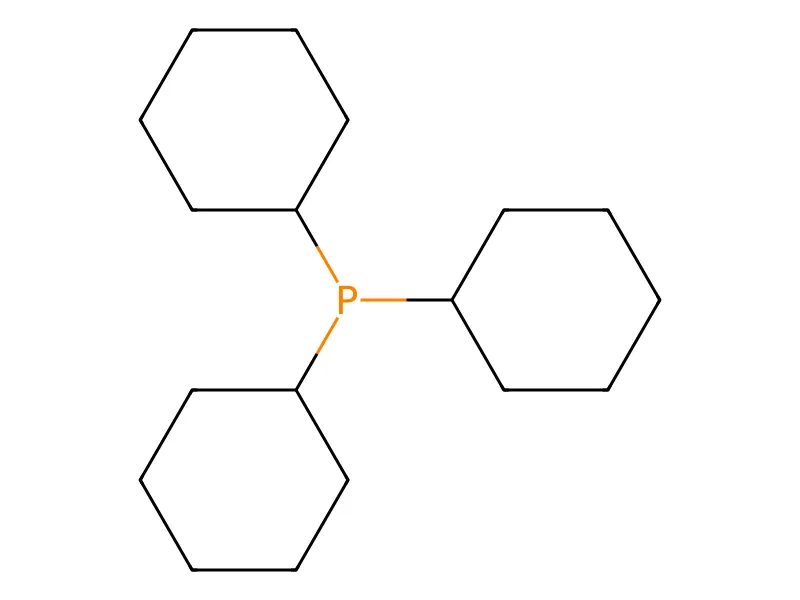
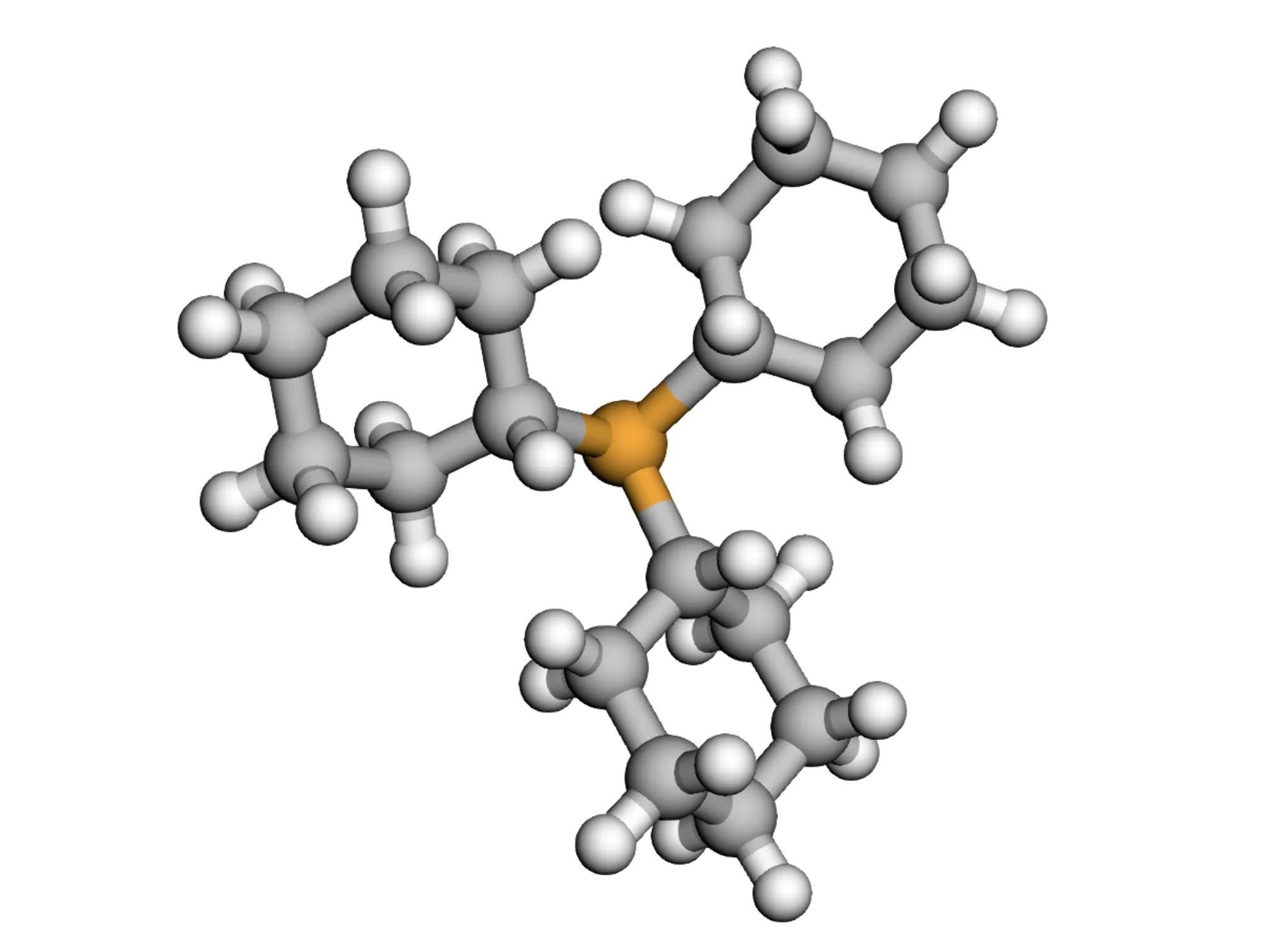
Dataset Subsets
| Subset | Count | Description |
|---|---|---|
| Drugs-75K | 75,099 molecules | Drug-like molecules with at least 5 rotatable bonds |
| Kraken | 1,552 molecules | Monodentate organophosphorus (III) ligands |
| EE | 872 reactions | Rhodium (Rh)-bound atropisomeric catalyst-substrate pairs derived from chiral bisphosphine |
| BDE | 5,915 reactions | Organometallic catalysts ML$_1$L$_2$ with electronic binding energies |
Benchmarks
Ionization Potential (Drugs-75K)
Predict ionization potential from molecular structure
| Rank | Model | MAE (eV) |
|---|---|---|
| 🥇 1 | Ensemble - GemNet GemNet on full conformer ensemble | 0.4066 |
| 🥈 2 | 3D - GemNet Geometry-enhanced message passing (single conformer) | 0.4069 |
| 🥉 3 | Ensemble - DimeNet++ DimeNet++ on full conformer ensemble | 0.4126 |
| 4 | Ensemble - LEFTNet LEFTNet on full conformer ensemble | 0.4149 |
| 5 | 3D - LEFTNet Local Environment Feature Transformer (single conformer) | 0.4174 |
| 6 | Ensemble - ClofNet ClofNet on full conformer ensemble | 0.428 |
| 7 | 2D - GraphGPS Graph Transformer with positional encodings | 0.4351 |
| 8 | 2D - GIN Graph Isomorphism Network | 0.4354 |
| 9 | 2D - GIN+VN GIN with Virtual Nodes | 0.4361 |
| 10 | 3D - ClofNet Conformation-ensemble learning network (single conformer) | 0.4393 |
| 11 | 3D - SchNet Continuous-filter convolutional network (single conformer) | 0.4394 |
| 12 | 3D - DimeNet++ Directional message passing network (single conformer) | 0.4441 |
| 13 | Ensemble - SchNet SchNet on full conformer ensemble | 0.4452 |
| 14 | Ensemble - PaiNN PaiNN on full conformer ensemble | 0.4466 |
| 15 | 3D - PaiNN Polarizable Atom Interaction Network (single conformer) | 0.4505 |
| 16 | 2D - ChemProp Message Passing Neural Network | 0.4595 |
| 17 | 1D - LSTM LSTM on SMILES sequences | 0.4788 |
| 18 | 1D - Random forest Random Forest on Morgan fingerprints | 0.4987 |
| 19 | 1D - Transformer Transformer on SMILES sequences | 0.6617 |
Electron Affinity (Drugs-75K)
Predict electron affinity from molecular structure
| Rank | Model | MAE (eV) |
|---|---|---|
| 🥇 1 | Ensemble - GemNet GemNet on full conformer ensemble | 0.391 |
| 🥈 2 | 3D - GemNet Geometry-enhanced message passing (single conformer) | 0.3922 |
| 🥉 3 | Ensemble - DimeNet++ DimeNet++ on full conformer ensemble | 0.3944 |
| 4 | Ensemble - LEFTNet LEFTNet on full conformer ensemble | 0.3953 |
| 5 | 3D - LEFTNet Local Environment Feature Transformer (single conformer) | 0.3964 |
| 6 | Ensemble - ClofNet ClofNet on full conformer ensemble | 0.4033 |
