A Systematization of Transformer-Based Chemical Language Models
This paper is a Systematization (literature review) that surveys the landscape of transformer-based chemical language models (CLMs) operating on SMILES representations. It organizes the field into three architectural categories (encoder-only, decoder-only, encoder-decoder), discusses tokenization strategies, pre-training and fine-tuning methodologies, and identifies open challenges and future research directions. The review covers approximately 30 distinct CLMs published through early 2024.
Why Review Transformer CLMs for SMILES?
The chemical space is vast, with databases like ZINC20 exceeding 5.5 billion compounds, and the amount of unlabeled molecular data far outstrips available labeled data for specific tasks like toxicity prediction or binding affinity estimation. Traditional molecular representations (fingerprints, descriptors, graph-based methods) require expert-engineered features and extensive domain knowledge.
Transformer-based language models, originally developed for NLP, have emerged as a compelling alternative. By treating SMILES strings as a “chemical language,” these models can leverage large-scale unsupervised pre-training on abundant unlabeled molecules, then fine-tune on small labeled datasets for specific downstream tasks. Earlier approaches like Seq2Seq and Seq3Seq fingerprint methods used RNN-based encoder-decoders, but these suffered from vanishing gradients and sequential processing bottlenecks when handling long SMILES sequences.
The authors motivate this review by noting that no prior survey has comprehensively organized transformer-based CLMs by architecture type while simultaneously covering tokenization, embedding strategies, and downstream application domains.
Architectural Taxonomy: Encoder, Decoder, and Encoder-Decoder Models
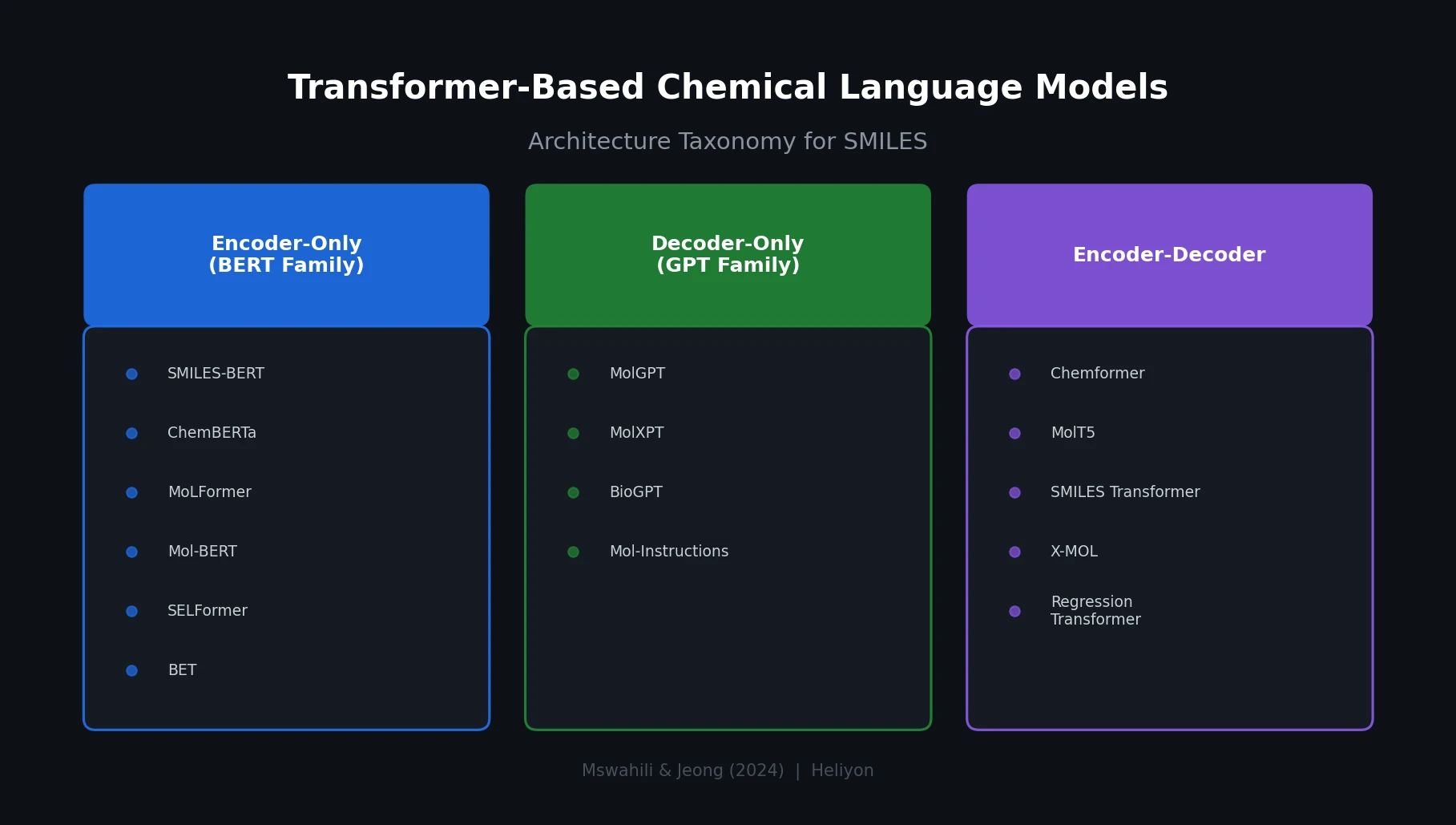
The core organizational contribution is a three-way taxonomy of transformer CLMs based on their architectural backbone.
