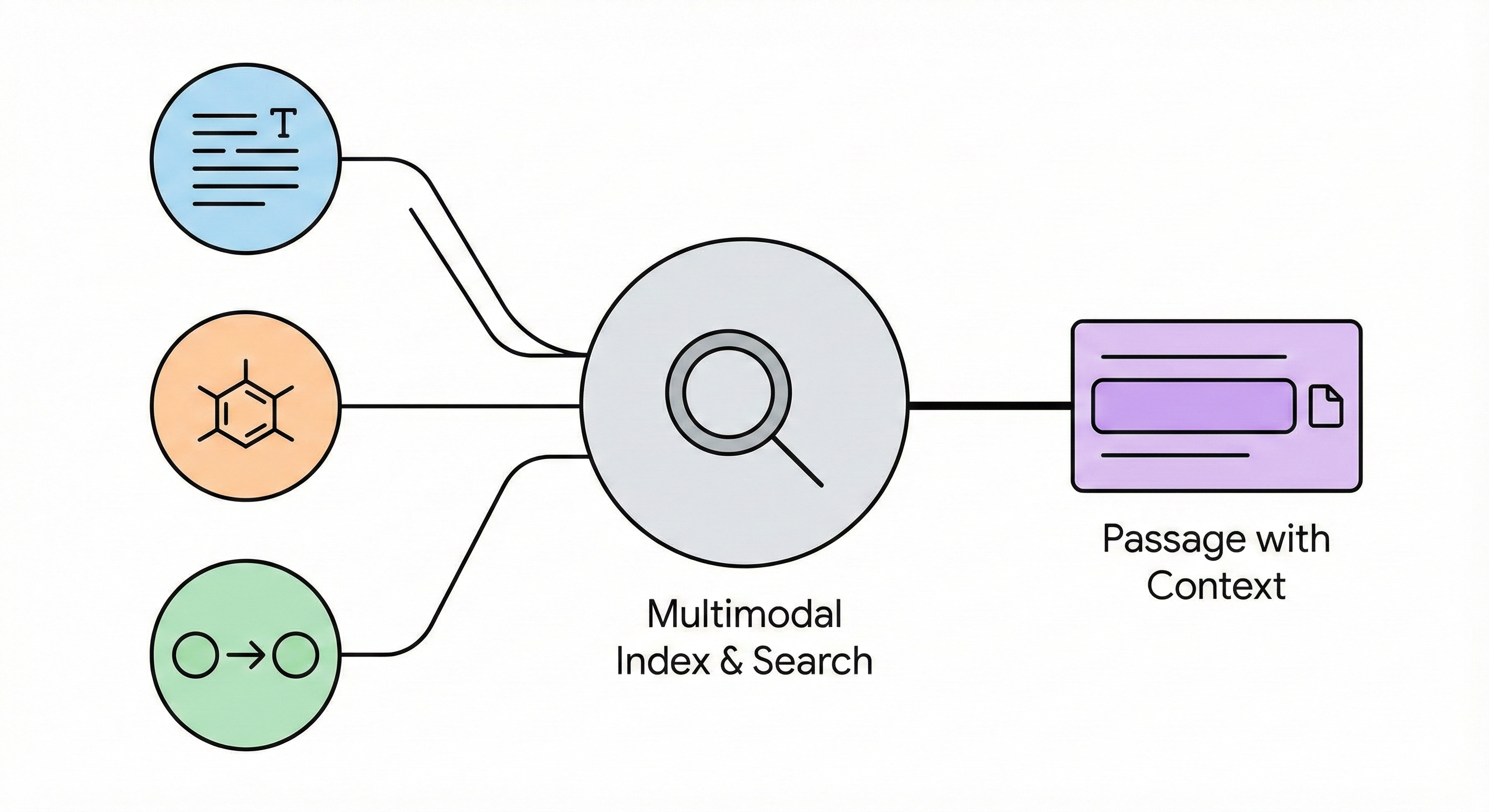
Contribution: Multimodal Synthesis Retrieval
This paper represents a $\Psi_{\text{Method}}$ projection that proposes a novel architectural pipeline for indexing and searching chemical literature. The framework unifies text, molecular diagrams, and structured reaction records. It also contains a secondary $\Psi_{\text{Resource}}$ projection, providing a functional demonstration tool and curating a specific benchmark dataset for Suzuki coupling reactions.