| 7 | 2D - GraphGPS Graph Transformer with positional encodings | 0.4085 |
| 8 | 2D - GIN Graph Isomorphism Network | 0.4169 |
| 9 | 2D - GIN+VN GIN with Virtual Nodes | 0.4169 |
| 10 | 3D - SchNet Continuous-filter convolutional network (single conformer) | 0.4207 |
| 11 | 3D - DimeNet++ Directional message passing network (single conformer) | 0.4233 |
| 12 | Ensemble - SchNet SchNet on full conformer ensemble | 0.4232 |
| 13 | 3D - ClofNet Conformation-ensemble learning network (single conformer) | 0.4251 |
| 14 | Ensemble - PaiNN PaiNN on full conformer ensemble | 0.4269 |
| 15 | 2D - ChemProp Message Passing Neural Network | 0.4417 |
| 16 | 3D - PaiNN Polarizable Atom Interaction Network (single conformer) | 0.4495 |
| 17 | 1D - LSTM LSTM on SMILES sequences | 0.4648 |
| 18 | 1D - Random forest Random Forest on Morgan fingerprints | 0.4747 |
| 19 | 1D - Transformer Transformer on SMILES sequences | 0.585 |
Electronegativity (Drugs-75K)
Predict electronegativity (χ) from molecular structure
| Rank | Model | MAE (eV) |
|---|---|---|
| 🥇 1 | 3D - GemNet Geometry-enhanced message passing (single conformer) | 0.197 |
| 🥈 2 | Ensemble - GemNet GemNet on full conformer ensemble | 0.2027 |
| 🥉 3 | Ensemble - LEFTNet LEFTNet on full conformer ensemble | 0.2069 |
| 4 | 3D - LEFTNet Local Environment Feature Transformer (single conformer) | 0.2083 |
| 5 | Ensemble - ClofNet ClofNet on full conformer ensemble | 0.2199 |
| 6 | 2D - GraphGPS Graph Transformer with positional encodings | 0.2212 |
| 7 | 3D - SchNet Continuous-filter convolutional network (single conformer) | 0.2243 |
| 8 | Ensemble - SchNet SchNet on full conformer ensemble | 0.2243 |
| 9 | 2D - GIN Graph Isomorphism Network | 0.226 |
| 10 | 2D - GIN+VN GIN with Virtual Nodes | 0.2267 |
| 11 | Ensemble - DimeNet++ DimeNet++ on full conformer ensemble | 0.2267 |
| 12 | Ensemble - PaiNN PaiNN on full conformer ensemble | 0.2294 |
| 13 | 3D - PaiNN Polarizable Atom Interaction Network (single conformer) | 0.2324 |
| 14 | 3D - ClofNet Conformation-ensemble learning network (single conformer) | 0.2378 |
| 15 | 3D - DimeNet++ Directional message passing network (single conformer) | 0.2436 |
| 16 | 2D - ChemProp Message Passing Neural Network | 0.2441 |
| 17 | 1D - LSTM LSTM on SMILES sequences | 0.2505 |
| 18 | 1D - Random forest Random Forest on Morgan fingerprints | 0.2732 |
| 19 | 1D - Transformer Transformer on SMILES sequences | 0.4073 |
B₅ Sterimol Parameter (Kraken)
Predict B₅ sterimol descriptor for organophosphorus ligands
| Rank | Model | MAE |
|---|---|---|
| 🥇 1 | Ensemble - PaiNN PaiNN on full conformer ensemble | 0.2225 |
