An Empirical Comparison of Sequence Architectures for Molecular Generation
This is an Empirical paper that systematically compares two dominant sequence modeling architectures, recurrent neural networks (RNNs) and the Transformer, for chemical language modeling. The primary contribution is a controlled experimental comparison across three generative tasks of increasing complexity, combined with an evaluation of two molecular string representations (SMILES and SELFIES). The paper does not propose a new method; instead, it provides practical guidance on when each architecture is more appropriate for molecular generation.
Why Compare RNNs and Transformers for Molecular Design?
Exploring unknown molecular space and designing molecules with target properties is a central goal in computational drug design. Language models trained on molecular string representations (SMILES, SELFIES) have shown the capacity to learn complex molecular distributions. RNN-based models, including LSTM and GRU variants, were the first widely adopted architectures for this task. Models like CharRNN, ReLeaSE, and conditional RNNs demonstrated success in generating focused molecular libraries. More recently, self-attention-based Transformer models (Mol-GPT, LigGPT) have gained popularity due to their parallelizability and ability to capture long-range dependencies.
Despite the widespread adoption of Transformers across NLP, it was not clear whether they uniformly outperform RNNs for molecular generation. Prior work by Dollar et al. showed that RNN-based models achieved higher validity than Transformer-based models in some settings. Flam-Shepherd et al. demonstrated that RNN language models could learn complex molecular distributions across challenging generative tasks. This paper extends that comparison by adding the Transformer architecture to the same set of challenging tasks and evaluating both SMILES and SELFIES representations.
Experimental Design: Three Tasks, Two Architectures, Two Representations
The core experimental design uses a 2x2 setup: two architectures (RNN and Transformer) crossed with two molecular representations (SMILES and SELFIES), yielding four model variants: SM-RNN, SF-RNN, SM-Transformer, and SF-Transformer.
Three generative tasks
The three tasks, drawn from Flam-Shepherd et al., are designed with increasing complexity:
Penalized LogP task: Generate molecules with high penalized LogP scores (LogP minus synthetic accessibility and long-cycle penalties). The dataset is built from ZINC15 molecules with penalized LogP > 4.0. Molecule sequences are relatively short (50-75 tokens).
Multidistribution task: Learn a multimodal molecular weight distribution constructed from four distinct subsets: GDB13 (MW <= 185), ZINC (185 <= MW <= 425), Harvard Clean Energy Project (460 <= MW <= 600), and POLYMERS (MW > 600). This tests the ability to capture multiple modes simultaneously.
Large-scale task: Generate large molecules from PubChem with more than 100 heavy atoms and MW ranging from 1250 to 5000. This tests long-sequence generation capability.
Model configuration
Models are compared with matched parameter counts (5.2-5.3M to 36.4M parameters). Hyperparameter optimization uses random search over learning rate [0.0001, 0.001], hidden units (500-1000 for RNNs, 376-776 for Transformers), layer number [3, 5], and dropout [0.0, 0.5]. A regex-based tokenizer replaces character-by-character tokenization, reducing token lengths from 10,000 to under 3,000 for large molecules.
Evaluation metrics
The evaluation covers multiple dimensions:
- Standard metrics: validity, uniqueness, novelty
- Molecular properties: FCD, LogP, SA, QED, Bertz complexity (BCT), natural product likeness (NP), molecular weight (MW)
- Wasserstein distance: measures distributional similarity between generated and training molecules for each property
- Tanimoto similarity: structural and scaffold similarity between generated and training molecules
- Token length (TL): comparison of generated vs. training sequence lengths
For each task, 10,000 molecules are generated and evaluated.
Key Results Across Tasks
Penalized LogP task
| Model | FCD | LogP | SA | QED | BCT | NP | MW | TL |
|---|---|---|---|---|---|---|---|---|
| SM-RNN | 0.56 | 0.12 | 0.02 | 0.01 | 16.61 | 0.09 | 5.90 | 0.43 |
| SF-RNN | 1.63 | 0.25 | 0.42 | 0.02 | 36.43 | 0.23 | 2.35 | 0.40 |
| SM-Transformer | 0.83 | 0.18 | 0.02 | 0.01 | 23.77 | 0.09 | 7.99 | 0.84 |
| SF-Transformer | 1.97 | 0.22 | 0.47 | 0.02 | 44.43 | 0.28 | 5.04 | 0.53 |
RNN-based models achieve smaller Wasserstein distances across most properties. The authors attribute this to LogP being computed as a sum of atomic contributions (a local property), which aligns with RNNs’ strength in capturing local structural features. RNNs also generated ring counts closer to the training distribution (4.10 for SM-RNN vs. 4.04 for SM-Transformer, with training data at 4.21). The Transformer performed better on global structural similarity (higher Tanimoto similarity to training data).
Multidistribution task
| Model | FCD | LogP | SA | QED | BCT | NP | MW | TL |
|---|---|---|---|---|---|---|---|---|
| SM-RNN | 0.16 | 0.07 | 0.03 | 0.01 | 18.34 | 0.02 | 7.07 | 0.81 |
| SF-RNN | 1.46 | 0.38 | 0.55 | 0.03 | 110.72 | 0.24 | 10.00 | 1.58 |
| SM-Transformer | 0.16 | 0.16 | 0.03 | 0.01 | 39.94 | 0.02 | 10.03 | 1.28 |
| SF-Transformer | 1.73 | 0.37 | 0.63 | 0.04 | 107.46 | 0.30 | 17.57 | 2.40 |
Both SMILES-based models captured all four modes of the MW distribution well. While RNNs had smaller overall Wasserstein distances, the Transformer fitted the higher-MW modes better. This aligns with the observation that longer molecular sequences (which correlate with higher MW) favor the Transformer’s global attention mechanism over the RNN’s sequential processing.
Large-scale task
| Model | FCD | LogP | SA | QED | BCT | NP | MW | TL |
|---|---|---|---|---|---|---|---|---|
| SM-RNN | 0.46 | 1.89 | 0.20 | 0.01 | 307.09 | 0.03 | 105.29 | 12.05 |
| SF-RNN | 1.65 | 1.78 | 0.43 | 0.01 | 456.98 | 0.14 | 100.79 | 15.26 |
| SM-Transformer | 0.36 | 1.64 | 0.07 | 0.01 | 172.93 | 0.02 | 59.04 | 7.41 |
| SF-Transformer | 1.91 | 2.82 | 0.47 | 0.01 | 464.75 | 0.18 | 92.91 | 11.57 |
The Transformer demonstrates a clear advantage on large molecules. SM-Transformer achieves substantially lower Wasserstein distances than SM-RNN across nearly all properties, with particularly large improvements in BCT (172.93 vs. 307.09) and MW (59.04 vs. 105.29). The Transformer also produces better Tanimoto similarity scores and more accurate token length distributions.
Standard metrics across all tasks
| Task | Metric | SM-RNN | SF-RNN | SM-Transformer | SF-Transformer |
|---|---|---|---|---|---|
| LogP | Valid | 0.90 | 1.00 | 0.89 | 1.00 |
| LogP | Uniqueness | 0.98 | 0.99 | 0.98 | 0.99 |
| LogP | Novelty | 0.75 | 0.71 | 0.71 | 0.71 |
| Multi | Valid | 0.95 | 1.00 | 0.97 | 1.00 |
| Multi | Uniqueness | 0.96 | 1.00 | 1.00 | 1.00 |
| Multi | Novelty | 0.91 | 0.98 | 0.91 | 0.98 |
| Large | Valid | 0.84 | 1.00 | 0.88 | 1.00 |
| Large | Uniqueness | 0.99 | 0.99 | 0.98 | 0.99 |
| Large | Novelty | 0.85 | 0.92 | 0.86 | 0.94 |
SELFIES achieves 100% validity across all tasks by construction, while SMILES validity drops for large molecules. The Transformer achieves slightly higher validity than the RNN for SMILES-based models, particularly on the large-scale task (0.88 vs. 0.84).
Conclusions and Practical Guidelines
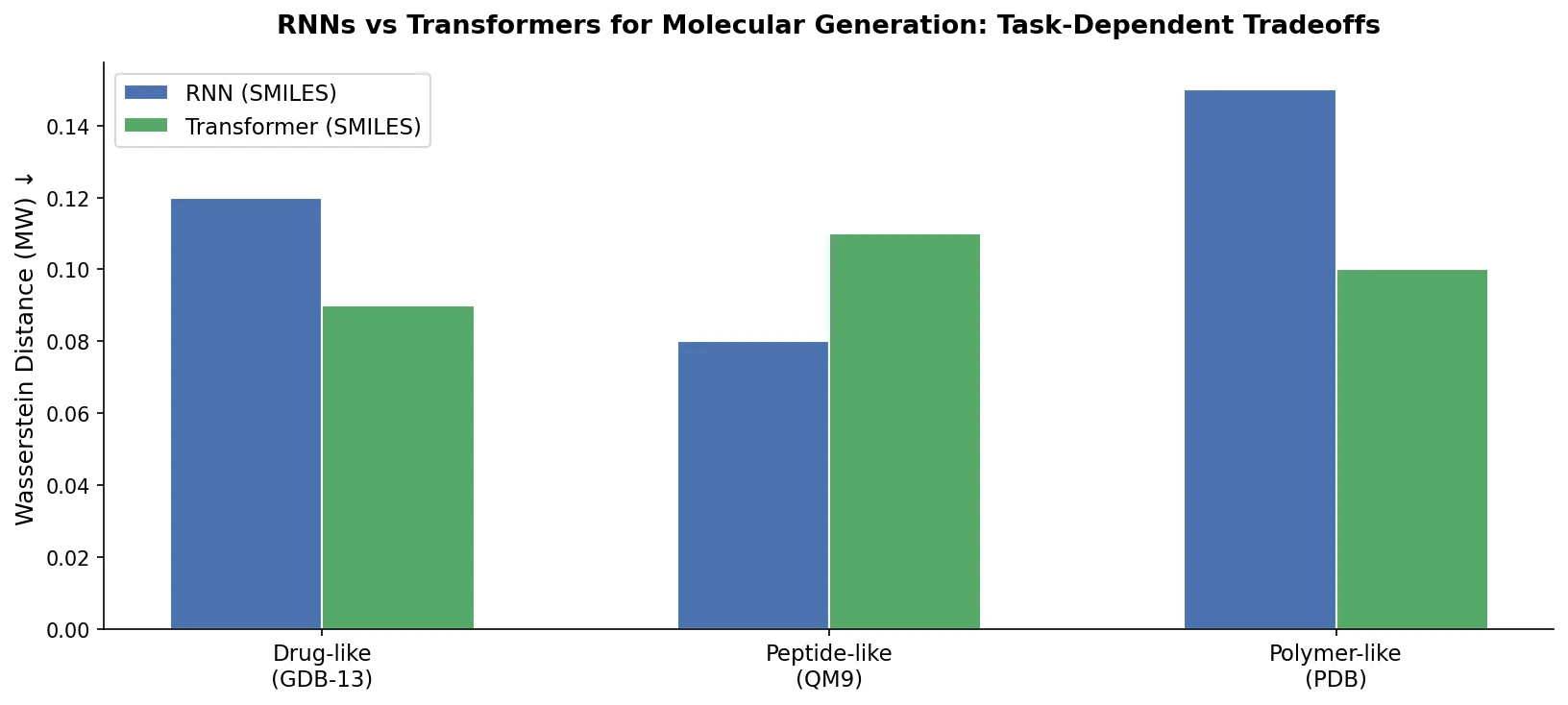
The central finding is that neither architecture universally dominates. The choice between RNNs and Transformers should depend on the characteristics of the molecular data:
RNNs are preferred when molecular properties depend on local structural features (e.g., LogP, ring counts) and when sequences are relatively short. They better capture local fragment distributions.
