A Systematization of Transformer Architectures for Molecular Science
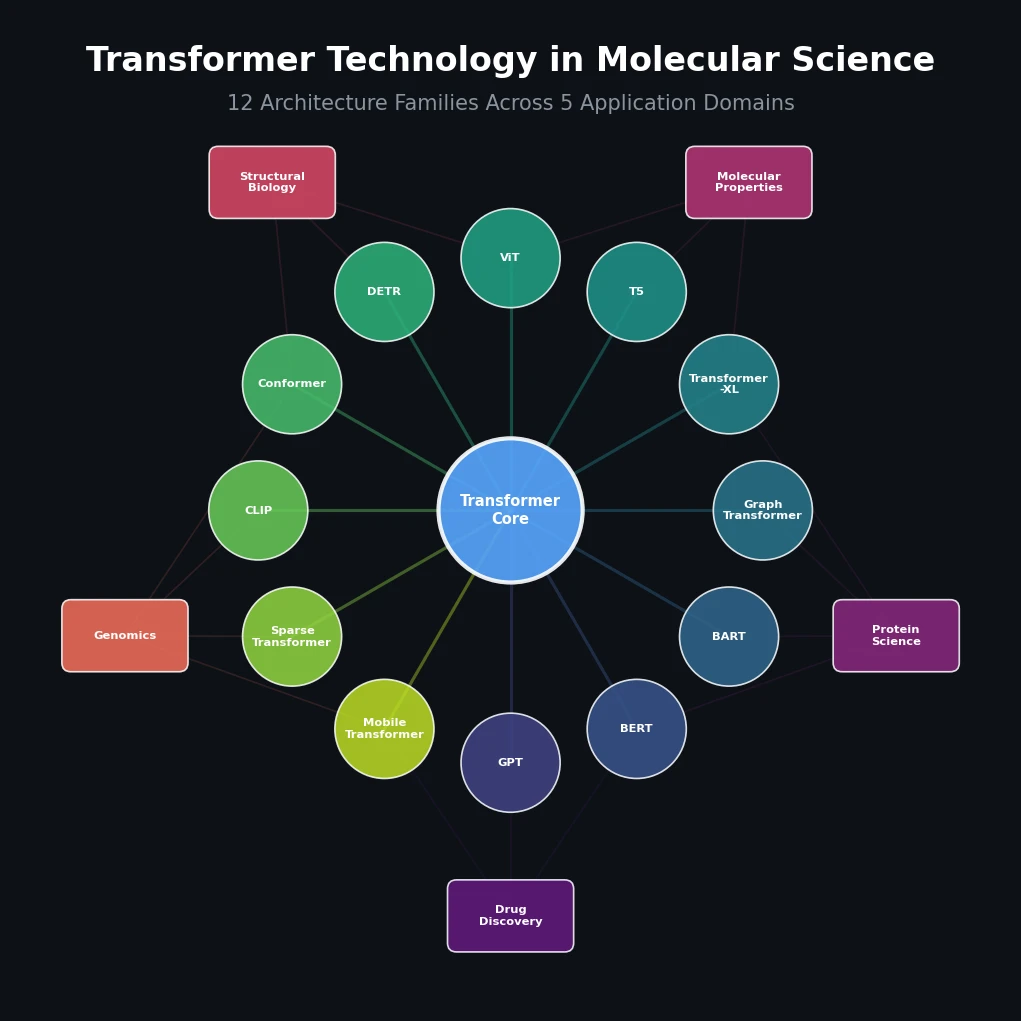
This paper is a Systematization review. It organizes and taxonomizes 12 families of transformer architectures that have been applied across molecular science, including chemistry, biology, and drug discovery. The primary contribution is not a new method or dataset, but a structured technical overview of the algorithmic internals of each transformer variant and their specific applications to molecular problems. The review covers 201 references and provides a unified treatment of how these architectures capture molecular patterns from sequential, graphical, and image-based data.
