A Systematization of Transformers in Chemistry
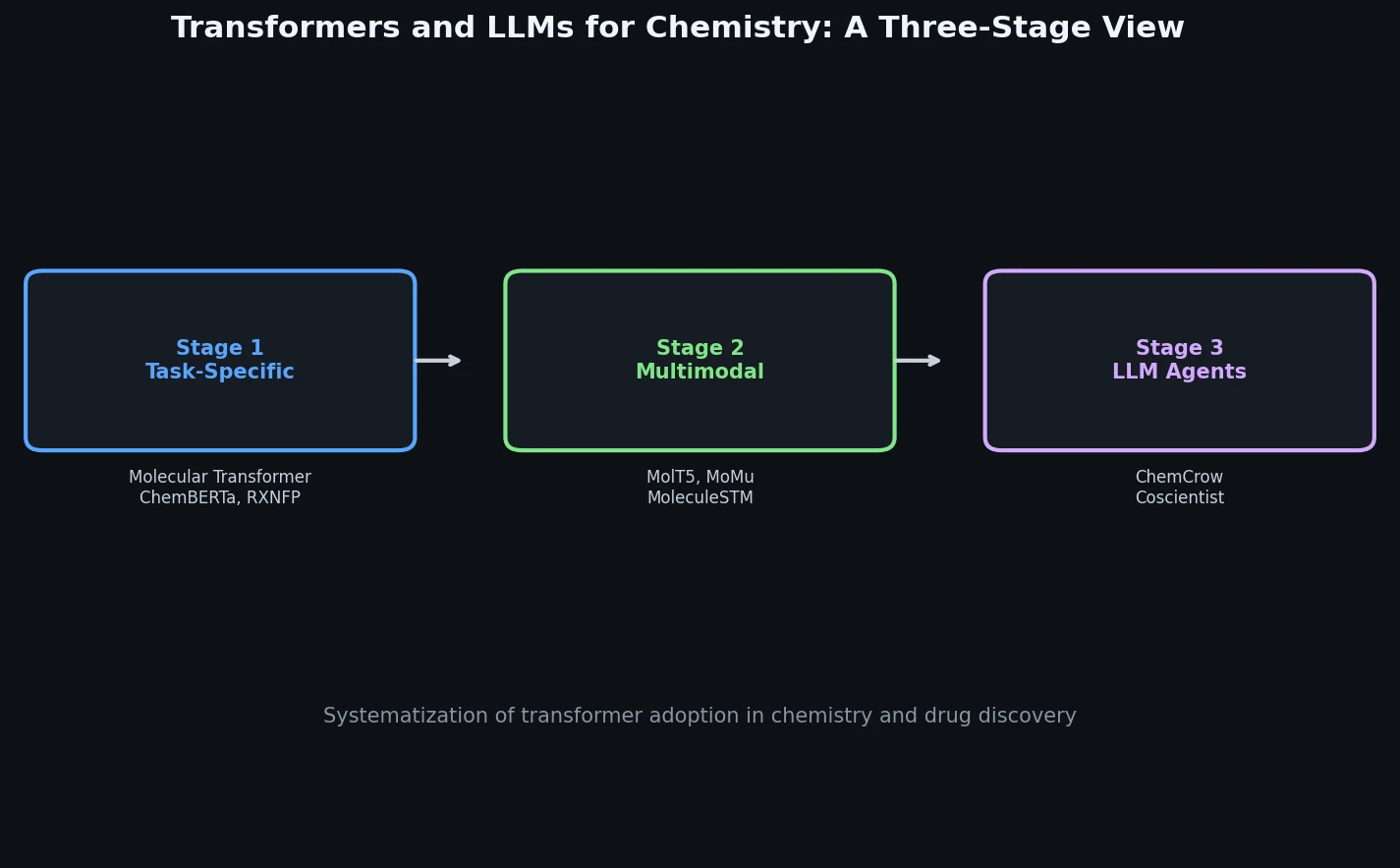
This book chapter by Bran and Schwaller is a Systematization paper that organizes the growing body of work applying transformer architectures to chemistry and drug discovery. Rather than proposing a new method, the authors trace a three-stage evolution: (1) task-specific single-modality models operating on SMILES and reaction strings, (2) multimodal models bridging molecular representations with spectra, synthesis actions, and natural language, and (3) large language models and LLM-powered agents capable of general chemical reasoning.