Transformers are preferred when dealing with large molecules (high MW, long sequences) where global attention can capture the overall distribution more effectively. RNNs suffer from information obliteration on long sequences.
SMILES outperforms SELFIES on property distribution metrics across nearly all tasks and models. While SELFIES guarantees 100% syntactic validity, its generated molecules show worse distributional fidelity to training data. The authors argue that validity is a less important concern than property fidelity, since invalid SMILES can be filtered easily.
The authors acknowledge that longer sequences remain challenging for both architectures. For Transformers, the quadratic growth of the attention matrix limits scalability. For RNNs, the vanishing gradient problem limits effective context length.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Task 1 | ZINC15 (penalized LogP > 4.0) | Not specified | High penalized LogP molecules |
| Task 2 | GDB-13 + ZINC + CEP + POLYMERS | ~200K | Multimodal MW distribution |
| Task 3 | PubChem (>100 heavy atoms) | Not specified | MW range 1250-5000 |
Data processing code available at https://github.com/danielflamshep/genmoltasks (from the original Flam-Shepherd et al. study).
Algorithms
- Tokenization: Regex-based tokenizer (not character-by-character)
- Hyperparameter search: Random search over learning rate [0.0001, 0.001], hidden units, layers [3, 5], dropout [0.0, 0.5]
- Selection: Top 20% by sum of valid + unique + novelty, then final selection on all indicators
- Generation: 10K molecules per model per task
Models
| Model | Parameters | Architecture |
|---|---|---|
| RNN variants | 5.2M - 36.4M | RNN (LSTM/GRU) |
| Transformer variants | 5.3M - 36.4M | Transformer decoder |
Evaluation
Wasserstein distance for property distributions (FCD, LogP, SA, QED, BCT, NP, MW, TL), Tanimoto similarity (molecular and scaffold), validity, uniqueness, novelty.
Hardware
Not specified in the paper.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| trans_language | Code | Not specified | Transformer implementation by the authors |
| genmoltasks | Code/Data | Apache-2.0 | Dataset construction from Flam-Shepherd et al. |
Paper Information
Citation: Chen, Y., Wang, Z., Zeng, X., Li, Y., Li, P., Ye, X., & Sakurai, T. (2023). Molecular language models: RNNs or transformer? Briefings in Functional Genomics, 22(4), 392-400. https://doi.org/10.1093/bfgp/elad012
@article{chen2023molecular,
title={Molecular language models: RNNs or transformer?},
author={Chen, Yangyang and Wang, Zixu and Zeng, Xiangxiang and Li, Yayang and Li, Pengyong and Ye, Xiucai and Sakurai, Tetsuya},
journal={Briefings in Functional Genomics},
volume={22},
number={4},
pages={392--400},
year={2023},
publisher={Oxford University Press},
doi={10.1093/bfgp/elad012}
}