| 🥈 2 | Ensemble - GemNet GemNet on full conformer ensemble | 0.2313 |
| 🥉 3 | Ensemble - DimeNet++ DimeNet++ on full conformer ensemble | 0.263 |
| 4 | Ensemble - LEFTNet LEFTNet on full conformer ensemble | 0.2644 |
| 5 | Ensemble - SchNet SchNet on full conformer ensemble | 0.2704 |
| 6 | 3D - GemNet Geometry-enhanced message passing (single conformer) | 0.2789 |
| 7 | 3D - LEFTNet Local Environment Feature Transformer (single conformer) | 0.3072 |
| 8 | 2D - GIN Graph Isomorphism Network | 0.3128 |
| 9 | Ensemble - ClofNet ClofNet on full conformer ensemble | 0.3228 |
| 10 | 3D - SchNet Continuous-filter convolutional network (single conformer) | 0.3293 |
| 11 | 3D - PaiNN Polarizable Atom Interaction Network (single conformer) | 0.3443 |
| 12 | 2D - GraphGPS Graph Transformer with positional encodings | 0.345 |
| 13 | 3D - DimeNet++ Directional message passing network (single conformer) | 0.351 |
| 14 | 2D - GIN+VN GIN with Virtual Nodes | 0.3567 |
| 15 | 1D - Random forest Random Forest on Morgan fingerprints | 0.476 |
| 16 | 2D - ChemProp Message Passing Neural Network | 0.485 |
| 17 | 3D - ClofNet Conformation-ensemble learning network (single conformer) | 0.4873 |
| 18 | 1D - LSTM LSTM on SMILES sequences | 0.4879 |
| 19 | 1D - Transformer Transformer on SMILES sequences | 0.9611 |
L Sterimol Parameter (Kraken)
Predict L sterimol descriptor for organophosphorus ligands
| Rank | Model | MAE |
|---|---|---|
| 🥇 1 | Ensemble - GemNet GemNet on full conformer ensemble | 0.3386 |
| 🥈 2 | Ensemble - DimeNet++ DimeNet++ on full conformer ensemble | 0.3468 |
| 🥉 3 | Ensemble - PaiNN PaiNN on full conformer ensemble | 0.3619 |
| 4 | Ensemble - LEFTNet LEFTNet on full conformer ensemble | 0.3643 |
| 5 | 3D - GemNet Geometry-enhanced message passing (single conformer) | 0.3754 |
| 6 | 2D - GIN Graph Isomorphism Network | 0.4003 |
| 7 | 3D - DimeNet++ Directional message passing network (single conformer) | 0.4174 |
| 8 | 1D - Random forest Random Forest on Morgan fingerprints | 0.4303 |
| 9 | Ensemble - SchNet SchNet on full conformer ensemble | 0.4322 |
| 10 | 2D - GIN+VN GIN with Virtual Nodes | 0.4344 |
| 11 | 2D - GraphGPS Graph Transformer with positional encodings | 0.4363 |
| 12 | 3D - PaiNN Polarizable Atom Interaction Network (single conformer) | 0.4471 |
| 13 | Ensemble - ClofNet ClofNet on full conformer ensemble | 0.4485 |
| 14 | 3D - LEFTNet Local Environment Feature Transformer (single conformer) | 0.4493 |
| 15 | 1D - LSTM LSTM on SMILES sequences | 0.5142 |
| 16 | 2D - ChemProp Message Passing Neural Network | 0.5452 |
