Dataset Examples
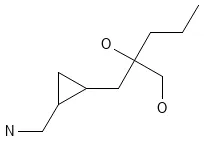
CCCC(O)(CO)CC1CC1CN)Dataset Subsets
| Subset | Size | Description |
|---|---|---|
| C/N/O Set | ~910.1M | Molecules containing up to 13 atoms of Carbon, Nitrogen, and Oxygen. |
| Cl/S Set | ~67.3M | Molecules containing up to 13 atoms, adding Sulfur (aromatic heterocycles, sulfones, sulfonamides, thioureas) and Chlorine (aromatic substituents). |
Related Datasets
| Dataset | Relationship | Link |
|---|---|---|
| GDB-11 | Predecessor | Notes |
| GDB-17 | Successor | Notes |
Key Contribution
The creation and release of the 977.5 million-compound GDB-13, a significant expansion in molecular size (up to 13 atoms) and elemental diversity (including S and Cl) made possible by key algorithmic optimizations that significantly accelerated the enumeration process.
Overview
GDB-13 extends the systematic enumeration of drug-like chemical space to molecules containing up to 13 atoms of Carbon, Nitrogen, Oxygen, Sulfur, and Chlorine. Building on the methodology established in GDB-11, this database represents a 37-fold increase in size while maintaining 100% Lipinski compliance for virtual screening applications. The enumeration results in a vast array of cyclic topologies, where 54% of the database comprises molecules with at least one three- or four-membered ring.
Strengths
- Systematic coverage of structures with up to 13 atoms
- High drug-likeness: 100% Lipinski compliance and 99.5% Vieth compliance
- High proportion of leadlike (98.9%) and fragmentlike (45.1%) molecules
- Structural novelty providing fragments absent from established databases like ZINC, ACX, and PubChem
Limitations
- Limited to small molecules with up to 13 atoms of C, N, O, S, and Cl
- Omits 66.2% of known chemical space up to 13 atoms found in external databases
- Excludes specific nonenumerated elements (F, Br, I, P, Si, metals) and functional groups (chlorine on nonaromatic carbons, mercaptans, sulfoxides, enamines, allenes)
- Excludes highly strained molecules and highly polar combinations
- Consists entirely of computer-generated structures pending experimental validation
Technical Notes
Algorithmic Approach
Type: Rule-Based Combinatorial Graph Enumeration
This approach relies on combinatorial enumeration. It utilizes a rule-based graph generation algorithm (GENG) paired with chemical stability filters to construct the dataset.
Process:
- Start with mathematical graphs representing saturated hydrocarbons up to 13 nodes using GENG (non-planar graphs discarded)
- Apply topological filters to remove highly strained small ring systems (e.g., fused cyclopropanes and bridgehead 3/4-membered rings)
- Generate 3D structures via CORINA or ChemAxon to apply a 3D volume-based strain filter. The local strain of a tetravalent carbon is estimated by the volume $V$ of the tetrahedron formed by extending a $1 \text{ \AA}$ line along its four single bonds. Hydrocarbons with planar or pyramidal carbon centers are discarded if: $$ V < 0.345 \text{ \AA}^3 $$
- Introduce unsaturations and heteroatoms through systematic substitution
- Apply chemical rule filters and element-ratio heuristics to ensure stability and drug-likeness
- Apply post-processing algorithms to introduce nitro groups, nitriles, aromatic chlorines, thiophenes, sulfonamides, and thioureas
Key Optimization: Replaced computationally expensive MM2 minimization (used in GDB-11) with a fast geometry-based estimation of strained polycyclic ring systems, combined with fast “element-ratio” filters. This achieved a 6.4-fold speedup in structure validation early in the pipeline.
Differences from GDB-11
- Element Selection: Fluorine removed from allowed elements; sulfur and chlorine added for higher drug relevance (e.g., thiophenes, sulfonamides).
- Optimization Method: MM2-based structure optimization replaced with a much faster, custom geometry-based estimation of local strain (measuring the tetrahedron volume of carbon centers).
- Heuristic Filters: Fast elemental ratio filters added to quickly reject highly polar, unstable combinations early in the pipeline.
Reproducibility Details
Paper & Data Availability
- Paper Access: The original paper is published in the Journal of the American Chemical Society (JACS) and is closed-access/paywalled. No open-access preprint exists on arXiv or ChemRxiv.
- Data Access: The full GDB-13 database and its subsets are freely available via the Reymond Group Downloads Page and are persistently hosted on Zenodo.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| GDB-13 Database (Reymond Group) | Dataset | Free download | Official download page hosted by the Reymond Group |
| GDB-13 on Zenodo | Dataset | Unknown | Persistent archival copy |
Source Code & Algorithms
The exact custom source code (e.g., GENG orchestration, local strain filters) is not publicly available. Researchers must re-implement the rules strictly described in the paper and supplementary materials.
Heuristic Filters
Implemented element-ratio filters derived from analyzing known compound databases to reject chemically unstable or highly polar molecules early in the generation pipeline:
$$ \begin{aligned} \frac{N + O}{C} &< 1.0 \\ \frac{N}{C} &< 0.571 \\ \frac{O}{C} &< 0.666 \end{aligned} $$
Excluded Functional Groups
- O-O bonds (peroxides)
- Hemiacetals, aminals, acyclic imines, non-aromatic enols
- Compounds containing both primary/secondary amines and aldehydes/ketones
- Nonenumerated elements (F, Br, I, P, Si, metals)
- High-heteroatom ratio structures (e.g., mannitol)
Hardware & Compute
- Compute Cost: ~40,000 CPU hours for the 910 million C/N/O structures.
- Infrastructure: Executed in parallel on a 500-node cluster
- Assembly Optimization: The switch from MM2 minimization to geometry-based estimation of strained polycyclic ring systems, alongside element-ratio filters, reduced assembly time 6.4-fold comparing GDB-11 workloads (1600 CPU hours to 250 CPU hours).
Citation
@article{blum2009gdb13,
title={970 million druglike small molecules for virtual screening in the chemical universe database GDB-13},
author={Blum, Lorenz C and Reymond, Jean-Louis},
journal={Journal of the American Chemical Society},
volume={131},
number={25},
pages={8732--8733},
year={2009},
publisher={ACS Publications},
doi={10.1021/ja902302h}
}
