Pioneering Seq2Seq Translation for Reaction Prediction
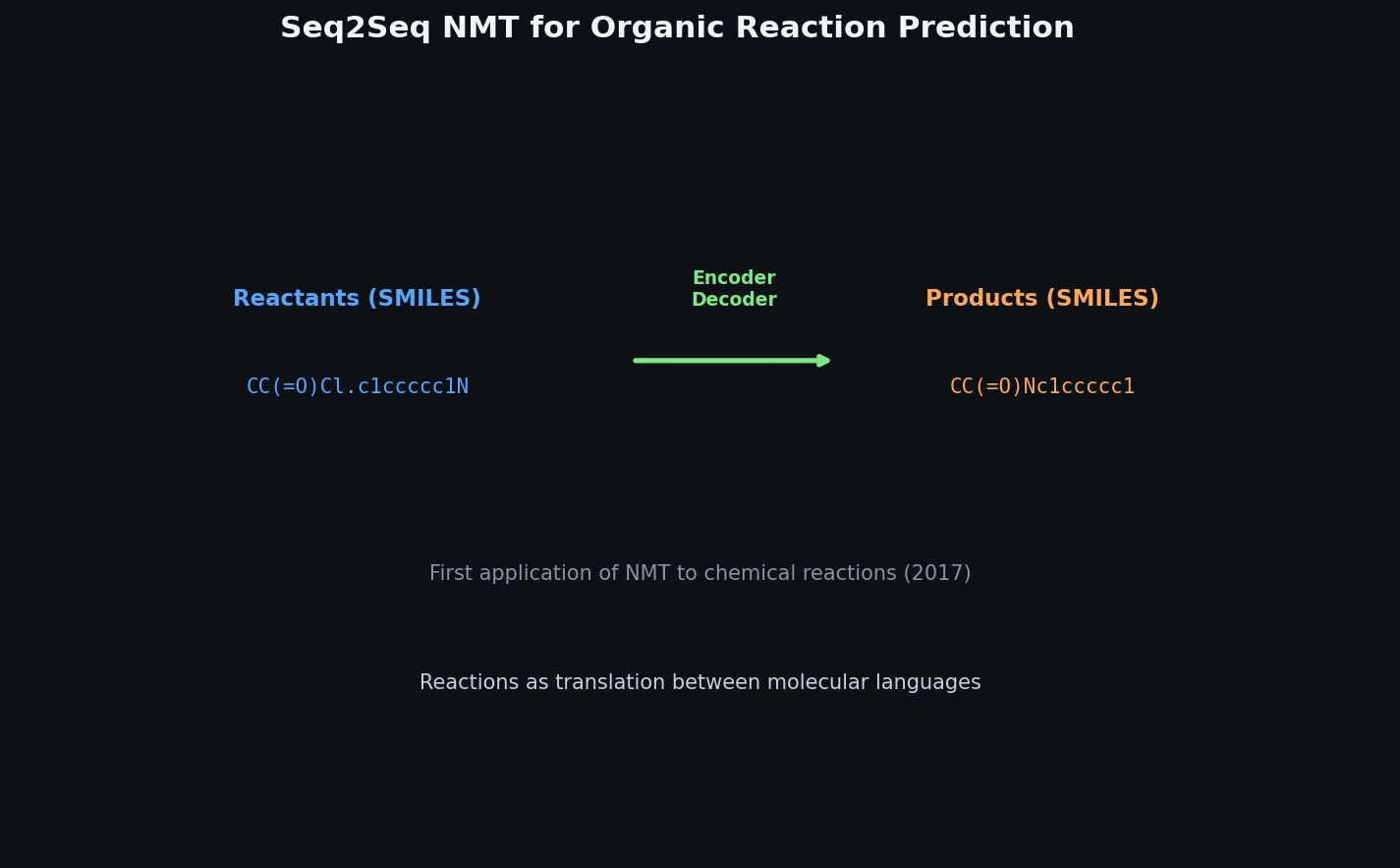
This is a Method paper. It introduces the idea of applying neural machine translation (NMT) to organic chemistry reaction prediction by framing product prediction as a sequence-to-sequence translation problem from reactant/reagent SMILES to product SMILES. This was one of the earliest works to demonstrate that a data-driven encoder-decoder model could predict reaction products without any hand-coded reaction rules or SMARTS transformations.