Why Transformers for Chemistry?
The authors motivate the review by drawing analogies between natural language and chemical language. Just as text can be decomposed into subwords and tokens, molecules can be linearized into SMILES or SELFIES strings, and chemical reactions can be encoded as reaction SMILES. This structural parallel enabled direct transfer of transformer architectures, originally designed for machine translation, to chemical prediction tasks.
Several factors accelerated this adoption:
- The publication of open chemical databases and benchmarks (e.g., MoleculeNet, Open Reaction Database, Therapeutics Data Commons)
- Improvements in compute infrastructure and training algorithms
- The success of attention mechanisms at capturing context-dependent relationships, which proved effective for learning chemical grammar and atom-level correspondences
The review positions the transformer revolution in chemistry as a natural extension of NLP advances, noting that the gap between chemical and natural language is progressively closing.
Molecular Representations as Language
A key section of the review covers text-based molecular representations that make transformer applications possible:
- SMILES (Simplified Molecular Input Line Entry System): The dominant linearization scheme since the 1980s, encoding molecular graphs as character sequences with special symbols for bonds, branches, and rings.
- SELFIES (Self-Referencing Embedded Strings): A newer representation that guarantees every string maps to a valid molecule, addressing the robustness issues of SMILES in generative settings.
- Reaction SMILES: Extends molecular representations to encode full chemical reactions in the format “A.B > catalyst.reagent > C.D”, enabling reaction prediction as a sequence-to-sequence task.
The authors note that while IUPAC names, InChI, and DeepSMILES exist as alternatives, SMILES and SELFIES dominate practical applications.
Stage 1: Task-Specific Transformer Models
The first stage of transformer adoption focused on clearly defined chemical tasks, with models trained on a single data modality (molecular strings).
Chemical Translation Tasks
The encoder-decoder architecture was directly applied to tasks framed as translation:
- Molecular Transformer (Schwaller et al.): Treated reaction prediction as translation from reactant SMILES to product SMILES, becoming a leading method for forward synthesis prediction.
- Retrosynthetic planning: The reverse task, predicting reactants from products, with iterative application to construct full retrosynthetic trees mapping to commercially available building blocks.
- Chemformer (Irwin et al.): A pre-trained model across multiple chemical tasks, offering transferability to new applications with improved performance.
- Graph-to-sequence models (Tu and Coley): Used a custom graph encoder with a transformer decoder, achieving improvements through permutation-invariant molecular graph encoding.
Representation Learning and Feature Extraction
Encoder-only transformers proved valuable for generating molecular and reaction embeddings:
- Reaction representations (Wang et al., SMILES-BERT): Trained models to generate reaction vectors that outperformed hand-engineered features on downstream regression tasks.
- Reaction classification (Schwaller et al.): Replaced the decoder with a classification layer to map chemical reactions by class, revealing clustering patterns by reaction type, data source, and molecular properties.
- Yield prediction: Regression heads attached to encoders achieved strong results on high-throughput experimentation datasets.
- Protein language models (Rives et al., ESM): Trained on 250 million protein sequences using unsupervised learning, achieving strong performance on protein property prediction and structure forecasting.
- RXNMapper (Schwaller et al.): A notable application where attention weight analysis revealed that transformers internally learn atom-to-atom mappings in chemical reactions, leading to an open-source atom mapping algorithm that outperformed existing approaches.
Stage 2: Multimodal Chemical Models
The second stage extended transformers beyond molecular strings to incorporate additional data types:
- Molecular captioning: Describing molecules in natural language, covering scaffolds, sources, drug interactions, and other features (Edwards et al.).
- Bidirectional molecule-text conversion: Models capable of generating molecules from text queries and performing molecule-to-molecule tasks (Christofidellis et al.).
- Experimental procedure prediction: Generating actionable synthesis steps from reaction SMILES (Vaucher et al.), bridging the gap between retrosynthetic planning and laboratory execution.
- Structural elucidation from IR spectra: Encoding IR spectra as text sequences alongside chemical formulas, then predicting SMILES from these inputs (Alberts et al.), achieving 45% accuracy in structure prediction and surpassing prior approaches for functional group identification.
Stage 3: Large Language Models and Chemistry Agents
The most recent stage builds on foundation models pre-trained on vast text corpora, adapted for chemistry through fine-tuning and in-context learning.
