A General-Purpose RNN for Chemical Property Prediction from SMILES
SMILES2Vec is a Method paper that introduces a deep recurrent neural network architecture for predicting chemical properties directly from SMILES text representations. The primary contributions are: (1) a Bayesian-optimized CNN-GRU architecture that serves as a general-purpose predictor for diverse chemical properties (toxicity, activity, solubility, solvation energy), (2) an explanation mask mechanism that provides interpretable predictions by identifying which SMILES characters drive the network’s decisions, and (3) evidence that representation learning from raw SMILES can match or outperform models using hand-crafted molecular descriptors.
Motivation: Beyond Engineered Features in Chemical Modeling
At the time of writing (2017), deep learning models in chemistry relied heavily on engineered molecular descriptors and fingerprints as input features. Over 5,000 molecular descriptors had been developed since the late 1940s, and QSAR/QSPR modeling remained the dominant paradigm. The authors identified two key limitations with this approach:
- Restricted search space: Engineered features limit the neural network’s ability to discover potentially useful representations that domain experts have not anticipated.
- Incomplete domain knowledge: For complex properties where first-principles understanding is incomplete, the lack of appropriate descriptors constrains model performance.
In contrast, computer vision and NLP had shown that deep learning models trained on raw data (unaltered images, raw text) could learn powerful representations without feature engineering. The chemical SMILES notation, a text-based encoding of molecular structure that serves as the standard interchange format in cheminformatics, provided a natural analog to text data for NLP-style modeling.
A secondary motivation was interpretability. Most ML and DL models for chemistry operated as black boxes, which posed particular problems for regulated applications like FDA drug approval where mechanistic explanations are required.
Core Innovation: CNN-GRU Architecture with Explanation Masks
Architecture Design via Bayesian Optimization
SMILES2Vec treats SMILES strings as character-level text input. The network processes one-hot encoded characters (padded to length 250, covering 99.9% of the ChEMBL database) through three stages:
- Embedding layer: Maps one-hot character vectors to a learned embedding space (size 50)
- 1D convolutional layer: 192 filters with kernel size 3, stride 1
- Bidirectional GRU layers: Two layers with 224 and 384 units respectively
The authors explored four architectural classes (GRU, LSTM, CNN-GRU, CNN-LSTM) using Bayesian optimization via SigOpt. Each class was evaluated over 60 trials, optimizing embedding size, convolutional filter count, and RNN layer widths. The CNN-GRU class was selected as the best compromise: CNN-LSTM performed best on classification (Tox21), while GRU-based networks excelled at regression (FreeSolv). The final architecture is summarized by the hyperparameters:
| Component | Parameter | Value |
|---|---|---|
| Embedding | Size | 50 |
| Conv1D | Filters | 192 |
| BiGRU Layer 1 | Units | 224 |
| BiGRU Layer 2 | Units | 384 |
Explanation Mask for Interpretability
The explanation mask is a post-hoc interpretability mechanism. Given a trained (frozen) SMILES2Vec base model, a separate explanation network learns to produce a per-character mask over the input SMILES string. The mask is trained to preserve the base model’s output while masking as much input as possible. The loss function for a single sample is:
$$ \text{Loss}_i = | f(\text{SMILES}_i, \theta) - \text{Sol}(\text{SMILES}_i) |_2 + 10^{-6} | \text{MASK}_i |_2 + 0.05 , H(\text{MASK}_i) $$
where $f(\text{SMILES}_i, \theta)$ is the base network prediction, $\text{Sol}(\text{SMILES}_i)$ is the ground truth solubility, $H$ is the entropy of the normalized mask, and $\text{MASK}_i$ is the per-character mask vector. The L2 term encourages sparsity and the entropy term penalizes uniform attention distributions.
The explanation network itself is a 20-layer residual network with SELU activations, ending in a 1D convolution of length 1, batch normalization, and a softplus activation. The softplus output ranges from 0 (fully masked) to infinity (amplified attention), allowing the mask to both suppress and emphasize specific SMILES characters.
Experimental Setup and Baseline Comparisons
Datasets
The model was evaluated on four datasets from the MoleculeNet benchmark and the ESOL solubility dataset:
| Dataset | Property | Task | Size |
|---|---|---|---|
| Tox21 | Toxicity | Multi-task classification | 8,014 |
| HIV | Activity | Single-task classification | 41,193 |
| FreeSolv | Solvation energy | Single-task regression | 643 |
| ESOL | Solubility | Single-task regression | 1,128 |
SMILES strings longer than 250 characters were excluded. Classification datasets (Tox21, HIV) used 1/6 test split with minority class oversampling; regression datasets (FreeSolv, ESOL) used 1/10 test split. All experiments used 5-fold cross-validation.