Limitations of Existing Reaction Prediction Methods
Prior computational approaches to reaction prediction fell into three categories, each with significant drawbacks:
Rule-based methods (e.g., CAMEO, EROS) relied on manually encoded reaction rules. They performed well on reactions covered by the rules but required continuous manual encoding as new reaction types were discovered. Many older systems became outdated for this reason.
Physical calculation methods computed energies of transition states from plausible reaction pathways using quantum mechanics. While principled, these approaches carried high computational cost. Simplified approaches (ToyChem, ROBIA) traded accuracy for speed.
Machine learning methods at the time either predicted individual mechanistic steps (requiring tree search for multi-step reactions) or classified reaction types and applied SMARTS transformations to generate products. The classification-based approach of Wei et al. still required manual encoding of SMARTS transformations for new reaction types and struggled with ambiguous reaction classes.
The key gap was the absence of a method that could predict reaction products directly from input molecules, learn from data alone, and generalize to new reaction types without manual rule encoding.
Core Innovation: Reactions as Machine Translation
The central insight is that SMILES strings can be treated as a language with grammatical specifications. Predicting reaction products then becomes a problem of translating “reactant and reagent” sentences into “product” sentences.
The model uses a GRU-based encoder-decoder architecture with attention:
- Encoder: 3 layers of GRU cells that process the reversed, tokenized SMILES string of reactants and reagents
- Decoder: 3 layers of GRU cells that generate product SMILES tokens autoregressively
- Attention mechanism: allows the decoder to attend to relevant encoder states at each generation step
- Embedding dimension: 600
- Vocabulary: 311 input tokens (reactants/reagents), 180 output tokens (products)
- Bucketed sequences: four bucket sizes handle variable-length inputs and outputs: (54, 54), (70, 60), (90, 65), (150, 80)
The SMILES tokenization uses a PEG-based parser that splits SMILES strings into atoms, bonds, branching symbols, and ring closure numbers. Input sequences are reversed before feeding to the encoder, following standard practice in NMT at the time.
The translation objective finds the product sequence $\mathbf{y}$ that maximizes the conditional probability:
$$p(\mathbf{y} \mid \mathbf{x}) = \prod_{t=1}^{T} p(y_t \mid y_1, \ldots, y_{t-1}, \mathbf{x})$$
where $\mathbf{x}$ is the tokenized reactant/reagent sequence and $T$ is the product sequence length.
Training Data and Experimental Evaluation
Training Sets
Two training sets were constructed:
| Source | Size | Description |
|---|---|---|
| Patent reactions (“real”) | 1,094,235 | USPTO patent applications (2001-2013), filtered by length |
| Generated reactions (“gen”) | 865,118 | 75 reaction types from Wade’s organic chemistry textbook, applied to GDB-11 molecules (1-10 atoms) |
The “real” set was filtered to exclude reactions with reactant/reagent strings longer than 150 characters, product strings longer than 80 characters, or more than four products. The “gen” set was composed by iterating reaction templates (as SMARTS) over small molecules from GDB-11, covering five substrate types: acid derivatives, alcohols, aldehydes/ketones, alkenes, and alkynes.
Two models were compared: a “gen” model (trained only on generated reactions) and a “real+gen” model (trained on both sets).
Textbook Problem Evaluation
The models were tested on 10 problem sets from Wade’s textbook, following the evaluation approach of Wei et al. Each problem set contained 6-15 reactions. Evaluation metrics included the ratio of fully correct predictions and the average Tanimoto similarity between Morgan fingerprints of predicted and actual products.
The “real+gen” model outperformed the “gen” model on most problem sets. On problem set 17-44 (aromatic compound reactions, only present in the “real” training set), the “real+gen” model correctly answered 4 out of 11 problems while the “gen” model answered 2. The “gen” model’s ability to correctly predict some aromatic reactions despite never being trained on them suggests the model can extrapolate to unseen reaction patterns.
For Diels-Alder reactions (problem set 15-30), neither model achieved fully correct predictions for all problems, though the “real+gen” model showed better Tanimoto scores, indicating partially correct structural predictions even when the exact product was missed.
