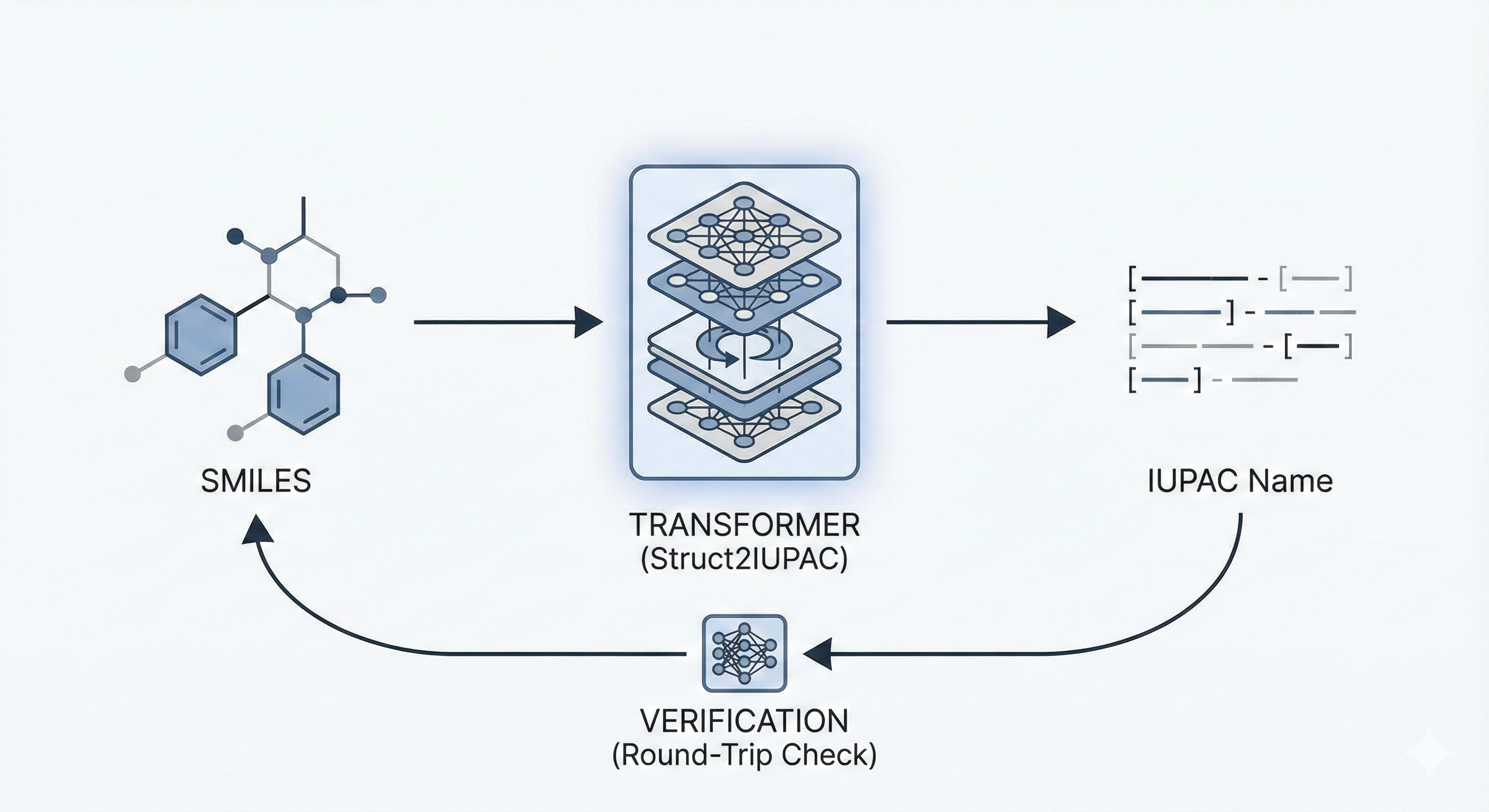
Struct2IUPAC as a Methodological Shift
This is primarily a Method paper with significant elements of Position.
- Method: The authors propose a specific neural architecture (Transformer with custom tokenization) and a verification pipeline (round-trip check) to solve the SMILES $\leftrightarrow$ IUPAC translation task. They rigorously benchmark this against rule-based baselines (OPSIN).
- Position: The authors explicitly argue for a paradigm shift, suggesting that “heavy” neural architectures should replace complex, costly rule-based legacy systems even for “exact” algorithmic tasks.
The Cost of Rule-Based Chemical Naming
- Complexity of Naming: Generating IUPAC names manually is error-prone and requires deep algorithmic knowledge.
- Lack of Open Source Tools: While open-source tools exist for Name-to-Structure (e.g., OPSIN), there were no open-source tools for the inverse “Structure-to-Name” conversion at the time of writing.
- Cost of Development: Developing rule-based converters “from scratch” is prohibitively expensive and time-consuming compared to training a neural model on existing data.
Struct2IUPAC Core Innovation
- Struct2IUPAC: The first effective open-source neural model for converting SMILES to IUPAC names, treating chemical translation as a Neural Machine Translation (NMT) problem.
- Verification Loop: A novel inference pipeline that generates multiple candidates via beam search and validates them using a reverse converter (OPSIN) to ensure the generated name maps back to the original structure.
- Custom Tokenization: A manually curated rule-based tokenizer for IUPAC names that handles specific chemical suffixes, prefixes, and stereochemical markers.
Experimental Setup and Stress Testing
- Accuracy Benchmarking: The models were tested on a held-out subset of 100,000 molecules from PubChem. The authors measured accuracy across different beam sizes (1, 3, 5).