Training Protocol
- Optimizer: RMSprop with learning rate $10^{-3}$, $\rho = 0.9$, $\epsilon = 10^{-8}$
- Batch size: 32
- Epochs: 250 with early stopping (patience of 25 epochs based on validation loss)
- Classification loss: Binary cross-entropy
- Regression loss: Mean absolute error
- Metrics: AUC for classification, RMSE for regression
Baselines
SMILES2Vec was compared against:
- MLP with engineered features: Standard multi-layer perceptron using molecular fingerprints (from MoleculeNet)
- Molecular graph convolutions: Graph-based neural network from MoleculeNet
- Chemception: CNN operating on 2D chemical images
Bayesian Optimization Protocol
Only two datasets were used for architecture optimization: the nr-ahr toxicity task from Tox21 (classification) and FreeSolv (regression). The remaining datasets (full Tox21, HIV, ESOL) served purely for generalization evaluation. A fixed test set was held out during optimization, and correlation between validation and test metrics (0.54 for Tox21, 0.78 for FreeSolv) confirmed limited overfitting to the validation set.
Results: Competitive Accuracy with Interpretable Predictions
Property Prediction Performance
SMILES2Vec achieved the following validation metrics (with a pre-training approach from ChemNet improving performance slightly):
| Dataset | Metric | SMILES2Vec | SMILES2Vec + Pre-training | Graph Conv |
|---|---|---|---|---|
| Tox21 | AUC | 0.80 | 0.81 | 0.81 |
| HIV | AUC | 0.78 | 0.80 | 0.80 |
| FreeSolv | RMSE (kcal/mol) | 1.4 | 1.2 | 1.3 |
| ESOL | RMSE | 0.63 | - | - |
Exact numbers for MLP and Chemception baselines were reported only in a bar chart (Figure 6) and not as precise values. The paper states that MLP with fingerprints performed worst across all tasks, and Chemception fell between MLP and the graph/SMILES methods.
Key findings:
- SMILES2Vec outperformed MLP models using engineered features across all tasks, despite using no feature engineering.
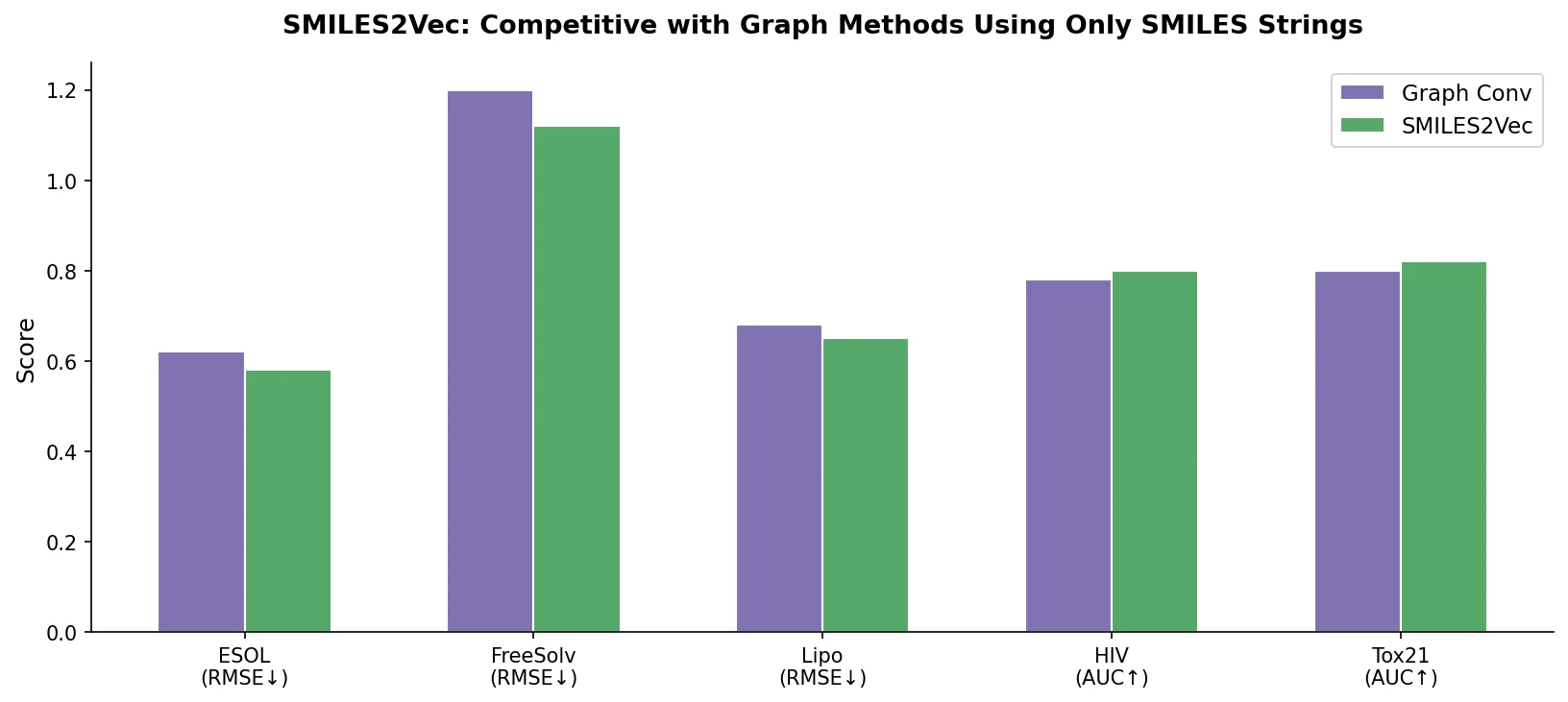
- Against graph convolutions (the state-of-the-art at the time), SMILES2Vec matched on classification (Tox21: 0.81 vs 0.81, HIV: 0.80 vs 0.80) and outperformed on regression (FreeSolv: 1.2 vs 1.3).