| 17 | 3D - SchNet Continuous-filter convolutional network (single conformer) | 0.5458 |
| 18 | 3D - ClofNet Conformation-ensemble learning network (single conformer) | 0.6417 |
| 19 | 1D - Transformer Transformer on SMILES sequences | 0.8389 |
Buried B₅ Parameter (Kraken)
Predict buried B₅ sterimol descriptor for organophosphorus ligands
| Rank | Model | MAE |
|---|---|---|
| 🥇 1 | Ensemble - GemNet GemNet on full conformer ensemble | 0.1589 |
| 🥈 2 | Ensemble - PaiNN PaiNN on full conformer ensemble | 0.1693 |
| 🥉 3 | 2D - GIN Graph Isomorphism Network | 0.1719 |
| 4 | 3D - GemNet Geometry-enhanced message passing (single conformer) | 0.1782 |
| 5 | Ensemble - DimeNet++ DimeNet++ on full conformer ensemble | 0.1783 |
| 6 | Ensemble - SchNet SchNet on full conformer ensemble | 0.2024 |
| 7 | Ensemble - LEFTNet LEFTNet on full conformer ensemble | 0.2017 |
| 8 | 2D - GraphGPS Graph Transformer with positional encodings | 0.2066 |
| 9 | 3D - DimeNet++ Directional message passing network (single conformer) | 0.2097 |
| 10 | Ensemble - ClofNet ClofNet on full conformer ensemble | 0.2178 |
| 11 | 3D - LEFTNet Local Environment Feature Transformer (single conformer) | 0.2176 |
| 12 | 3D - SchNet Continuous-filter convolutional network (single conformer) | 0.2295 |
| 13 | 3D - PaiNN Polarizable Atom Interaction Network (single conformer) | 0.2395 |
| 14 | 2D - GIN+VN GIN with Virtual Nodes | 0.2422 |
| 15 | 1D - Random forest Random Forest on Morgan fingerprints | 0.2758 |
| 16 | 1D - LSTM LSTM on SMILES sequences | 0.2813 |
| 17 | 3D - ClofNet Conformation-ensemble learning network (single conformer) | 0.2884 |
| 18 | 2D - ChemProp Message Passing Neural Network | 0.3002 |
| 19 | 1D - Transformer Transformer on SMILES sequences | 0.4929 |
Buried L Parameter (Kraken)
Predict buried L sterimol descriptor for organophosphorus ligands
| Rank | Model | MAE |
|---|---|---|
| 🥇 1 | Ensemble - GemNet GemNet on full conformer ensemble | 0.0947 |
| 🥈 2 | Ensemble - DimeNet++ DimeNet++ on full conformer ensemble | 0.1185 |
| 🥉 3 | 2D - GIN Graph Isomorphism Network | 0.12 |
| 4 | Ensemble - PaiNN PaiNN on full conformer ensemble | 0.1324 |
| 5 | Ensemble - LEFTNet LEFTNet on full conformer ensemble | 0.1386 |
| 6 | Ensemble - SchNet SchNet on full conformer ensemble | 0.1443 |
| 7 | 3D - LEFTNet Local Environment Feature Transformer (single conformer) | 0.1486 |
| 8 | 2D - GraphGPS Graph Transformer with positional encodings | 0.15 |
| 9 | 1D - Random forest Random Forest on Morgan fingerprints | 0.1521 |
| 10 | 3D - DimeNet++ Directional message passing network (single conformer) | 0.1526 |
| 11 | Ensemble - ClofNet ClofNet on full conformer ensemble | 0.1548 |