- Comparison to Rules: The neural IUPAC2Struct model was compared directly against the rule-based OPSIN tool.
- Stress Testing:
- Sequence Length: Evaluated performance across varying token lengths, identifying a “sweet spot” (10-60 tokens) and failure modes for very short (e.g., methane) or long molecules.
- Stereochemistry: Tested on “stereo-dense” compounds. The authors define a “stereo-density” index ($I$) as the ratio of stereocenters ($S$) to total tokens ($N$): $$I = \frac{S}{N}$$ They observed a performance drop for these dense molecules, though the model still handled many stereocenters robustly.
- Tautomers: Verified the model’s ability to handle different tautomeric forms (e.g., Guanine and Uracil variants).
- Latency Analysis: Benchmarked inference speeds on CPU vs. GPU relative to output sequence length.
Benchmarks and Outcomes
- High Accuracy: The Struct2IUPAC model achieved 98.9% accuracy (Beam 5 with verification). The reverse model (IUPAC2Struct) achieved 99.1%, comparable to OPSIN’s 99.4%.
- Distribution Modeling vs. Intuition: The authors claim the model infers “chemical logic,” because it correctly generates multiple valid IUPAC names for single molecules where naming ambiguity exists (e.g., parent group selection). However, this more likely reflects the Transformer successfully modeling the high-frequency conditional probability distribution of synonymous names present in the PubChem training data, rather than learning intrinsic chemical rules.
- Production Readiness: Inference on GPU takes less than 0.5 seconds even for long names, making it viable for production use.
- Paradigm Shift: The authors conclude that neural networks are a viable, cost-effective alternative to developing rule-based algorithms for legacy notation conversion.
Reproducibility Details
Data
The study utilized the PubChem database.
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Total | PubChem | ~95M | Filtered for RDKit compatibility |
| Training | Split A | 47,312,235 | Random 50% split |
| Testing | Split B | 47,413,850 | Random 50% split |
- Cleaning: Molecules that could not be processed by RDKit were removed. Molecules containing tokens not in the tokenizer (e.g., aromatic selenium) were excluded.
- Availability: A subset of 100,000 test molecules is available on GitHub (
data/test_100000.csv) and Zenodo. The full train/test splits are not explicitly provided.
Algorithms
- Tokenization:
- SMILES: Character-based tokenization.
- IUPAC: Custom rule-based tokenizer splitting suffixes (
-one,-al), prefixes (-oxy,-di), and special symbols ((,),R(S)).
- Verification Step:
- Generate $N$ names using Beam Search ($N=5$).
- Reverse translate the candidate name using OPSIN.
- Check if the OPSIN structure matches the original input SMILES.
- Display the first verified match; otherwise, report failure.
Models
- Architecture: Standard Transformer with 6 encoder layers and 6 decoder layers.
- Hyperparameters:
- Attention Heads: 8
- Attention Dimension ($d_{\text{model}}$): 512
- Feed-Forward Dimension ($d_{\text{ff}}$): 2048
- Training Objective: The models were trained using standard autoregressive cross-entropy loss over the target token sequence $y$ given the input string $x$: $$\mathcal{L} = - \sum_{t=1}^{T} \log P(y_t \mid y_{<t}, x)$$
- Training: Two separate models were trained:
Struct2IUPAC(SMILES $\to$ IUPAC) andIUPAC2Struct(IUPAC $\to$ SMILES). - Availability: Code for model architecture is provided in the GitHub repository. Pre-trained weights for the IUPAC2Struct model are available, but the Struct2IUPAC model weights are not publicly released, meaning researchers would need to retrain that model on their own PubChem data to reproduce those results.
Evaluation
Evaluation was performed on a random subset of 100,000 molecules from the test set.
| Metric | Task | Beam Size | Accuracy |
|---|---|---|---|
| Exact Match | Struct2IUPAC | 1 | 96.1% |
| Exact Match | Struct2IUPAC | 5 | 98.9% |
| Exact Match | IUPAC2Struct | 1 | 96.6% |
| Exact Match | IUPAC2Struct | 5 | 99.1% |
- Robustness: Accuracy drops significantly for augmented (non-canonical) SMILES (37.16%) and stereo-enriched compounds (66.52%).
Hardware
- Training Infrastructure: 4 $\times$ Tesla V100 GPUs and 36 CPUs.
- Training Time: Approximately 10 days under full load.
- Inference Speed: <0.5s per molecule on GPU; scale is linear with output token length.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| IUPAC2Struct (GitHub) | Code | MIT | Transformer code and pre-trained IUPAC2Struct model |
| Test data (Zenodo) | Dataset | Unknown | 100k test molecules, OPSIN failure cases, model failure cases |
| Struct2IUPAC web demo | Other | N/A | Online interface for SMILES to IUPAC conversion |
Paper Information
Citation: Krasnov, L., Khokhlov, I., Fedorov, M. V., & Sosnin, S. (2021). Transformer-based artificial neural networks for the conversion between chemical notations. Scientific Reports, 11(1), 14798. https://doi.org/10.1038/s41598-021-94082-y
Publication: Scientific Reports 2021
@article{krasnovTransformerbasedArtificialNeural2021a,
title = {Transformer-Based Artificial Neural Networks for the Conversion between Chemical Notations},
author = {Krasnov, Lev and Khokhlov, Ivan and Fedorov, Maxim V. and Sosnin, Sergey},
year = 2021,
month = jul,
journal = {Scientific Reports},
volume = {11},
number = {1},
pages = {14798},
publisher = {Nature Publishing Group},
doi = {10.1038/s41598-021-94082-y}
}
Additional Resources: