Paper Information
Citation: Born, J. & Manica, M. (2023). Regression Transformer enables concurrent sequence regression and generation for molecular language modelling. Nature Machine Intelligence, 5(4), 432-444. https://doi.org/10.1038/s42256-023-00639-z
Publication: Nature Machine Intelligence, April 2023
Additional Resources:
A Multitask Model That Unifies Regression and Generation
The Regression Transformer (RT) is a Method paper. It introduces a single model architecture that can both predict continuous molecular properties and conditionally generate molecules with desired property values. The core idea is to reformulate regression as a sequence modelling task: instead of training a dedicated regression head, continuous property values are tokenized into sequences of digits and predicted alongside molecular tokens using a cross-entropy loss.
Closing the Gap Between Predictors and Generators
Existing transformer-based approaches in computational chemistry develop property predictors and generative models as separate systems. Even when a single architecture like Chemformer (Irwin et al., 2022) addresses both tasks, it does so through task-specific heads. This means the two capabilities remain disjoint, and the generative model cannot use its own property prediction ability during generation.
The RT addresses three specific gaps:
- No true multitask entanglement: Prior work either tunes separate heads for prediction and generation or limits communication between modules to a reward signal.
- No inductive bias for continuous properties: Molecular generative models lack mechanisms to condition generation on floating-point property values.
- Disconnected workflows: Property predictors cannot generate molecules, and generators cannot assess whether their outputs satisfy property constraints.
Core Innovation: Regression as Conditional Sequence Modelling
The RT’s key insight is that regression can be cast as sequential classification over digit tokens while preserving predictive accuracy. This is achieved through three components:
Numerical Tokenization
Floating-point property values are split into individual digit tokens that preserve decimal order. Each token $t_{v,p}$ encodes a digit value $v \in [0, 9]$ and its decimal place $p \in \mathbb{Z}$. For example, the value 12.3 becomes the token sequence [1_1, 2_0, 3_-1].
Numerical Encodings
To provide an inductive bias about the semantic proximity of digit tokens (which cross-entropy loss cannot convey), the RT introduces Numerical Encodings (NEs), analogous to positional encodings. For a token $t_{v,p}$ at embedding dimension $j$:
$$ \text{NE}_{\text{Float}}(v, p, j) = (-1)^j \cdot \frac{v \cdot 10^p}{j + 1} $$
These encodings ensure that pairwise distances between digit tokens decay monotonically with their floating-point proximity. The model can also learn digit orderings from data alone, but NEs provide a useful inductive bias.
Alternating Training with Self-Consistency
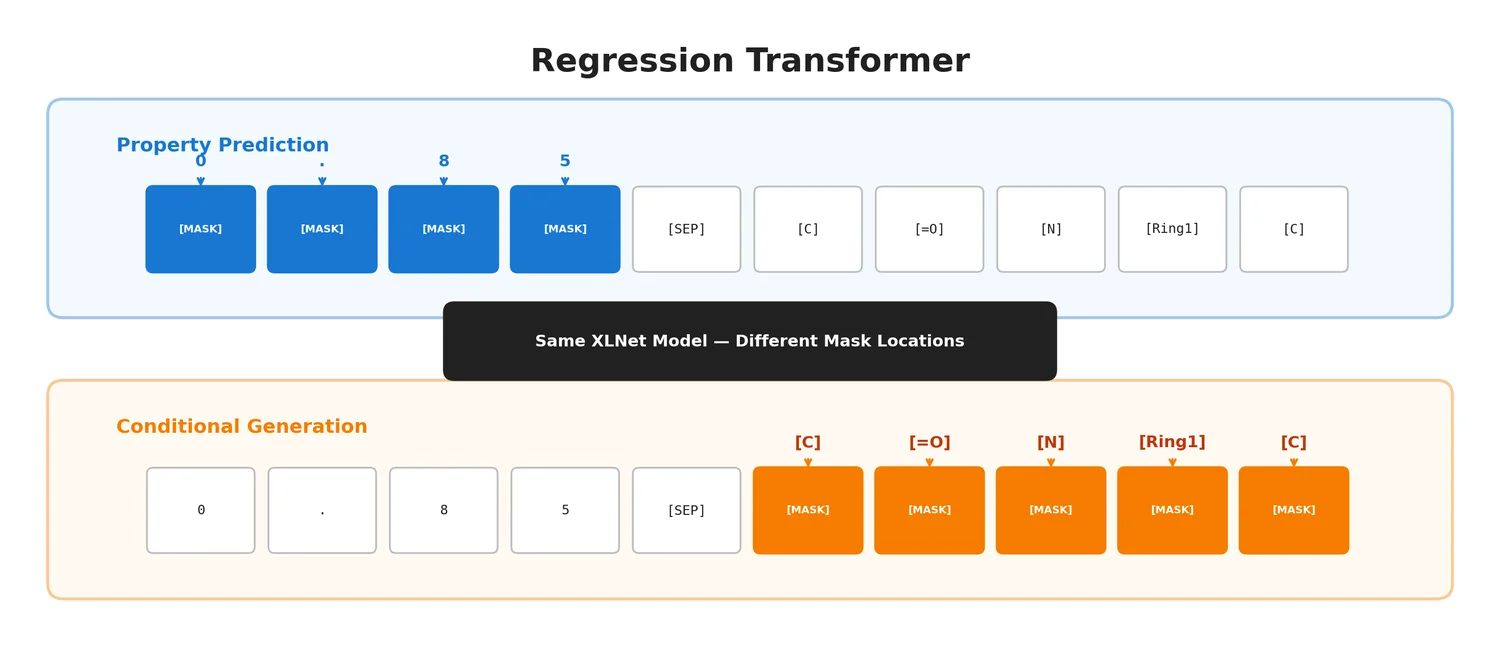
The RT uses an XLNet backbone trained with permutation language modelling (PLM). The key is that the same model serves two roles depending on which tokens are masked:
- Mask numerical tokens: the model performs property prediction (regression)