Scaling Laws and Emergent Capabilities
The authors discuss how model scaling leads to emergent capabilities relevant to chemistry:
- Below certain compute thresholds, model performance on chemistry tasks appears random.
- Above critical sizes, sudden improvements emerge, along with capabilities like chain-of-thought (CoT) reasoning and instruction following.
- These emergent abilities enable chemistry tasks that require multi-step reasoning without explicit training on chemical data.
LLMs as Chemistry Tools
Key applications of LLMs in chemistry include:
- Fine-tuning for low-data chemistry (Jablonka et al.): GPT-3 fine-tuned on limited chemistry datasets performed comparably to, and sometimes exceeded, specialized models with engineered features for tasks like predicting transition wavelengths and phase classification.
- In-context learning: Providing LLMs with a few examples enables prediction on chemistry tasks without any parameter updates, particularly valuable when data is scarce.
- Bayesian optimization with LLMs (Ramos et al.): Using GPT models for uncertainty-calibrated regression, enabling catalyst and molecular optimization directly from synthesis procedures without feature engineering.
- 3D structure generation (Flam-Shepherd and Aspuru-Guzik): Using language models to generate molecular structures with three-dimensional atomic positions in XYZ, CIF, and PDB formats, matching graph-based algorithms while overcoming representation limitations.
LLM-Powered Chemistry Agents
The review highlights the agent paradigm as the most impactful recent development:
- 14 LLM use-cases (Jablonka et al.): A large-scale collaborative effort demonstrating applications from computational tool wrappers to reaction optimization assistants and scientific question answering.
- ChemCrow (Bran, Cox et al.): An LLM-powered agent equipped with curated computational chemistry tools, capable of planning and executing tasks across drug design, materials design, and synthesis. ChemCrow demonstrated that tool integration overcomes LLM hallucination issues by grounding responses in reliable data sources.
- Autonomous scientific research (Boiko et al.): Systems with focus on cloud laboratory operability.
The agent paradigm offers tool composability through natural language interfaces, allowing users to chain multiple computational tools into custom pipelines.
Outlook and Limitations
The authors identify several themes for the future:
- The three stages represent increasing generality, from task-specific single-modality models to open-ended agents.
- Natural language interfaces are progressively closing the gap between chemical and human language.
- Tool integration through agents provides grounding that mitigates hallucination, a known limitation of direct LLM application to chemistry.
- The review acknowledges that LLMs have a “high propensity to generate false and inaccurate content” on chemical tasks, making tool-augmented approaches preferable to direct application.
The chapter does not provide quantitative benchmarks or systematic comparisons across the methods discussed, as its goal is to organize the landscape rather than evaluate individual methods.
Reproducibility Details
This is a review/survey chapter and does not introduce new models, datasets, or experiments. The reproducibility assessment applies to the referenced works rather than the review itself.
Key Referenced Resources
Several open-source tools and datasets discussed in the review are publicly available:
| Artifact | Type | License | Notes |
|---|---|---|---|
| RXNMapper | Code | MIT | Attention-based atom mapping |
| ChemCrow | Code | MIT | LLM-powered chemistry agent |
| MoleculeNet | Dataset | Various | Molecular ML benchmarks |
| Open Reaction Database | Dataset | CC-BY-SA-4.0 | Curated reaction data |
| Therapeutics Data Commons | Dataset | MIT | Drug discovery ML datasets |
Reproducibility Classification
Not applicable (review paper). Individual referenced works range from Highly Reproducible (open-source models like RXNMapper, ChemCrow) to Partially Reproducible (some models without released code) to Closed (proprietary LLMs like GPT-3/GPT-4 used in fine-tuning studies).
Paper Information
Citation: Bran, A. M., & Schwaller, P. (2024). Transformers and Large Language Models for Chemistry and Drug Discovery. In Drug Development Supported by Informatics (pp. 143-163). Springer Nature Singapore. https://doi.org/10.1007/978-981-97-4828-0_8
@incollection{bran2024transformers,
title={Transformers and Large Language Models for Chemistry and Drug Discovery},
author={Bran, Andres M. and Schwaller, Philippe},
booktitle={Drug Development Supported by Informatics},
pages={143--163},
year={2024},
publisher={Springer Nature Singapore},
doi={10.1007/978-981-97-4828-0_8}
}