Bridging the Gap Between Transformer Variants and Molecular Applications
Transformer-based models have become widespread in molecular science, yet the authors identify a gap: there is no organized taxonomy linking these diverse techniques in the existing literature. Individual papers introduce specific architectures or applications, but practitioners lack a unified reference that explains the technical differences between GPT, BERT, BART, graph transformers, and other variants in the context of molecular data. The review aims to fill this gap by providing an in-depth investigation of the algorithmic components of each model family, explaining how their architectural innovations contribute to processing complex molecular data. The authors note that the success of transformers in molecular science stems from several factors: the sequential nature of chemical and biological molecules (DNA, RNA, proteins, SMILES strings), the attention mechanism’s ability to capture long-range dependencies within molecular structures, and the capacity for transfer learning through pre-training on large chemical and biological datasets.
Twelve Transformer Families and Their Molecular Mechanisms
The review covers transformer preliminaries before diving into 12 specific architecture families. The core self-attention mechanism computes:
$$ \text{Attention}(Q, K, V) = \text{softmax}\left(\frac{QK^T}{\sqrt{d_k}}\right)V $$
where $d_k$ is the dimension of the key vectors. The position-wise feed-forward network is:
$$ \text{FFN}(x) = \max(0, xW_1 + b_1)W_2 + b_2 $$
The 12 architecture families covered are:
GPT (Generative Pre-trained Transformer): Uses the decoder part of the transformer for autoregressive generation. Applications include MolGPT for molecular generation, DrugGPT for protein-ligand binding, and cMolGPT for target-specific de novo molecular generation.
BERT (Bidirectional Encoder Representations from Transformers): Uses transformer encoders with masked language modeling and next-sentence prediction for pre-training. Molecular applications include FP-BERT for molecular property prediction using composite fingerprint representations, Graph-BERT for protein-protein interaction identification, SMILES-BERT, and Mol-BERT.
BART (Bidirectional and Auto-Regressive Transformers): Functions as a denoising autoencoder with both encoder and decoder. Molecular applications include Chemformer for sequence-to-sequence chemistry tasks, MS2Mol for mass spectrometry analysis, and MolBART for molecular feature learning.
Graph Transformer: Leverages self-attention on graph-structured data to capture global context. Applications include GraphSite for protein-DNA binding site prediction (using AlphaFold2 structure predictions), KPGT for knowledge-guided molecular graph pre-training, and PAGTN for establishing long-range dependencies in molecular graphs.
Transformer-XL: Incorporates relative positional encoding for modeling long sequences. Used for small molecule retention time prediction, drug design with CHEMBL data (1.27 million molecules), and Heck reaction generation.
T5 (Text-to-Text Transfer Transformer): Unifies NLP tasks into text-to-text mapping. T5Chem was pre-trained on 97 million molecules from PubChem and achieved 99.5% accuracy on reaction classification (USPTO 500 MT). C5T5 uses IUPAC naming for molecular optimization in drug discovery.
Vision Transformer (ViT): Applies transformer architecture to image patches. Used for organic molecule classification (97% accuracy with WGAN-generated data), bacterial identification via SERS, and molecular property prediction from mass spectrometry data (TransG-Net).
DETR (Detection Transformer): End-to-end object detection using transformers. Applied to cryo-EM particle picking (TransPicker), molecular structure image recognition (IMG2SMI), and cell segmentation (Cell-DETR).
Conformer: Integrates convolutional modules into transformer structure. Used for DNA storage error correction (RRCC-DNN), drug-target affinity prediction (NG-DTA with Davis and Kiba datasets).
CLIP (Contrastive Language-Image Pre-training): Multimodal learning linking text and images. Applied to peptide design (Cut&CLIP for protein degradation), gene identification (pathCLIP), and drug discovery (CLOOME for zero-shot transfer learning).
Sparse Transformers: Use sparse attention matrices to reduce complexity to $O(n\sqrt{n})$. Applied to drug-target interaction prediction with gated cross-attention mechanisms.
Mobile and Efficient Transformers: Compressed variants (TinyBERT, MobileBERT) for resource-constrained environments. Molormer uses ProbSparse self-attention for drug-drug interaction prediction. LOGO is a lightweight pre-trained language model for non-coding genome interpretation.
Survey Organization and Coverage of Molecular Domains
As a survey paper, this work does not present new experiments. Instead, it catalogues existing applications across multiple molecular domains:
Drug Discovery and Design: GPT-based ligand design (DrugGPT), BART-based molecular generation (Chemformer, MolBART), graph transformer pre-training for molecular property prediction (KPGT), T5-based chemical reaction prediction (T5Chem), and sparse transformer methods for drug-target interactions.
Protein Science: BERT-based protein-protein interaction prediction (Graph-BERT), graph transformer methods for protein-DNA binding (GraphSite with AlphaFold2 integration), conformer-based drug-target affinity prediction (NG-DTA), and CLIP-based peptide design (Cut&CLIP).
Molecular Property Prediction: FP-BERT for fingerprint-based prediction, SMILES-BERT and Mol-BERT for end-to-end prediction from SMILES, KPGT for knowledge-guided graph pre-training, and Transformer-XL for property modeling with relative positional encoding.
Structural Biology: DETR-based cryo-EM particle picking (TransPicker), vision transformer applications in cell imaging, and Cell-DETR for instance segmentation in microscopy.
Genomics: Conformer-based DNA storage error correction (RRCC-DNN), LOGO for non-coding genome interpretation, and MetaTransformer for metagenomic sequencing analysis.
Future Directions and Limitations of the Survey
The review concludes with four future directions:
ChatGPT integration into molecular science: Using LLMs for data analysis, literature review, and hypothesis generation in chemistry and biology.
Multifunction transformers: Models that extract features across diverse molecular structures and sequences simultaneously.
Molecular-aware transformers: Architectures that handle multiple data types (text, sequence, structure, image, energy, molecular dynamics, function) in a unified framework.
Self-assessment transformers and superintelligence: Speculative discussion of models that learn from seemingly unrelated data sources.
The review has several limitations worth noting. The coverage is broad but shallow: each architecture family receives only 1-2 pages of discussion, and the paper largely describes existing work rather than critically evaluating it. The review does not systematically compare the architectures against each other on common benchmarks. The future directions section (particularly the superintelligence discussion) is speculative and lacks concrete proposals. The paper also focuses primarily on technical architecture descriptions rather than analyzing failure modes, scalability challenges, or reproducibility concerns across the surveyed methods. As a review article, no new data were created or analyzed.
Reproducibility Details
Data
This is a survey paper. No new datasets were created or used. The paper reviews applications involving datasets such as PubChem (97 million molecules for T5Chem), CHEMBL (1.27 million molecules for Transformer-XL drug design), USPTO 500 MT (reaction classification), ESOL (5,328 molecules for property prediction), and Davis/Kiba (drug-target affinity).
Algorithms
No new algorithms are introduced. The paper provides mathematical descriptions of the core transformer components (self-attention, positional encoding, feed-forward networks, layer normalization) and describes how 12 architecture families modify these components.
Models
No new models are presented. The paper surveys existing models including MolGPT, DrugGPT, FP-BERT, SMILES-BERT, Chemformer, MolBART, GraphSite, KPGT, T5Chem, TransPicker, Cell-DETR, CLOOME, and Molormer, among others.
Evaluation
No new evaluation is performed. Performance numbers cited from the literature include: T5Chem reaction classification accuracy of 99.5%, ViT organic molecule classification at 97%, Transformer-XL property prediction RMSE of 0.6 on ESOL, and Heck reaction generation feasibility rate of 47.76%.
Hardware
No hardware requirements are specified, as this is a survey paper.
| Artifact | Type | License | Notes |
|---|---|---|---|
| Paper (open access) | Paper | CC-BY-NC-ND | Open access via Wiley |
Paper Information
Citation: Jiang, J., Ke, L., Chen, L., Dou, B., Zhu, Y., Liu, J., Zhang, B., Zhou, T., & Wei, G.-W. (2024). Transformer technology in molecular science. WIREs Computational Molecular Science, 14(4), e1725. https://doi.org/10.1002/wcms.1725
@article{jiang2024transformer,
title={Transformer technology in molecular science},
author={Jiang, Jian and Ke, Lu and Chen, Long and Dou, Bozheng and Zhu, Yueying and Liu, Jie and Zhang, Bengong and Zhou, Tianshou and Wei, Guo-Wei},
journal={WIREs Computational Molecular Science},
volume={14},
number={4},
pages={e1725},
year={2024},
publisher={Wiley},
doi={10.1002/wcms.1725}
}