- Mask textual tokens: the model performs conditional sequence generation
The base PLM objective is:
$$ \mathcal{L}_{\text{PLM}} = \mathbb{E}_{\mathbf{z} \sim \mathcal{Z}_T} \left[ \sum_{i=c+1}^{T} \log p_\theta(x_{z_i} \mid \mathbf{x}_{\mathbf{z}_{< i}}) \right] $$
This is refined into two specialized objectives: a property prediction objective $\mathcal{L}_P$ that masks only numerical tokens, and a generation objective $\mathcal{L}_G$ that masks only textual tokens. Training alternates between these every 50 steps.
The self-consistency (SC) loss adds a critical feedback loop. After generating a candidate molecule $\hat{\mathbf{x}}$, the model re-evaluates it by predicting the property of the generated sequence:
$$ \mathcal{L}_{\text{SC}} = \mathcal{L}_G(\mathbf{x}) + \alpha \cdot \mathcal{L}_P(\hat{\mathbf{x}}) $$
This rewards generating molecules whose predicted properties match the primed property value, exploiting the RT’s dual capability as both predictor and generator.
Experiments Across Molecules, Proteins, and Reactions
Drug Likeness (QED)
Initial validation on a synthetic QED dataset (~1.4M molecules from ChEMBL) demonstrated that the RT can simultaneously learn to predict QED scores (RMSE < 0.06) and generate novel molecules conditioned on desired QED values (Spearman’s $\rho$ up to 0.517 between primers and generated molecule properties). Novelty exceeded 99% across all configurations. The alternating training scheme with SC loss outperformed both single-task models and the vanilla PLM objective.
SELFIES representations proved comparable to SMILES for property prediction and far superior for generation (~100% validity vs. ~40% for SMILES).
MoleculeNet Regression Benchmarks
On ESOL, FreeSolv, and Lipophilicity, the RT outperformed XGBoost and MPNN baselines despite using only a classification loss. It performed on par with XLNet using a conventional regression head, and was only mildly inferior to models like BERT and BART that used large-scale self-supervised pre-training with regression losses.
Critically, only the RT could also conditionally generate molecules for these tasks. External validation with Grover (a self-supervised Graph Transformer) confirmed high correlation with the RT’s own property predictions (0.86, 0.84, and 0.75 for ESOL, FreeSolv, and Lipophilicity respectively).
Constrained Property Optimization
On the penalized logP (plogP) benchmark with similarity constraints, the RT outperformed JT-VAE and GCPN by large margins. At similarity threshold $\delta = 0.4$, the RT achieved 3.16 average improvement with 97.1% success rate, while also predicting plogP with PCC of 0.92. Competing methods cannot perform property prediction at all.
| Model | Improvement ($\delta$=0.4) | Success | Property Prediction |
|---|---|---|---|
| JT-VAE | 0.84 | 83.6% | Unfeasible |
| GCPN | 2.49 | 100% | Unfeasible |
| MoFlow | 4.71 | 85.7% | Unfeasible |
| RT | 3.16 | 97.1% | PCC = 0.92 |
The comparison is not strictly fair: all competing methods are trained specifically to maximize plogP, and some (GCPN, JT-VAE) apply gradient optimization at inference time. The RT is only trained to reconstruct molecules with similar predicted plogP to the seed, so its training objective is property-agnostic rather than directly optimizing for higher plogP values.