- SMILES2Vec outperformed Chemception (2D image CNN) on classification tasks but slightly underperformed on regression, which the authors attributed to SMILES lacking explicit atomic number information.
Interpretability Evaluation
On the ESOL solubility dataset, the explanation mask was evaluated against first-principles chemical knowledge. The authors separated compounds into soluble (> 1.0) and insoluble (< -5.0) categories and defined ground truth: soluble compounds should attend to hydrophilic atoms (O, N) while insoluble compounds should attend to hydrophobic atoms (C, F, Cl, Br, I). The top-3 character accuracy was 88%, confirming that SMILES2Vec learned representations consistent with known functional group chemistry.
Qualitative analysis of the masks showed that for low-solubility molecules, characters corresponding to hydrophobic groups (c, C, Cl) received high attention, while high-solubility molecules showed attention focused on hydrophilic groups (O, N).
Limitations
- The interpretability evaluation was limited to solubility, a well-understood property with simple first-principles rules. The authors acknowledged that quantifying interpretability for complex properties (toxicity, activity) where no simple ground truth exists is nontrivial.
- The Bayesian optimization used only a subset of datasets, so the architecture may not be globally optimal across all chemical tasks.
- SMILES strings lack explicit atomic number information, which may limit performance on physical property prediction compared to image or graph representations.
- The explanation mask approach requires training a separate 20-layer network per property, adding computational overhead.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Architecture optimization | Tox21 (nr-ahr task) | 8,014 | Single toxicity task for Bayesian optimization |
| Architecture optimization | FreeSolv | 643 | Solvation free energy regression |
| Evaluation | Tox21 (full, 12 tasks) | 8,014 | Multi-task classification |
| Evaluation | HIV | 41,193 | Single-task classification |
| Evaluation | ESOL | 1,128 | Solubility regression, also used for interpretability |
All datasets are publicly available through MoleculeNet. The ESOL dataset is from Delaney (2004).
Algorithms
- Bayesian optimization via SigOpt (60 trials per architectural class, 4 classes, 6 manually seeded initial designs per class)
- RMSprop optimizer with standard settings
- Explanation mask trained with Adam, learning rate annealed from $10^{-2}$ to $10^{-6}$
Models
- Final architecture: Embedding(50) -> Conv1D(192, kernel=3, stride=1) -> BiGRU(224) -> BiGRU(384)
- Explanation network: 20-layer residual network with SELU activations
- No pre-trained weights or code were released
Evaluation
| Metric | Dataset | Value | Notes |
|---|---|---|---|
| AUC | Tox21 | 0.81 | With pre-training |
| AUC | HIV | 0.80 | With pre-training |
| RMSE | FreeSolv | 1.2 kcal/mol | With pre-training |
| RMSE | ESOL | 0.63 | Base model |
| Top-3 accuracy | ESOL interpretability | 88% | Explanation mask |
Hardware
The authors report using TensorFlow with GPU acceleration via NVIDIA cuDNN libraries. Specific GPU models and training times were not reported.
Artifacts
No code, models, or data artifacts were released by the authors. The datasets used are publicly available through MoleculeNet.
Paper Information
Citation: Goh, G. B., Hodas, N. O., Siegel, C., & Vishnu, A. (2017). SMILES2Vec: An Interpretable General-Purpose Deep Neural Network for Predicting Chemical Properties. arXiv preprint arXiv:1712.02034.
@article{goh2017smiles2vec,
title={SMILES2Vec: An Interpretable General-Purpose Deep Neural Network for Predicting Chemical Properties},
author={Goh, Garrett B. and Hodas, Nathan O. and Siegel, Charles and Vishnu, Abhinav},
journal={arXiv preprint arXiv:1712.02034},
year={2017},
doi={10.48550/arxiv.1712.02034}
}