Paper Information
Citation: Schoenmaker, L., Béquignon, O. J. M., Jespers, W., & van Westen, G. J. P. (2023). UnCorrupt SMILES: a novel approach to de novo design. Journal of Cheminformatics, 15, 22.
Publication: Journal of Cheminformatics, 2023
Additional Resources:
@article{schoenmaker2023uncorrupt,
title={UnCorrupt SMILES: a novel approach to de novo design},
author={Schoenmaker, Linde and B{\'e}quignon, Olivier J. M. and Jespers, Willem and van Westen, Gerard J. P.},
journal={Journal of Cheminformatics},
volume={15},
pages={22},
year={2023},
publisher={Springer},
doi={10.1186/s13321-023-00696-x}
}
A Transformer-Based SMILES Error Corrector
This is a Method paper that proposes a post hoc approach to fixing invalid SMILES produced by de novo molecular generators. Rather than trying to prevent invalid outputs through alternative representations (SELFIES) or constrained architectures (graph models), the authors train a transformer model to translate invalid SMILES into valid ones. The corrector is framed as a sequence-to-sequence translation task, drawing on techniques from grammatical error correction (GEC) in natural language processing.
The Problem of Invalid SMILES in Molecular Generation
SMILES-based generative models produce some percentage of invalid outputs that cannot be converted to molecules. The invalidity rate varies substantially across model types:
- RNN models (DrugEx): 5.7% invalid (pretrained) and 4.7% invalid (target-directed)
- GANs (ORGANIC): 9.5% invalid
- VAEs (GENTRL): 88.9% invalid
These invalid outputs represent wasted computation and potentially introduce bias toward molecules that are easier to generate correctly. Previous approaches to this problem include using alternative representations (DeepSMILES, SELFIES) or graph-based models, but these either limit the search space or increase computational cost. The authors propose a complementary strategy: fix the errors after generation.
Error Taxonomy Across Generator Types
The paper classifies invalid SMILES errors into six categories based on RDKit error messages:
- Syntax errors: malformed SMILES grammar
- Unclosed rings: unmatched ring closure digits
- Parentheses errors: unbalanced open/close parentheses
- Bond already exists: duplicate bonds between the same atoms
- Aromaticity errors: atoms incorrectly marked as aromatic or kekulization failures
- Valence errors: atoms exceeding their maximum bond count
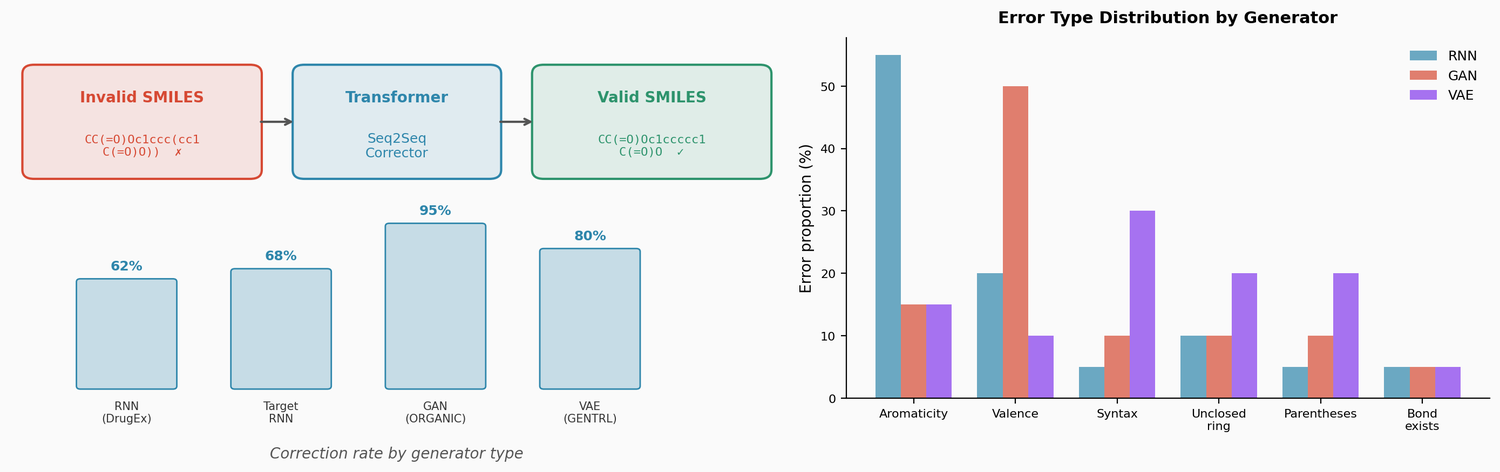
The distribution of error types differs across generators. RNN-based models primarily produce aromaticity errors, suggesting they learn SMILES grammar well but struggle with chemical validity. The GAN (ORGANIC) produces mostly valence errors. The VAE (GENTRL) produces more grammar-level errors (syntax, parentheses, unclosed rings), indicating that sampling from the continuous latent space often produces sequences that violate basic SMILES structure.