| 12 | 3D - GemNet Geometry-enhanced message passing (single conformer) | 0.1635 |
| 13 | 3D - PaiNN Polarizable Atom Interaction Network (single conformer) | 0.1673 |
| 14 | 2D - GIN+VN GIN with Virtual Nodes | 0.1741 |
| 15 | 3D - SchNet Continuous-filter convolutional network (single conformer) | 0.1861 |
| 16 | 1D - LSTM LSTM on SMILES sequences | 0.1924 |
| 17 | 2D - ChemProp Message Passing Neural Network | 0.1948 |
| 18 | 3D - ClofNet Conformation-ensemble learning network (single conformer) | 0.2529 |
| 19 | 1D - Transformer Transformer on SMILES sequences | 0.2781 |
Enantioselectivity (EE)
Predict enantiomeric excess for Rh-catalyzed asymmetric reactions
| Rank | Model | MAE (%) |
|---|---|---|
| 🥇 1 | Ensemble - GemNet GemNet on full conformer ensemble | 11.61 |
| 🥈 2 | Ensemble - DimeNet++ DimeNet++ on full conformer ensemble | 12.03 |
| 🥉 3 | Ensemble - PaiNN PaiNN on full conformer ensemble | 13.56 |
| 4 | Ensemble - ClofNet ClofNet on full conformer ensemble | 13.96 |
| 5 | Ensemble - SchNet SchNet on full conformer ensemble | 14.22 |
| 6 | 3D - DimeNet++ Directional message passing network (single conformer) | 14.64 |
| 7 | 3D - SchNet Continuous-filter convolutional network (single conformer) | 17.74 |
| 8 | 3D - GemNet Geometry-enhanced message passing (single conformer) | 18.03 |
| 9 | Ensemble - LEFTNet LEFTNet on full conformer ensemble | 18.42 |
| 10 | 3D - LEFTNet Local Environment Feature Transformer (single conformer) | 19.8 |
| 11 | 3D - PaiNN Polarizable Atom Interaction Network (single conformer) | 20.24 |
| 12 | 3D - ClofNet Conformation-ensemble learning network (single conformer) | 33.95 |
| 13 | 2D - ChemProp Message Passing Neural Network | 61.03 |
| 14 | 1D - Random forest Random Forest on Morgan fingerprints | 61.3 |
| 15 | 2D - GraphGPS Graph Transformer with positional encodings | 61.63 |
| 16 | 1D - Transformer Transformer on SMILES sequences | 62.08 |
| 17 | 2D - GIN Graph Isomorphism Network | 62.31 |
| 18 | 2D - GIN+VN GIN with Virtual Nodes | 62.38 |
| 19 | 1D - LSTM LSTM on SMILES sequences | 64.01 |
Bond Dissociation Energy (BDE)
Predict metal-ligand bond dissociation energy for organometallic catalysts
| Rank | Model | MAE (kcal/mol) |
|---|---|---|
| 🥇 1 | 3D - DimeNet++ Directional message passing network (single conformer) | 1.45 |
| 🥈 2 | Ensemble - DimeNet++ DimeNet++ on full conformer ensemble | 1.47 |
| 🥉 3 | 3D - LEFTNet Local Environment Feature Transformer (single conformer) | 1.53 |
| 4 | Ensemble - LEFTNet LEFTNet on full conformer ensemble | 1.53 |
| 5 | Ensemble - GemNet GemNet on full conformer ensemble | 1.61 |
