Transformer-Based SMILES Embeddings for Property Prediction
This is a Method paper that introduces Transformer-CNN, a two-stage architecture for QSAR (Quantitative Structure-Activity Relationship) modeling. The primary contribution is a transfer learning approach: a Transformer model is first trained on the task of SMILES canonicalization (mapping non-canonical SMILES to canonical forms), and the encoder’s internal representations are then used as “dynamic SMILES embeddings” for downstream property prediction via a convolutional neural network (TextCNN). The authors also contribute an interpretability framework based on Layer-wise Relevance Propagation (LRP) that traces predictions back to individual atom contributions.
From Descriptors to Learned Embeddings in QSAR
Traditional QSAR methods rely on hand-engineered molecular descriptors (fragment counts, physicochemical features) coupled with feature selection and classical ML algorithms. While deep learning approaches that operate on raw SMILES strings or molecular graphs have reduced the need for manual feature engineering, they typically require large training datasets to learn effective representations from scratch. QSAR datasets, in contrast, often contain only hundreds of molecules, making it difficult to train end-to-end deep models.
The authors identify two specific gaps. First, existing SMILES-based autoencoders such as CDDD (Continuous and Data-Driven molecular Descriptors) produce fixed-length latent vectors, discarding positional information that could be useful for property prediction and interpretation. Second, QSAR models built on deep architectures generally lack interpretability, making it hard to verify that predictions rely on chemically meaningful structural features rather than spurious correlations.
Dynamic SMILES Embeddings via Canonicalization Pre-training
The core insight is that training a Transformer to perform SMILES canonicalization (a Seq2Seq task mapping non-canonical SMILES to canonical SMILES) produces an encoder whose internal states serve as information-rich, position-dependent molecular embeddings.
Pre-training on SMILES Canonicalization
The Transformer encoder-decoder is trained on approximately 17.7 million canonicalization pairs derived from the ChEMBL database (SMILES with length up to 110 characters). Each molecule is augmented 10 times by generating non-canonical SMILES variants, plus one identity pair where both sides are canonical. The training uses character-level tokenization with a 66-symbol vocabulary covering drug-like molecules including stereochemistry, charges, and inorganic ions.
The Transformer architecture follows Vaswani et al. with 3 layers and 10 self-attention heads. The learning rate schedule follows:
$$\lambda = \text{factor} \cdot \min(1.0,; \text{step} / \text{warmup}) / \max(\text{step},; \text{warmup})$$
where factor = 20, warmup = 16,000 steps, and $\lambda$ is clipped at a minimum of $10^{-4}$. Training runs for 10 epochs (275,907 batches per epoch) without early stopping.
On validation with 500,000 generated ChEMBL-like SMILES, the model correctly canonicalizes 83.6% of all samples. Performance drops for stereochemistry (37.2% for @-containing SMILES) and cis/trans notation (73.9%).
From Encoder States to QSAR Predictions
After pre-training, the encoder’s output for a molecule with $N$ characters is a matrix of dimensions $(N, \text{EMBEDDINGS})$. Unlike fixed-length CDDD descriptors, these “dynamic embeddings” preserve positional information, meaning equivalent characters receive different embedding values depending on their context and position.
To handle variable-length embeddings, the authors use a TextCNN architecture (from DeepChem) with 1D convolutional filters at kernel sizes (1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 15, 20) producing (100, 200, 200, 200, 200, 100, 100, 100, 100, 100, 160, 160) filters respectively. After GlobalMaxPool and concatenation, the features pass through Dropout (rate = 0.25), a Dense layer ($N = 512$), a Highway layer, and finally an output layer (1 neuron for regression, 2 for classification).
The Transformer weights are frozen during QSAR training. The Adam optimizer is used with a fixed learning rate of $10^{-4}$ and early stopping on a 10% held-out validation set. Critically, SMILES augmentation ($n = 10$) is applied during both training and inference, with the final prediction being the average over augmented SMILES for each molecule.