Protein Language Modelling
On the TAPE benchmark, the RT matched or outperformed conventional transformers on fluorescence and stability prediction tasks, despite those baselines being pre-trained on 24-106 million protein sequences (vs. 2.6 million for the RT). The RT also performed conditional protein generation, a task that none of the TAPE baselines can address.
Chemical Reaction Modelling
The RT was applied to reaction yield prediction on Buchwald-Hartwig amination and Suzuki coupling datasets. It matched Yield-BERT performance ($R^2$ = 0.939 and 0.81 respectively) while also enabling novel capabilities: reconstructing missing precursors from partial reactions and decorating existing reactions to achieve higher predicted yields. Across both datasets, over 40% of top-five predicted sequences contained reactions with novel precursors and higher predicted yield.
Key Findings and Limitations
Key Findings
- Regression can be successfully reformulated as sequential classification over digit tokens without losing predictive accuracy compared to models using regression losses.
- The alternating training scheme with self-consistency loss enables cross-task benefits, where the model outperforms single-task variants at both prediction and generation.
- A single ~27M parameter model handles property prediction, conditional molecular generation, conditional protein generation, and reaction yield prediction with precursor generation.
- The model learns the natural ordering of digits from data: 47% of embedding dimensions for the tenths place directly encode digit ordering even without explicit numerical encodings.
Limitations
- No large-scale pre-training: The RT uses ~27M parameters trained from scratch on task-specific datasets, unlike BARTSmiles or MoLFormer which pre-train on billions of molecules. Scaling up could improve results.
- Fine-grained regression precision: The model sometimes struggles with intra-mode precision (e.g., on the fluorescence dataset where predictions cluster around bright/dark modes rather than capturing continuous variation).
- Single-property focus: All reported experiments use a single continuous property, though the framework naturally extends to multi-property settings.
- SELFIES validity caveats: While SELFIES are always syntactically valid, they can produce degenerate short molecules (~1.9% defective generations where the output has less than 50% of the seed’s atoms).
- XLNet backbone limitations: Results on MoleculeNet regression are slightly below models using BART or BERT backbones with large-scale pre-training, suggesting the RT framework could benefit from stronger base models.
Reproducibility Details
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| Regression Transformer (GitHub) | Code | MIT | Training and evaluation scripts |
| GT4SD Integration | Code + Models | MIT | Pre-trained model inference pipelines |
| HuggingFace Demo | Demo | - | Interactive inference webapp |
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Drug likeness | ChEMBL (QED) | ~1.4M molecules | Synthetic QED labels computed with RDKit |
| Regression benchmark | MoleculeNet (ESOL, FreeSolv, Lipo) | 642-4,200 compounds | 16x SMILES augmentation, 3 random splits |
| Property optimization | ZINC (plogP) | 215,381 train / 799 test | Fixed split from Jin et al. (2018) |
| Protein pre-training | UniProt (Boman) | 2,648,205 peptides | 15-45 amino acid peptides |
| Protein benchmarks | TAPE (Fluorescence, Stability) | 21,446-53,416 samples | Fixed splits |
| Reaction pre-training | USPTO | 2,830,616 reactions | Molecular weight as numerical property |
| Reaction yield | Buchwald-Hartwig / Suzuki | 3,955 / 5,760 reactions | Ten 70/30 random splits |
Algorithms
- Architecture: XLNet (32 hidden layers, 256 hidden dim, 1024 FFN dim, 16 attention heads, 20% dropout)
- Parameters: ~27 million
- Training: Permutation language modelling pre-training, then alternating objectives (property prediction + conditional generation with SC loss)
- Decoding: Greedy for property prediction, beam search for sequence generation
Evaluation
| Task | Metric | RT Result | Notes |
|---|---|---|---|
| QED prediction | RMSE | 0.037 | Best config (NE + SC) |
| QED generation | Spearman’s $\rho$ | 0.517 | Between primers and generated QED |
| ESOL | RMSE | Comparable to XLNet | Within s.d. of regression-loss XLNet |
| plogP optimization ($\delta$=0.4) | Improvement | 3.16 | Outperforms JT-VAE, GCPN |
| Protein fluorescence | Spearman’s $\rho$ | 0.72 | Outperforms TAPE baselines |
| BH yield prediction | $R^2$ | 0.939 | Near Yield-BERT (0.951) |
Hardware
- All models trained on single GPUs (NVIDIA A100 or V100)
- Training time: ~4 days for pre-training, ~1 day for fine-tuning
- Framework: PyTorch 1.3.1 with HuggingFace Transformers 3.1.0
Citation
@article{born2023regression,
title={Regression Transformer enables concurrent sequence regression and generation for molecular language modelling},
author={Born, Jannis and Manica, Matteo},
journal={Nature Machine Intelligence},
volume={5},
number={4},
pages={432--444},
year={2023},
publisher={Nature Publishing Group}
}