Architecture and Training
The SMILES corrector uses a standard encoder-decoder transformer architecture based on Vaswani et al., with learned positional encodings. Key specifications:
- Embedding dimension: 256
- Encoder/decoder layers: 3 each
- Attention heads: 8
- Feed-forward dimension: 512
- Dropout: 0.1
- Optimizer: Adam (learning rate 0.0005)
- Training: 20 epochs, batch size 16
Since no dataset of manually corrected invalid-valid SMILES pairs exists, the authors create synthetic training data by introducing errors into valid SMILES from the Papyrus bioactivity dataset (approximately 1.3M pairs). Errors are introduced through random perturbations following SMILES syntax rules: character substitutions, bond order changes, fragment additions from the GDB-8 database to atoms with full valence, and other structural modifications.
Training with Multiple Errors Improves Correction
A key finding is that training the corrector on inputs with multiple errors per SMILES substantially improves performance on real generator outputs. The baseline model (1 error per input) fixes 35-80% of invalid outputs depending on the generator. Increasing errors per training input to 12 raises this to 62-95%:
| Generator | 1 error/input | 12 errors/input |
|---|---|---|
| RNN (DrugEx) | ~60% fixed | 62% fixed |
| Target-directed RNN | ~60% fixed | 68% fixed |
| GAN (ORGANIC) | ~80% fixed | 95% fixed |
| VAE (GENTRL) | ~35% fixed | 80% fixed |
Training beyond 12 errors per input yields diminishing returns (80% average at 20 errors vs. 78% at 12). The improvement from multi-error training is consistent with GEC literature, where models learn to “distrust” inputs more when exposed to higher error rates.
The model also shows low overcorrection: only 14% of valid SMILES are altered during translation, comparable to overcorrection rates in spelling correction systems.
Fixed Molecules Are Comparable to Generator Outputs
The corrected molecules are evaluated against both the training set and the readily generated (valid) molecules from each generator:
- Uniqueness: 97% of corrected molecules are unique
- Novelty vs. generated: 97% of corrected molecules are novel compared to the valid generator outputs
- Similarity to nearest neighbor (SNN): 0.45 between fixed and generated sets, indicating the corrected molecules explore different parts of chemical space
- Property distributions: KL divergence scores between fixed molecules and the training set are comparable to those between generated molecules and the training set
This demonstrates that SMILES correction produces molecules that are as chemically reasonable as the generator’s valid outputs while exploring complementary regions of chemical space.
Local Chemical Space Exploration via Error Introduction
Beyond fixing generator errors, the authors propose using the SMILES corrector for analog generation. The workflow is:
- Take a known active molecule
- Introduce random errors into its SMILES (repeated 1000 times)
- Correct the errors using the trained corrector
This “local sequence exploration” generates novel analogs with 97% validity. The uniqueness (39%) and novelty (16-37%) are lower than for generator correction because the corrector often regenerates the original molecule. However, the approach produces molecules that are structurally similar to the starting compound (SNN of 0.85 to known ligands).
The authors demonstrate this on selective Aurora kinase B (AURKB) inhibitors. The generated analogs occupy the same binding site region as the co-crystallized ligand VX-680 in docking studies, with predicted bioactivities similar to known compounds. Compared to target-directed RNN generation, SMILES exploration produces molecules closer to known actives (higher SNN, scaffold similarity, and KL divergence scores).
Limitations
The corrector performance drops when applied to real generator outputs compared to synthetic test data, because the synthetic error distribution does not perfectly match the errors that generators actually produce. Generator-specific correctors trained on actual invalid outputs could improve performance. The local exploration approach has limited novelty since the corrector frequently regenerates the original molecule. The evaluation uses predicted rather than experimental bioactivities for the Aurora kinase case study.
Reproducibility
| Artifact | Type | License | Notes |
|---|---|---|---|
| LindeSchoenmaker/SMILES-corrector | Code + Data | MIT | Training code, synthetic error generation, and evaluation scripts |
Data: Synthetic training pairs derived from the Papyrus bioactivity dataset (v5.5). Approximately 1.3M invalid-valid pairs per error-count setting.
Code: Transformer implemented in PyTorch, adapted from Ben Trevett’s seq2seq tutorial. Generative model baselines use DrugEx, GENTRL, and ORGANIC.
Evaluation: Validity assessed with RDKit. Similarity metrics (SNN, fragment, scaffold) and KL divergence computed following MOSES and GuacaMol benchmark protocols.