Interpretability via Layer-wise Relevance Propagation
The LRP algorithm propagates relevance scores from the output back through the CNN layers to the Transformer encoder output (which is position-wise). The relevance conservation property holds:
$$y = R = f(x) = \sum_{l \in (L)} R_{l} = \sum_{l \in (L-1)} R_{l} = \cdots = \sum_{l \in (1)} R_{l}$$
In practice, biases absorb some relevance, so the total propagated to the input is less than the output:
$$\sum_{l \in (L)} R_{l} = \sum_{l \in (L-1)} R_{l} + B$$
For gated connections in the Highway block, the authors implement the signal-take-all redistribution rule. The interpretation algorithm generates one SMILES per non-hydrogen atom (each drawn starting from that atom), runs LRP on each, and averages contributions. If more than 50% of relevance dissipates on biases, the interpretation may be unreliable, serving as an applicability domain indicator.
Benchmarks Across 18 Regression and Classification Datasets
The authors evaluate on the same 18 datasets (9 regression, 9 classification) used in their previous SMILES augmentation study, enabling direct comparison. All experiments use five-fold cross-validation.
Regression Results ($r^2$)
| Dataset | Descriptor-based | SMILES-based (augm=10) | Transformer-CNN (no augm) | Transformer-CNN (augm=10) | CDDD |
|---|---|---|---|---|---|
| MP (19,104) | 0.83 | 0.85 | 0.83 | 0.86 | 0.85 |
| BP (11,893) | 0.98 | 0.98 | 0.97 | 0.98 | 0.98 |
| BCF (378) | 0.85 | 0.85 | 0.71 | 0.85 | 0.81 |
| FreeSolv (642) | 0.94 | 0.93 | 0.72 | 0.91 | 0.93 |
| LogS (1,311) | 0.92 | 0.92 | 0.85 | 0.91 | 0.91 |
| Lipo (4,200) | 0.70 | 0.72 | 0.60 | 0.73 | 0.74 |
| BACE (1,513) | 0.73 | 0.72 | 0.66 | 0.76 | 0.75 |
| DHFR (739) | 0.62 | 0.63 | 0.46 | 0.67 | 0.61 |
| LEL (483) | 0.19 | 0.25 | 0.20 | 0.27 | 0.23 |
Classification Results (AUC)
| Dataset | Descriptor-based | SMILES-based (augm=10) | Transformer-CNN (no augm) | Transformer-CNN (augm=10) | CDDD |
|---|---|---|---|---|---|
| HIV (41,127) | 0.82 | 0.78 | 0.81 | 0.83 | 0.74 |
| AMES (6,542) | 0.86 | 0.88 | 0.86 | 0.89 | 0.86 |
| BACE (1,513) | 0.88 | 0.89 | 0.89 | 0.91 | 0.90 |
| ClinTox (1,478) | 0.77 | 0.76 | 0.71 | 0.77 | 0.73 |
| Tox21 (7,831) | 0.79 | 0.83 | 0.81 | 0.82 | 0.82 |
| BBBP (2,039) | 0.90 | 0.91 | 0.90 | 0.92 | 0.89 |
| JAK3 (886) | 0.79 | 0.80 | 0.70 | 0.78 | 0.76 |
| BioDeg (1,737) | 0.92 | 0.93 | 0.91 | 0.93 | 0.92 |
| RP AR (930) | 0.85 | 0.87 | 0.83 | 0.87 | 0.86 |
Key Comparisons
Baselines include descriptor-based methods (the best from LibSVM, Random Forest, XGBoost, ASNN, and DNNs), direct SMILES-based models with augmentation, and CDDD descriptors analyzed by the same classical ML methods. CDDD descriptors come from the Sml2canSml autoencoder approach, which produces fixed 512-dimensional vectors.
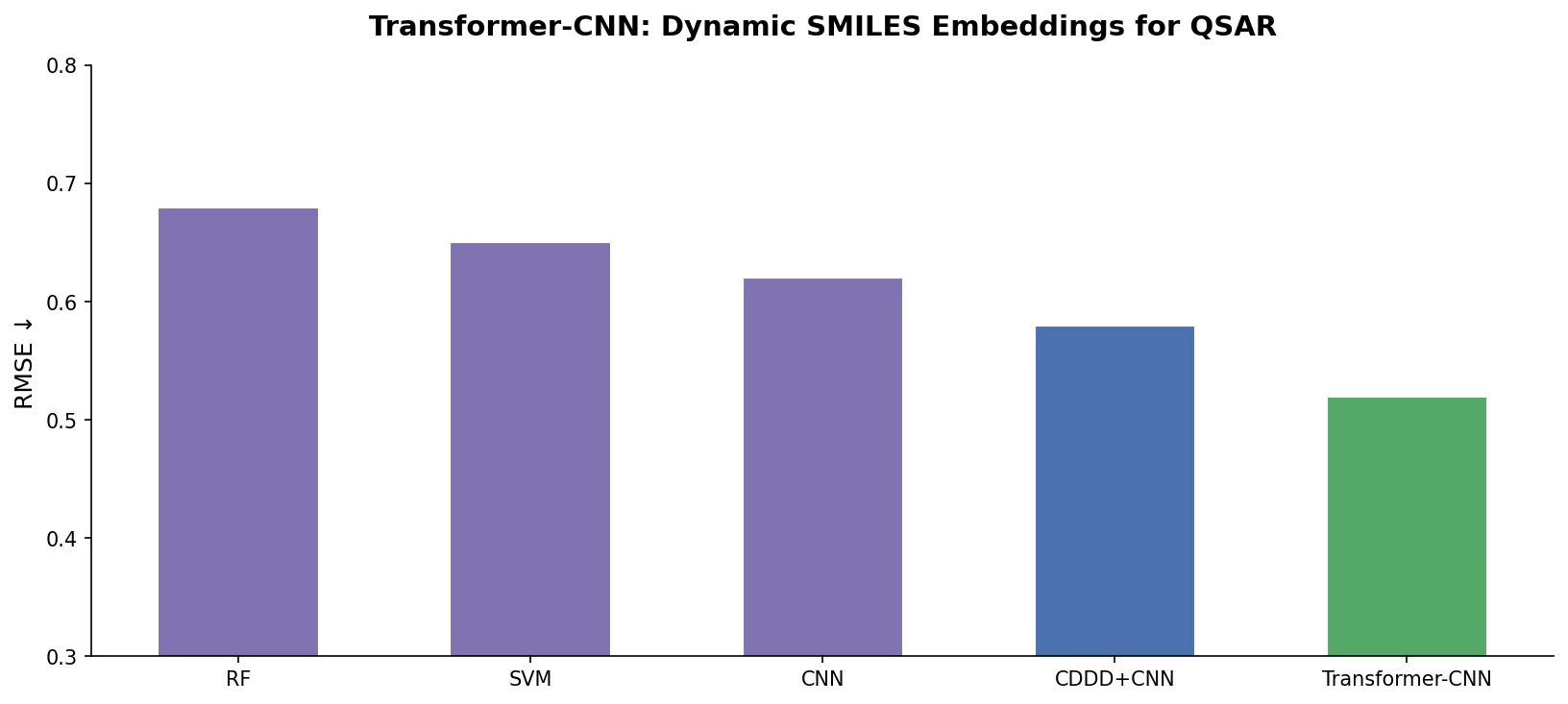
Transformer-CNN with augmentation matches or exceeds all baselines on 14 of 18 datasets. The effect of augmentation is dramatic: without it, Transformer-CNN underperforms substantially (e.g., BCF drops from 0.85 to 0.71, JAK3 from 0.78 to 0.70). This confirms that the internal consensus from multiple SMILES representations is essential to the method’s effectiveness.