The Gap in Passage-Level Chemical Retrieval
Scientific literature documents chemical reactions through a combination of text and visual diagrams. Textual descriptions detail parameters like yield and operational temperature, whereas diagrams graphically model these structural transformations. Existing tools such as SciFinder or Reaxys perform document-level or individual compound retrieval. They fail to explicitly link molecular figures to localized textual descriptions. This structure prevents researchers from directly extracting a corresponding reaction diagram alongside the exact textual protocol. Researchers require passage-level retrieval of synthesis protocols to efficiently access complete reaction conditions.
Core Innovation: Unified Multimodal Indexing
The core methodological innovation is a multimodal passage-level indexing and linking pipeline.
- Unified Indexing: The framework processes text and diagrams in parallel and directly links them into a single index structure. This architecture supports search queries utilizing raw text, discrete SMILES strings, or multimodal combinations.
- Compound-Passage Linking: The mechanism applies conflict-resolution logic linking chemical diagrams to specific text citations using two parallel heuristics:
- Token-based Alignment: Matching parsed diagram labels against documented text strings (e.g., “compound 5”) using normalized Levenshtein distance.
- Fingerprint-based Alignment: Matching chemical structures against generated SMILES strings via structural Tanimoto Similarity.
- ReactionMiner Integration: The pipeline parses and incorporates formatted reaction records (reactants, products, catalysts, quantitative yields) directly derived from segmented text passages.
Methodology & Expert Evaluation
The authors evaluated the system utilizing a chemical case study targeting specific synthesis domains alongside qualitative expert assessment.
- Dataset: Evaluators processed a corpus of 7 research manuscripts and 6 supplementary data documents detailing Suzuki coupling reactions.
- Volume: The resulting index processed 1,282 extracted passages (indexing 538), extracted 383 unique SMILES, and logged 219 parsed reactions.
- Qualitative Evaluation: Practicing structural chemists developed real-world queries (such as cross-referencing the conceptual “Burke group” alongside an explicit structural SMARTS pattern) to gauge retrieval capability.
Key Findings & System Limitations
- Diagram-to-Text Linking: The pipeline accurately paired visual molecular diagrams with structurally derived text details, permitting testers to navigate directly from a molecule query card to the exact origin passage within the source PDF.
- Contextual Insight Extraction: Specialized chemists found the parsed reaction representations (yield metrics, isolated catalysts) functionally pragmatic as high-level extractive summaries.
- Extrapolative Retrieval: The architecture permitted the effective retrieval of targeted chemical derivatives (such as benzo[b]thiophen-2-ylboronic acid) via structurally related input queries (dibenzothiophene).
The system evaluation highlights several architectural restrictions:
- Domain-Restricted Validation: The initial validation is entirely qualitative and bounded to the specific subclass of Suzuki coupling reactions. The evaluation omits standardized quantitative retrieval baselines (e.g., MAP, NDCG) and lacks systematic ablation data for the fusion scoring mechanism.
- Algorithmic Transparency: The multimodal query routing mechanism does not clearly indicate the dominant retrieval feature. This hides whether keyword text or structural similarity actually drove the final result placement. This ambiguity limits operator control.
- Optical Processing Brittleness: The embedded vision inference and primitive parsing pipelines display inherent fragility, producing intermittent failures when associating text passages with correctly parsed molecular diagrams.
- Metadata Logging Incompleteness: Practicing chemists requested additional structured metadata targets (such as specific molar equivalents and parameterized mol% values) to successfully bridge the extracted data stream directly into digital electronic lab notebooks.
Reproducibility
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| ReactionMiner Demo | Other | Unknown | Online demo landing page; source code repository not publicly linked |
Data
- Source: The corpus features 7 primary research papers and 6 auxiliary supplementary information documents focusing on Suzuki coupling reactions, sourced from practicing chemists at UIUC. This evaluation dataset is strictly internal and not publicly available.
- Preprocessing:
- Engineers convert source PDFs to full-page raster images.