| 6 | 3D - GemNet Geometry-enhanced message passing (single conformer) | 1.65 |
| 7 | Ensemble - PaiNN PaiNN on full conformer ensemble | 1.87 |
| 8 | Ensemble - SchNet SchNet on full conformer ensemble | 1.97 |
| 9 | Ensemble - ClofNet ClofNet on full conformer ensemble | 2.01 |
| 10 | 3D - PaiNN Polarizable Atom Interaction Network (single conformer) | 2.13 |
| 11 | 2D - GraphGPS Graph Transformer with positional encodings | 2.48 |
| 12 | 3D - SchNet Continuous-filter convolutional network (single conformer) | 2.55 |
| 13 | 3D - ClofNet Conformation-ensemble learning network (single conformer) | 2.61 |
| 14 | 2D - GIN Graph Isomorphism Network | 2.64 |
| 15 | 2D - ChemProp Message Passing Neural Network | 2.66 |
| 16 | 2D - GIN+VN GIN with Virtual Nodes | 2.74 |
| 17 | 1D - LSTM LSTM on SMILES sequences | 2.83 |
| 18 | 1D - Random forest Random Forest on Morgan fingerprints | 3.03 |
| 19 | 1D - Transformer Transformer on SMILES sequences | 10.08 |
Related Datasets
| Dataset | Relationship | Link |
|---|---|---|
| GEOM | Source | Notes |
Strengths
- Domain diversity: Beyond drug-like molecules, includes organometallics and catalysts rarely covered in existing benchmarks
- Ensemble-based: Provides full conformer ensembles with statistical weights
- DFT-quality energies: Drugs-75K features DFT-level conformers and energies (higher accuracy than GEOM-Drugs)
- Realistic scenarios: BDE subset models the practical constraint of lacking DFT-computed conformers for large catalyst systems
- Comprehensive baselines: Benchmarks 18 models across 1D (SMILES), 2D (graph), 3D (single conformer), and ensemble methods
- Property diversity: Covers ionization potential, electron affinity, electronegativity, ligand descriptors, and catalytic properties
Limitations
- Regression only: All tasks evaluate regression metrics exclusively
- Chemical space coverage: The 76K molecules encapsulate a fraction of the expansive drug-like and catalyst chemical spaces
- Compute requirements: Working with large conformer ensembles demands significant computational resources
- Proprietary data: EE subset is proprietary (as of December 2025)
- DFT bottleneck: BDE demonstrates a practical limitation: single DFT optimization can take 2-3 days, making conformer-level DFT infeasible for large organometallics
- Uniform sampling baseline: The initial data augmentation strategy tested for handling ensembles samples conformers uniformly rather than by Boltzmann weight. This unprincipled physical assumption likely explains why the strategy occasionally introduces noise and fails to aid complex 3D architectures.