A practical advantage over CDDD is that Transformer-CNN imposes no constraints on molecular properties (CDDD requires logP in (-5, 7), molecular weight under 12,600, 3-50 heavy atoms, and organic molecules only), since the Transformer was trained on the full diversity of ChEMBL.
Interpretability Case Studies
For AMES mutagenicity, the LRP analysis of 1-Bromo-4-nitrobenzene correctly identifies the nitro group and halogen as structural alerts, consistent with known mutagenicity rules. For aqueous solubility of haloperidol, the model assigns positive contributions to hydroxyl, carbonyl, and aliphatic nitrogen groups (which increase solubility) and negative contributions to aromatic carbons (which decrease it). Both cases align with established chemical knowledge, supporting the trustworthiness of the model.
Effective Transfer Learning for Small QSAR Datasets
Transformer-CNN achieves competitive or superior QSAR performance across 18 diverse benchmarks by combining three ingredients: (1) Transformer-based pre-training via SMILES canonicalization, (2) SMILES augmentation during training and inference, and (3) a lightweight CNN head. The method requires minimal hyperparameter tuning, as the Transformer weights are frozen and the CNN architecture is fixed.
The authors acknowledge several limitations and future directions:
- Stereochemistry canonicalization accuracy is low (37.2%), which could impact models for stereo-sensitive properties
- The LRP interpretability depends on sufficient relevance propagation (at least 50% reaching the input layer)
- The variance among augmented SMILES predictions could serve as a confidence estimate, but this is left to future work
- Applicability domain assessment based on SMILES reconstruction quality is proposed but not fully developed
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Pre-training | ChEMBL (SMILES <= 110 chars) | 17.7M pairs | 10x augmentation + 1 identity pair per molecule |
| Validation (canon.) | Generated ChEMBL-like SMILES | 500,000 | From a molecular generator |
| QSAR benchmarks | 9 regression + 9 classification | 378-41,127 | Available on OCHEM (https://ochem.eu) |
Algorithms
- Transformer: 3 layers, 10 self-attention heads, character-level tokenization (66 symbols)
- TextCNN: 12 kernel sizes (1-10, 15, 20) with 100-200 filters each, GlobalMaxPool, Dense(512), Highway, Dropout(0.25)
- Augmentation: n=10 non-canonical SMILES per molecule during training and inference
- LRP: signal-take-all redistribution for Highway gates, standard LRP for Dense and Conv layers
Models
- Transformer encoder weights pre-trained on canonicalization task (frozen during QSAR training)
- QSAR CNN trained with Adam optimizer, learning rate $10^{-4}$, early stopping
- Pre-trained embeddings and standalone prediction models available in the GitHub repository
Evaluation
- Regression: coefficient of determination $r^2 = 1 - SS_{\text{res}} / SS_{\text{tot}}$
- Classification: Area Under the ROC Curve (AUC)
- Five-fold cross-validation with bootstrap standard errors
Hardware
- NVIDIA Quadro P6000, Titan Xp, and Titan V GPUs (donated by NVIDIA)
- TensorFlow v1.12.0, RDKit v2018.09.2
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| transformer-cnn | Code | MIT | Source code, pre-trained embeddings, standalone prediction models |
| OCHEM | Other | N/A | Online platform hosting the method, training datasets, and models |
Paper Information
Citation: Karpov, P., Godin, G., & Tetko, I. V. (2020). Transformer-CNN: Swiss knife for QSAR modeling and interpretation. Journal of Cheminformatics, 12, 17. https://doi.org/10.1186/s13321-020-00423-w
@article{karpov2020transformer,
title={Transformer-{CNN}: Swiss knife for {QSAR} modeling and interpretation},
author={Karpov, Pavel and Godin, Guillaume and Tetko, Igor V.},
journal={Journal of Cheminformatics},
volume={12},
number={1},
pages={17},
year={2020},
publisher={Springer},
doi={10.1186/s13321-020-00423-w}
}