A Multitask BERT Framework for Molecular Property Prediction
MTL-BERT is a Method paper that introduces a multitask learning framework built on BERT for predicting molecular properties from SMILES strings. The primary contribution is the combination of three strategies to address data scarcity in drug discovery: (1) masked token pretraining on 1.7 million unlabeled molecules from ChEMBL, (2) multitask fine-tuning across 60 property prediction datasets simultaneously, and (3) SMILES enumeration as a data augmentation technique applied during pretraining, fine-tuning, and inference. The model achieves strong performance across 60 ADMET and molecular property datasets (44 classification and 16 regression), outperforming baselines including GNNs, XGBoost with molecular fingerprints, and prior SMILES-BERT approaches.
Data Scarcity in Molecular Property Prediction
Deep learning methods for molecular property prediction face a fundamental tension: they require large amounts of labeled data to learn effectively, but labeled bioactivity data is scarce due to the cost and time of laboratory experiments. Existing approaches at the time of publication addressed this in isolation. Graph neural networks (GNNs) learn from molecular graphs but are typically shallow (2-3 layers) and prone to overfitting on small datasets. The original SMILES-BERT model applied masked language modeling to SMILES strings but fine-tuned separately for each task, missing opportunities to share information across related properties. Fixed molecular representations like CDDD (continuous and data-driven descriptors) cannot be further optimized for specific downstream tasks.
The authors identify three specific gaps: (1) single-task fine-tuning wastes the correlations between related ADMET properties (e.g., lipophilicity relates to many ADMET endpoints), (2) using only canonical SMILES limits the model’s ability to learn robust molecular features, and (3) no prior work had combined pretraining, multitask learning, and SMILES enumeration into a unified framework.
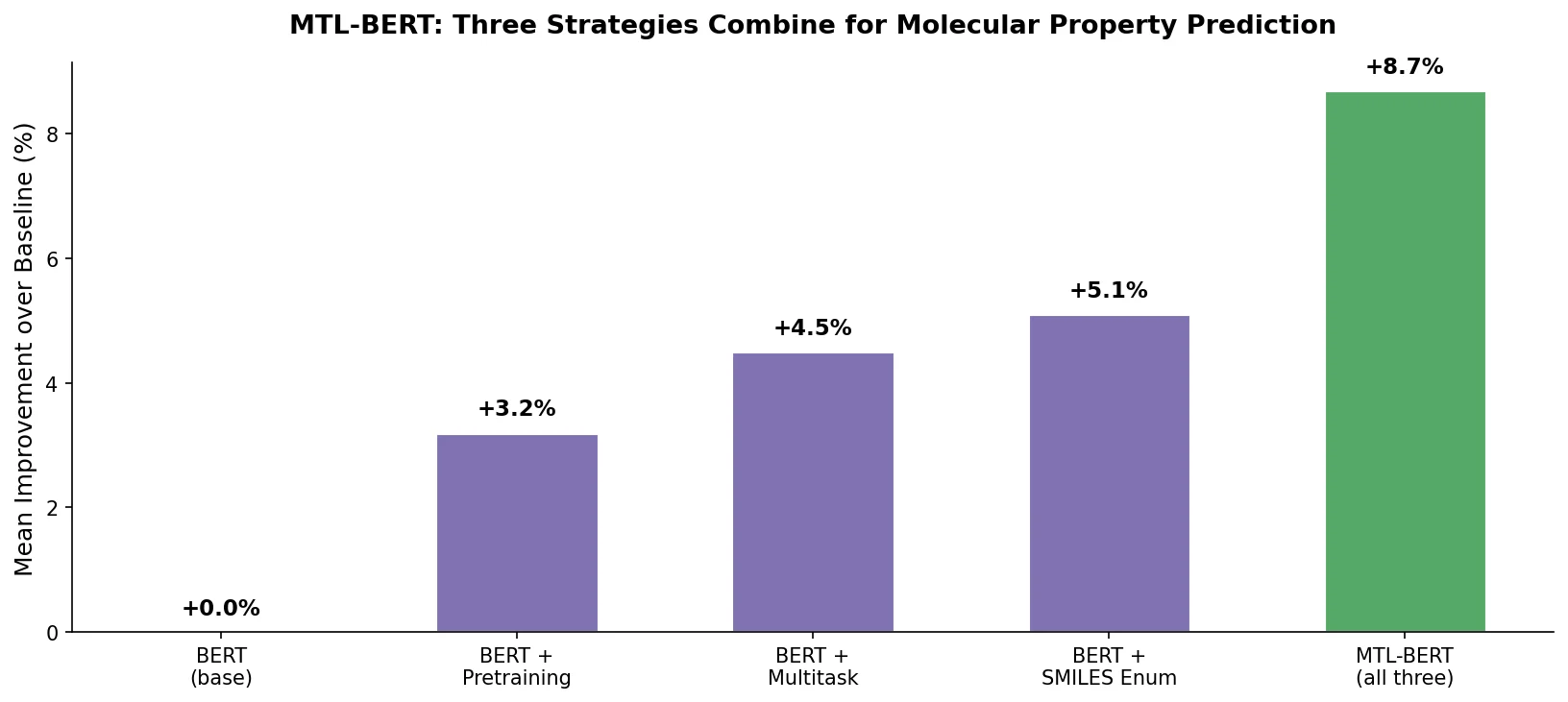
Three Strategies Combined: Pretraining, Multitask Learning, and SMILES Enumeration
The core innovation of MTL-BERT is the synergistic combination of three strategies in a single pipeline.
Masked SMILES Pretraining