Encoder-Only Models (BERT Family)
These models capture bidirectional context, making them well suited for extracting molecular representations for property prediction tasks. The review covers:
- BERT (Lee and Nam, 2022): Adapted for SMILES processing with linguistic knowledge infusion, using BPE tokenization
- MOLBERT (Fabian et al., 2020): Chemistry-specific BERT for physicochemical property and bioactivity prediction
- SMILES-BERT (Wang et al., 2019): BERT variant designed to learn molecular representations directly from SMILES without feature engineering
- ChemBERTa / ChemBERTa-2 (Chithrananda et al., 2020; Ahmad et al., 2022): RoBERTa-based models optimized for chemical property prediction, with ChemBERTa-2 exploring multi-task pre-training
- GPT-MolBERTa (Balaji et al., 2023): Combines GPT molecular features with a RoBERTa backbone
- MoLFormer (Ross et al., 2022): Large-scale model trained on 1.1 billion molecules, published in Nature Machine Intelligence
- SELFormer (Yuksel et al., 2023): Operates on SELFIES representations rather than SMILES
- Mol-BERT / MolRoPE-BERT (Li and Jiang, 2021; Liu et al., 2023): Differ in positional embedding strategy, with MolRoPE-BERT using rotary position embedding to handle longer sequences
- BET (Chen et al., 2021): Extracts predictive representations from hundreds of millions of molecules
Decoder-Only Models (GPT Family)
These models excel at generative tasks, including de novo molecular design:
- GPT-2-based model (Adilov, 2021): Generative pre-training from molecules
- MolXPT (Liu et al., 2023): Wraps molecules with text for generative pre-training, connecting chemical and natural language
- BioGPT (Luo et al., 2022): Focuses on biomedical text generation and mining
- MolGPT (Haroon et al., 2023): Uses relative attention to capture token distances and relationships for de novo drug design
- Mol-Instructions (Fang et al., 2023): Large-scale biomolecular instruction dataset for LLMs
Encoder-Decoder Models
These combine encoding and generation capabilities for sequence-to-sequence tasks:
- Chemformer (Irwin et al., 2022): BART-based model for reaction prediction and molecular property prediction
- MolT5 (adapted T5): Unified text-to-text framework for molecular tasks
- SMILES Transformer (Honda et al., 2019): Pre-trained molecular fingerprints for low-data drug discovery
- X-MOL (Xue et al., 2020): Large-scale pre-training for molecular understanding
- Regression Transformer (Born and Manica, 2023): Operates on SELFIES, enabling concurrent regression and generation
- TransAntivirus (Mao et al., 2023): Specialized for antiviral drug design using IUPAC nomenclature
Tokenization, Embedding, and Pre-Training Strategies
SMILES Tokenization
The review identifies tokenization as a critical preprocessing step that affects downstream performance. SMILES tokenization differs from standard NLP tokenization because SMILES strings lack whitespace and use parentheses for branching rather than sentence separation. The key approaches include:
| Strategy | Source | Description |
|---|---|---|
| Atom-in-SMILES (AIS) | Ucak et al. (2023) | Atom-level tokens preserving chemical identity |
| SMILES Pair Encoding (SPE) | Li and Fourches (2021) | BPE-inspired substructure tokenization |
| Byte-Pair Encoding (BPE) | Chithrananda et al. (2020); Lee and Nam (2022) | Standard subword tokenization adapted for SMILES |
| SMILESTokenizer | Chithrananda et al. (2020) | Character-level tokenization with chemical adjustments |
Positional Embeddings
The models use various positional encoding strategies: absolute, relative key, relative key-query, rotary (RoPE), and sinusoidal. Notably, SMILES-based models omit segmentation embeddings since SMILES data consists of single sequences rather than sentence pairs.