- The system extracts localized graphical layout and raw text via PyTesseract.
- The pipeline segments valid passage chunks emphasizing reaction-related sentences utilizing product-indicative lexicons and topic modeling.
Algorithms
- Diagram Extraction: A YOLOv8 model identifies and segments molecular regions within structured PDF pages.
- Diagram Parsing: The architecture relies on ChemScraper to infer structural semantics from raw diagrams:
- Born-digital PDFs: SymbolScraper extracts vector lines and polygons directly from bounding box definitions.
- Raster images: The system employs the Line Segment Detector (LSD) and watershed bounding algorithms to isolate native geometric primitives.
- Text Entity Extraction: The framework deploys ChemDataExtractor 2.0 to extract explicit molecular aliases. A translation layer maps these entities to string representations via OPSIN.
- Linking Logic (Fusion Score):
- Text Link: The algorithm calculates a normalized Levenshtein ratio connecting visual diagram labels against proximal text mentions based on calculated edit distance.
- Structure Link: The algorithm computes the discrete Tanimoto Similarity between generated 2048-bit Morgan fingerprints extracted from localized visual diagram features and baseline text SMILES queries: $$ T(A, B) = \frac{A \cdot B}{|A|^{2} + |B|^{2} - A \cdot B} $$ where $A$ and $B$ represent the boolean bit vectors of the respective fingerprint pairs.
- Conflict Resolution Protocol: The system fuses structural geometry bounds and discrete textual tokenization metrics, prioritizing the ranking sequence that yields a higher terminal similarity score. During final retrieval, the candidate subset is systematically re-ranked leveraging the hybrid calculation of the BM25 explicit metric and the localized count of exact SMILES pattern hits.
Models
- Reaction Extraction Parameters: The engineers configure a LLaMA-3.1-8b model fine-tuned entirely via LoRA targeting custom tokens representing reaction entities (compounds, reagents, thermal inputs) directly pulled from text sub-chunks. Exact prompt constraints, the fine-tuning dataset, and specific LoRA hyperparameters are omitted from the source text.
- Diagram Processing Bounds: The codebase incorporates a segmentation-aware multi-task neural network topology built into ChemScraper to execute low-level raster image parsing tasks.
Evaluation
- Search Engine Base: The authors implemented their indexing framework scaling atop PyTerrier.
- Text Feature Ranking: The metric utilizes standalone BM25 bounds mapping keyword-similarity.
- Structure Feature Operations: The topology operates RDKit bindings powering substructure coordinate mapping logic and exact molecular similarity searches.
- Multimodal Fusion Processing:
- The algorithm filters out terminal candidates mapping initial structural properties (SMILES queries) against the document-wide lexical properties (BM25 scores).
- The final fusion routing assigns the strongest positive weight to retrieved passages that accumulate dense local clusters of structurally exact verified SMILES patterns.
Hardware
- Compute Infrastructure: The hardware and parameter requirements to host the multi-stage vision extractors (YOLOv8, ChemScraper) alongside a local 8B LLM are entirely unspecified in the paper.
Paper Information
Citation: Shah, A. K., et al. (2025). Multimodal Search in Chemical Documents and Reactions. arXiv preprint arXiv:2502.16865. https://doi.org/10.48550/arXiv.2502.16865
Publication: SIGIR ‘25 (Demo Track), 2025
@misc{shahMultimodalSearchChemical2025,
title = {Multimodal {{Search}} in {{Chemical Documents}} and {{Reactions}}},
author = {Shah, Ayush Kumar and Dey, Abhisek and Luo, Leo and Amador, Bryan and Philippy, Patrick and Zhong, Ming and Ouyang, Siru and Friday, David Mark and Bianchi, David and Jackson, Nick and Zanibbi, Richard and Han, Jiawei},
year = 2025,
month = feb,
number = {arXiv:2502.16865},
eprint = {2502.16865},
primaryclass = {cs},
publisher = {arXiv},
doi = {10.48550/arXiv.2502.16865},
archiveprefix = {arXiv}
}
Additional Resources:
- Online Demo (Note: While the landing page advertises the system as open-source, the exact repository URL and installation prerequisites are omitted from the official manuscript.)