Following the BERT paradigm, MTL-BERT pretrains on 1.7 million unlabeled molecules from ChEMBL using a masked token recovery task. For each SMILES string, 15% of tokens are randomly selected: 80% are replaced with a [MASK] token, 10% are replaced with a random token, and 10% remain unchanged. The loss is computed only at masked positions. Unlike the original BERT, MTL-BERT omits the next-sentence prediction task since there is no sequential relationship between SMILES strings (following the RoBERTa finding that this task is unnecessary).
SMILES strings are tokenized with a regular expression that captures multi-character tokens (e.g., Si, Br, Cl) and common SMILES syntax. The model uses positional encoding to capture token order.
Transformer Architecture
The model uses a standard Transformer encoder with multihead self-attention. The scaled dot-product attention computes:
$$\mathbf{O}_h = \text{softmax}\left(\frac{\mathbf{Q}_h \mathbf{K}_h^T}{\sqrt{d_k}}\right) \mathbf{V}_h$$
where $\mathbf{Q}_h$, $\mathbf{K}_h$, and $\mathbf{V}_h$ are the query, key, and value matrices for head $h$, and $\sqrt{d_k}$ is a scaling factor. The outputs from all heads are concatenated and projected. Each attention sublayer is followed by a position-wise feedforward network with GELU activation, layer normalization, and residual connections.
Three model sizes were compared:
| Model | Layers | Heads | Embedding Size | FFN Size | Recovery Accuracy | Fine-tuning Performance |
|---|---|---|---|---|---|---|
| MTL-BERT_SMALL | 4 | 4 | 128 | 512 | 0.931 | 0.826 |
| MTL-BERT_MEDIUM | 8 | 8 | 256 | 1,024 | 0.962 | 0.852 |
| MTL-BERT_LARGE | 12 | 12 | 576 | 2,304 | 0.974 | 0.848 |
The medium model was selected for its best fine-tuning performance with lower computational cost, despite the large model achieving higher pretraining recovery accuracy. The slight performance drop for the large model suggests mild overfitting.
Multitask Fine-tuning with Task Tokens
During fine-tuning, task tokens ([T0], [T1], …) are prepended to each input SMILES string. The Transformer output at each task token position is passed through a task-specific two-layer feedforward network for the corresponding prediction task. An attention mask prevents direct information exchange between task tokens, allowing each task to learn directly from SMILES tokens without interference. This design also reduces the discrepancy between pretraining (no task tokens visible) and fine-tuning.
Cross-entropy loss is used for classification tasks and mean squared error for regression tasks. The total multitask loss is a simple sum of per-task losses without learned weighting.