Pre-Training and Fine-Tuning Pipeline
The standard workflow follows two phases:
- Pre-training: Unsupervised training on large unlabeled SMILES databases (ZINC, PubChem, ChEMBL) using masked language modeling (MLM), where the model learns to predict masked tokens within SMILES strings
- Fine-tuning: Supervised adaptation on smaller labeled datasets for specific tasks (classification or regression)
The self-attention mechanism, central to all transformer CLMs, is formulated as:
$$ Z = \text{Softmax}\left(\frac{(XW^Q)(XW^K)^T}{\sqrt{d_k}}\right) XW^V $$
where $X \in \mathbb{R}^{N \times M}$ is the input feature matrix, $W^Q$, $W^K$, $W^V \in \mathbb{R}^{M \times d_k}$ are learnable weight matrices, and $\sqrt{d_k}$ is the scaling factor.
Benchmark Datasets and Evaluation Landscape
The review catalogs the standard evaluation ecosystem for CLMs. Pre-training databases include ZINC, PubChem, and ChEMBL. Fine-tuning and evaluation rely heavily on MoleculeNet benchmarks:
| Category | Datasets | Task Type | Example Size |
|---|---|---|---|
| Physical Chemistry | ESOL, FreeSolv, Lipophilicity | Regression | 642 to 4,200 |
| Biophysics | PCBA, MUV, HIV, PDBbind, BACE | Classification/Regression | 11,908 to 437,929 |
| Physiology | BBBP, Tox21, ToxCast, SIDER, ClinTox | Classification | 1,427 to 8,575 |
The authors also propose four new fine-tuning datasets targeting diseases: COVID-19 drug compounds, cocrystal formation, antimalarial drugs (Plasmodium falciparum targets), and cancer gene expression/drug response data.
Challenges, Limitations, and Future Directions
Current Challenges
The review identifies several persistent limitations:
- Data efficiency: Despite transfer learning, transformer CLMs still require substantial pre-training data, and labeled datasets for specific tasks remain scarce
- Interpretability: The complexity of transformer architectures makes it difficult to understand how specific molecular features contribute to predictions
- Computational cost: Training large-scale models demands significant GPU resources, limiting accessibility
- Handling rare molecules: Models struggle with molecular structures that deviate significantly from training data distributions
- SMILES limitations: Non-unique representations, invalid strings, exceeded atom valency, and inadequate spatial information capture
SMILES Representation Issues
The authors highlight five specific problems with SMILES as an input representation:
- Non-canonical representations reduce string uniqueness for the same molecule
- Many symbol combinations produce chemically invalid outputs
- Valid SMILES strings can encode chemically impossible molecules (e.g., exceeded valency)
- Spatial information is inadequately captured
- Syntactic and semantic robustness is limited
Future Research Directions
The review proposes several directions:
- Alternative molecular representations: Exploring SELFIES, DeepSMILES, IUPAC, and InChI beyond SMILES
- Role of SMILES token types: Strategic masking of metals, non-metals, bonds, and branches during MLM pre-training to identify which components are most critical
- Few-shot learning: Combining few-shot approaches with large-scale pre-trained CLMs for data-scarce scenarios
- Drug repurposing: Training CLMs to distinguish identical compounds with different biological activity profiles across therapeutic domains
- Improved benchmarks: Incorporating disease-specific datasets (malaria, cancer, COVID-19) for more realistic evaluation
- Ethical considerations: Addressing dual-use risks, data biases, and responsible open-source release of CLMs
Reproducibility Details
This is a literature review paper. It does not introduce new models, code, or experimental results. The reproducibility assessment focuses on the accessibility of the reviewed works and proposed datasets.
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Pre-training | ZINC20 | 5.5B+ compounds | Publicly available |
| Pre-training | PubChem | 100M+ compounds | Publicly available |
| Pre-training | ChEMBL | 2M+ compounds | Publicly available |
| Fine-tuning | MoleculeNet (8 datasets) | 642 to 437,929 | Standard benchmark suite |
| Proposed | COVID-19 drug compounds | 740 | From Harigua-Souiai et al. (2021) |
| Proposed | Cocrystal formation | 3,282 | From Mswahili et al. (2021) |
| Proposed | Antimalarial drugs | 4,794 | From Mswahili et al. (2024) |
| Proposed | Cancer gene/drug response | 201 drugs, 734 cell lines | From Kim et al. (2021) |
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| DAI Lab website | Other | N/A | Authors’ research lab |
No code, models, or evaluation scripts are released with this review. The paper does not include a supplementary materials section or GitHub repository.
Hardware
Not applicable (literature review).
Paper Information
Citation: Mswahili, M. E., & Jeong, Y.-S. (2024). Transformer-based models for chemical SMILES representation: A comprehensive literature review. Heliyon, 10(20), e39038. https://doi.org/10.1016/j.heliyon.2024.e39038
@article{mswahili2024transformer,
title={Transformer-based models for chemical {SMILES} representation: A comprehensive literature review},
author={Mswahili, Medard Edmund and Jeong, Young-Seob},
journal={Heliyon},
volume={10},
number={20},
pages={e39038},
year={2024},
publisher={Elsevier},
doi={10.1016/j.heliyon.2024.e39038}
}