- Drugs-75K properties: The large-scale benchmark (Drugs-75K) specifically targets electronic properties (Ionization Potential, Electron Affinity, Electronegativity). As the authors explicitly highlight in Section 5.2, these properties are generally less sensitive to conformational rotations compared to steric or spatial interactions. This significantly confounds evaluating whether explicit conformer ensembles actually benefit large-scale regression tasks.
- Unrealistic single-conformer baselines: The 3D single-conformer models are exclusively evaluated on the lowest-energy conformer. This setup is inherently flawed for real-world application, as knowing the global minimum a priori requires exhaustively searching and computing energies for the entire conformer space.
Technical Notes
Data Generation Pipeline
Drugs-75K
Source: GEOM-Drugs subset
Filtering:
- Minimum 5 rotatable bonds (focus on flexible molecules)
- Allowed elements: H, C, N, O, F, Si, P, S, Cl
Conformer generation:
- DFT-level calculations for both conformers and energies
- Higher accuracy than original GEOM-Drugs (semi-empirical GFN2-xTB)
Properties: Ionization Potential (IP), Electron Affinity (EA), Electronegativity (χ)
Kraken
Source: Original Kraken dataset (1,552 monodentate organophosphorus(III) ligands)
Properties: 4 of 78 available properties (selected for high variance across conformer ensembles)
- $B_5$: Sterimol B5, maximum width of substituent (steric descriptor)
- $L$: Sterimol L, length of substituent (steric descriptor)
- $\text{Bur}B_5$: Buried Sterimol B5, steric effects within the first coordination sphere
- $\text{Bur}L$: Buried Sterimol L, steric effects within the first coordination sphere
EE (Enantiomeric Excess)
Generation method: Q2MM (Quantum-guided Molecular Mechanics)
Reactions: 872 catalyst-substrate pairs involving 253 Rhodium (Rh)-bound atropisomeric catalysts from chiral bisphosphine with 10 enamide substrates
Property: Enantiomeric excess (EE) for asymmetric catalysis
Availability: Proprietary-only (closed-source as of December 2025)
BDE (Bond Dissociation Energy)
Molecules: 5,915 organometallic catalysts (ML₁L₂ structure)
Initial conformers: OpenBabel with geometric optimization
Energies: DFT calculations
Property: Electronic binding energy (difference in minimum energies of bound-catalyst complex and unbound catalyst)
Key constraint: DFT optimization for full conformer ensembles computationally infeasible (2-3 days per molecule)
Benchmark Setup
Task: Predict molecular properties from structure using different representation strategies (1D/2D/3D/Ensemble). The ground-truth regression targets are calculated as the Boltzmann-averaged value of the property across the conformer ensemble:
$$ \langle y \rangle_{k_B} = \sum_{\mathbf{C}_i \in \mathcal{C}} p_i y_i $$
Where $p_i$ is the conformer probability (Boltzmann weight) under experimental conditions derived from the conformer energy $e_i$:
$$ p_i = \frac{\exp(-e_i / k_B T)}{\sum_j \exp(-e_j / k_B T)} $$
Data splits: Datasets are partitioned 70% train, 10% validation, and 20% test.
Model categories:
- 1D Models: SMILES-based (Random Forest on concatenated MACCS/ECFP/RDKit fingerprints, LSTM, Transformer).
- 2D Models: Graph-based (GIN, GIN+VN, ChemProp, GraphGPS).
- 3D Models: Single conformer (SchNet, DimeNet++, GemNet, PaiNN, ClofNet, LEFTNet). For evaluation, single 3D models exclusively ingest the lowest-energy conformer. This baseline setting often yields strong performance but is unrealistic in practice, as identifying the global minimum requires exhaustively searching the entire conformer space.
- Ensemble Models: Full conformer ensemble processing via explicit set encoders. For each conformer embedding $\mathbf{z}_i$, three aggregation strategies are evaluated:
Mean Pooling: $$ \mathbf{s}_{\text{MEAN}} = \frac{1}{|\mathcal{C}|} \sum_{i=1}^{|\mathcal{C}|} \mathbf{z}_i $$
DeepSets: $$ \mathbf{s}_{\text{DS}} = g\left(\sum_{i=1}^{|\mathcal{C}|} h(\mathbf{z}_i)\right) $$
Self-Attention: $$ \begin{aligned} \mathbf{s}_{\text{ATT}} &= \sum_{i=1}^{|\mathcal{C}|} \mathbf{c}_i, \quad \text{where} \quad \mathbf{c}_i = g\left( \sum_{j=1}^{|\mathcal{C}|} \alpha_{ij} h(\mathbf{z}_j) \right) \\ \alpha_{ij} &= \frac{\exp\left((\mathbf{W} h(\mathbf{z}_i))^\top (\mathbf{W} h(\mathbf{z}_j))\right)}{\sum_{k=1}^{|\mathcal{C}|} \exp\left((\mathbf{W} h(\mathbf{z}_i))^\top (\mathbf{W} h(\mathbf{z}_k))\right)} \end{aligned} $$
Evaluation metric: Mean Absolute Error (MAE) for all tasks.