SMILES Enumeration as Data Augmentation
A molecule can be represented by multiple valid SMILES strings by varying starting atoms and traversal orders. MTL-BERT applies SMILES enumeration at all three stages:
- Pretraining: Enumerated SMILES increase diversity of the self-supervised training data.
- Fine-tuning: Each dataset is augmented 20x with random SMILES variants, increasing data diversity and helping the model learn position-invariant features.
- Inference: Multiple SMILES are generated per test molecule, predictions are fused (averaged) for a more robust final prediction.
The 20x augmentation factor was chosen based on prior work showing diminishing returns beyond this level while significantly increasing computational cost.
Experimental Evaluation Across 60 Datasets
Setup
MTL-BERT was evaluated on 60 datasets (44 classification, 16 regression) covering ADMET properties and common molecular benchmarks. Datasets were sourced from ADMETlab and MoleculeNet. Each dataset was split 8:1:1 (train/validation/test), and experiments were repeated 10 times with random splits, reporting mean and standard deviation.
Classification tasks were evaluated with ROC-AUC and accuracy; regression tasks with $R^2$ and RMSE.
Baselines
Five baselines were compared:
- ECFP4-XGBoost: Extended-connectivity fingerprints (diameter 4) with gradient boosting
- Graph Attention Network (GAT)
- Graph Convolutional Network (GCN)
- AttentiveFP: A GNN with attention for molecular property prediction
- CDDD: Continuous and data-driven descriptors from a pretrained RNN auto-encoder
Ablation Study
Three model variants were compared to isolate contributions:
- MTL-BERT: Full model (pretraining + multitask + SMILES enumeration)
- STL-BERT: Single-task fine-tuning with SMILES enumeration (no multitask)
- Cano-BERT: Canonical SMILES only, single-task fine-tuning (equivalent to SMILES-BERT)
Cano-BERT showed more than 10% degradation on several datasets (CL, Fu, LC50DM) compared to STL-BERT, demonstrating the importance of SMILES enumeration. MTL-BERT outperformed STL-BERT on most datasets, with improvements exceeding 5% on $F_{20\%}$, SR-ARE, and SR-ATAD5, confirming that multitask learning provides additional benefit on top of enumeration.
Results vs. Baselines
MTL-BERT outperformed all baselines on nearly all 60 datasets. Specific findings:
- ECFP4-XGBoost performed inconsistently, doing well on some tasks (e.g., $F_{30\%}$, BACE, CL) but poorly on others, reflecting the limitation of fixed-length fingerprint representations.
- GNNs generally improved over fingerprints but still suffered from data scarcity, falling behind ECFP4-XGBoost by more than 3% on $F_{30\%}$, Carcinogenicity, CL, and VD.
- MTL-BERT surpassed all baselines except on CYP2C19-sub and BACE (by less than 1.1%).
- On 14 tasks (NR-ER, NR-PPAR-gamma, SR-ARE, SR-ATAD5, SR-HSE, SR-MMP, Bioconcentration Factor, Fu, LC50FM, Lipophilicity, CL, PPB, VD, LC50DM), MTL-BERT exceeded the best baseline by more than 5-10%.
- Improvements were statistically significant at the 95% confidence level (paired t-test, $P \leq 0.001$).
Representation Analysis
t-SNE visualization of pretrained token embeddings (from 1,000 randomly selected molecules, approximately 35,000 tokens) showed that:
- Tokens of the same type cluster together (capturing atomic type information).
- Within type clusters, sub-groups correspond to different chemical environments (e.g., oxygen atoms in nitrate groups vs. carbonyl groups).
- Nearby embeddings share similar molecular neighborhood environments.
Attention-based Interpretability
The model’s attention weights provide interpretability for predictions:
- For a solubility task (LogS/LogD), attention concentrated on polar groups, which are known determinants of aqueous solubility.
- For AMES (mutagenicity), attention focused on azide, nitrosamide, acylchloride, and nitrite groups, which are known mutagenic structural alerts.