Scalability Testing
A scalability test used generated reactions with substrate molecules containing 11-16 atoms (larger than the training set molecules with fewer than 11 atoms). Results showed:
- The “real+gen” model maintained Tanimoto scores around 0.7 and error rates around 0.4 as substrate atom count increased
- The ratio of fully correct predictions decreased as atom count increased, revealing that the recurrent network struggled with longer input sequences
- The “real+gen” model produced fewer invalid SMILES strings than the “gen” model, likely because training on more reactions improved the decoder’s ability to generate syntactically valid SMILES
Attention Analysis
Visualization of attention weights revealed a limitation: the decoder cells predominantly attended to the first few encoder cells rather than distributing attention across the full input sequence. This means the attention mechanism was not learning meaningful “alignment” between reactant atoms and product atoms. The authors note that if decoder cells generating tokens for unreactive sites could attend to the corresponding encoder cells (analogous to atom mapping), prediction quality on longer sequences could improve.
Token Embedding Analysis
t-SNE visualization of the learned token embeddings showed that encoder and decoder tokens clustered primarily by syntactic similarity rather than chemical properties. The model did not learn chemically meaningful embeddings, which the authors identify as an area for future improvement.
Key Findings, Limitations, and Impact
Key Findings
- Treating reaction prediction as NMT is viable: the seq2seq model can predict products without any hand-coded rules
- Training on real patent data significantly improves prediction over generated data alone
- The model can extrapolate to reaction types not seen during training (e.g., the “gen” model predicting aromatic reactions)
- Compared to the fingerprint-based approach of Wei et al., this method performed better on textbook problems and eliminated the need for manual SMARTS encoding
Limitations
- Invalid SMILES generation: the token-by-token generation process can produce syntactically invalid SMILES (e.g., mismatched parentheses), which the authors scored as zero
- Sequence length degradation: prediction accuracy dropped for longer SMILES strings, a known limitation of RNN-based seq2seq models at the time
- Poor attention alignment: attention weights collapsed to the first encoder positions rather than learning meaningful reactant-product correspondences
- Chemically naive embeddings: token embeddings did not capture chemical properties
- Multiple reaction pathways: reactions with competing pathways (e.g., substitution vs. elimination) were difficult for the model to handle
Historical Significance
This paper is historically significant as one of the first (alongside concurrent work) to propose the NMT framing for reaction prediction. This framing was later adopted and refined by the Molecular Transformer (Schwaller et al., 2019), which replaced GRUs with the Transformer architecture and achieved over 90% top-1 accuracy on standard benchmarks. The conceptual contribution of treating SMILES-to-SMILES translation as machine translation became the foundation of an entire subfield.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Training (real) | USPTO patent reactions | 1,094,235 | 2001-2013 applications, filtered by length |
| Training (gen) | Generated from Wade textbook templates | 865,118 | 75 reaction types, GDB-11 substrates |
| Testing (textbook) | Wade textbook problems | ~100 | 10 problem sets, 6-15 reactions each |
| Testing (scalability) | Generated from GDB-17 | 2,400 | 400 per atom count (11-16) |
Algorithms
- GRU-based encoder-decoder with attention mechanism
- PEG-based SMILES tokenizer
- Input sequence reversal
- Bucketed training with four bucket sizes
- TensorFlow seq2seq tutorial implementation with default learning rate
Models
| Parameter | Value |
|---|---|
| GRU layers | 3 |
| Embedding size | 600 |
| Input vocabulary | 311 tokens |
| Output vocabulary | 180 tokens |
| Buckets | (54,54), (70,60), (90,65), (150,80) |
Evaluation
| Metric | gen Model | real+gen Model | Notes |
|---|---|---|---|
| Textbook correct ratio | Variable by set | Higher on most sets | 10 problem sets |
| Average Tanimoto similarity | Variable | ~0.7 on scalability test | Morgan fingerprint based |
| Invalid SMILES ratio | Higher | ~0.4 on scalability test | Decreases with more training data |
Hardware
Not specified in the paper.
Paper Information
Citation: Nam, J. & Kim, J. (2016). Linking the Neural Machine Translation and the Prediction of Organic Chemistry Reactions. arXiv preprint, arXiv:1612.09529. https://arxiv.org/abs/1612.09529
Publication: arXiv preprint 2016
@article{nam2016linking,
title={Linking the Neural Machine Translation and the Prediction of Organic Chemistry Reactions},
author={Nam, Juno and Kim, Jurae},
journal={arXiv preprint arXiv:1612.09529},
year={2016},
doi={10.48550/arxiv.1612.09529}
}