Key Findings
Ensemble superiority (task-dependent): Across benchmarks, explicitly modeling the full conformer set using DeepSets often achieved top performance. However, these improvements are not uniform:
- Small-Scale Success: Ensemble methods show large improvements on tasks like Kraken (Ensemble PaiNN achieves 0.2225 on $B_5$ vs 0.3443 single) and EE (Ensemble GemNet achieves 11.61% vs 18.03% single).
- Large-Scale Plateau: The performance improvements did not strongly transfer to large subsets like Drugs-75K (best ensemble strategy for GemNet achieves 0.4066 eV on IP vs 0.4069 eV single). The authors conjecture that the computational burden of encoding all conformers in each ensemble alters learning dynamics and increases training difficulty.
Conformer Sampling for Noise: Data augmentation (randomly sampling one conformer from an ensemble during training) improves performance and robustness when underlying conformers are imprecise (e.g., the forcefield-generated conformers in the BDE subset).
3D vs 2D: 3D models generally outperform 2D graph models, especially for conformationally-sensitive properties, though 1D and 2D methods remain highly competitive on low-resource datasets or less rotation-sensitive properties.
Model architecture: No single model dominates all tasks. GemNet and LEFTNet excel on large-scale Drugs-75K, while DimeNet++ shows strong performance on smaller Kraken and reaction datasets. Model selection depends on dataset size and task characteristics.
Reproducibility Details
| Artifact | Type | License | Notes |
|---|---|---|---|
| SXKDZ/MARCEL | Code + Dataset | Apache-2.0 | Benchmark suite, dataset loaders, and hyperparameter configs |
| Drugs-75K | Dataset | Apache-2.0 | DFT-level conformers and energies derived from GEOM-Drugs |
| Kraken | Dataset | Copyright retained by original authors | Conformer ensembles and four steric descriptors |
| BDE | Dataset | Apache-2.0 | OpenBabel-generated conformers with DFT binding energies |
| EE | Dataset | Proprietary | Closed-source as of 2026 |
- Data: The Drugs-75K, Kraken, and BDE subsets are openly available via the project’s GitHub repository. The EE dataset remains closed-source/proprietary (as of 2026), making the EE suite of the benchmark currently irreproducible.
- Code: The benchmark suite and PyTorch-Geometric dataset loaders are open-sourced at GitHub (SXKDZ/MARCEL) under the Apache-2.0 license.
- Hardware: The authors trained models using Nvidia A100 (40GB) GPUs. Memory-intensive models (e.g., GemNet, LEFTNet) required Nvidia H100 (80GB) GPUs. Total computation across all benchmark experiments was approximately 6,000 GPU hours.
- Algorithms/Models: Hyperparameters for all 18 evaluated models are provided in the repository configuration files (
benchmarks/params). All baseline models use publicly available frameworks (e.g., PyTorch Geometric, OGB, RDKit). - Evaluation: Evaluation scripts are provided in the repository with consistent tracking of Mean Absolute Error (MAE) and proper configuration of benchmark splits.
Citation
@inproceedings{zhu2024learning,
title={Learning Over Molecular Conformer Ensembles: Datasets and Benchmarks},
author={Yanqiao Zhu and Jeehyun Hwang and Keir Adams and Zhen Liu and Bozhao Nan and Brock Stenfors and Yuanqi Du and Jatin Chauhan and Olaf Wiest and Olexandr Isayev and Connor W. Coley and Yizhou Sun and Wei Wang},
booktitle={The Twelfth International Conference on Learning Representations},
year={2024},
url={https://openreview.net/forum?id=NSDszJ2uIV}
}