Performance Gains from Combined Strategies with Interpretable Attention
MTL-BERT demonstrates that the combination of pretraining, multitask learning, and SMILES enumeration is more effective than any individual strategy for molecular property prediction. The ablation study provides clear evidence for the additive benefit of each component.
Key strengths include the breadth of evaluation (60 datasets covering diverse ADMET endpoints), the consistent improvement over multiple baseline types (fingerprints, GNNs, pretrained representations), and the interpretable attention mechanism that highlights chemically meaningful substructures.
Limitations to note: the simple sum of multitask losses (no learned task weighting) may not be optimal when tasks have very different scales or when some tasks are unrelated. The authors observe slight degradation on a few datasets (AMES, CYP1A2-Sub, FreeSolv), suggesting negative transfer in those cases. The 20x SMILES enumeration significantly increases computational cost during fine-tuning and inference. The paper does not report wall-clock training times or GPU hours, making it difficult to assess the practical cost of the enumeration strategy. Hardware details are not specified beyond acknowledgment of the High-Performance Computing Center at Central South University.
The hierarchical clustering of task representations reveals meaningful task groupings (e.g., LogD and LogP cluster together due to their shared relationship with water solubility), supporting the premise that multitask learning captures cross-task correlations.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Pretraining | ChEMBL | 1.7M molecules | Unlabeled SMILES; 10% held out for evaluation |
| Fine-tuning/Evaluation | ADMETlab + MoleculeNet | 60 datasets (44 classification, 16 regression) | 8:1:1 train/val/test split |
Algorithms
- Pretraining: Masked token prediction (15% masking rate: 80% [MASK], 10% random, 10% unchanged). Adam optimizer, learning rate 1e-4, batch size 512, 50 epochs.
- Fine-tuning: Adam optimizer, learning rate 5e-5, batch size 64, dropout 0.1. Cross-entropy for classification, MSE for regression. Early stopping with patience 20, max 200 epochs.
- SMILES enumeration: 20x augmentation. Repeated search up to 100 times if enumerated SMILES is identical to a previous one.
- Inference fusion: Predictions from multiple enumerated SMILES are averaged.
Models
- MTL-BERT_MEDIUM (selected model): 8 layers, 8 attention heads, 256 embedding size, 1,024 FFN size
- Pretraining recovery accuracy: 0.962
- 1,000 task tokens pre-allocated for future tasks
Evaluation
| Metric | Task Type | Notes |
|---|---|---|
| ROC-AUC | Classification | Primary metric |
| Accuracy | Classification | Secondary metric |
| $R^2$ | Regression | Primary metric |
| RMSE | Regression | Secondary metric |
All experiments repeated 10 times with random splits; mean and standard deviation reported.
Hardware
Hardware specifications are not reported in the paper. The authors acknowledge the High-Performance Computing Center of Central South University.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| MTL-BERT | Code | Not specified | Official implementation |
| ChEMBL | Dataset | CC BY-SA 3.0 | Pretraining data source |
| MoleculeNet | Dataset | MIT | Fine-tuning benchmark |
| ADMETlab | Dataset | Free for academic use | ADMET property datasets |
Paper Information
Citation: Zhang, X.-C., Wu, C.-K., Yi, J.-C., Zeng, X.-X., Yang, C.-Q., Lu, A.-P., Hou, T.-J., & Cao, D.-S. (2022). Pushing the boundaries of molecular property prediction for drug discovery with multitask learning BERT enhanced by SMILES enumeration. Research, 2022, Article 0004. https://doi.org/10.34133/research.0004
@article{zhang2022mtlbert,
title={Pushing the Boundaries of Molecular Property Prediction for Drug Discovery with Multitask Learning BERT Enhanced by SMILES Enumeration},
author={Zhang, Xiao-Chen and Wu, Cheng-Kun and Yi, Jia-Cai and Zeng, Xiang-Xiang and Yang, Can-Qun and Lu, Ai-Ping and Hou, Ting-Jun and Cao, Dong-Sheng},
journal={Research},
volume={2022},
pages={Article 0004},
year={2022},
doi={10.34133/research.0004},
publisher={American Association for the Advancement of Science (AAAS)}
}