This group covers models specifically designed for predicting molecular or crystal properties from chemical string representations. It includes SMILES-based QSAR architectures, transfer learning approaches, multitask prediction, hybrid prediction-generation models, and text-based crystal property prediction. Also included are studies on using language model perplexity as an intrinsic molecular scoring method and evaluations of how well language models capture complex molecular distributions.
| Paper | Year | Approach | Key Idea |
|---|---|---|---|
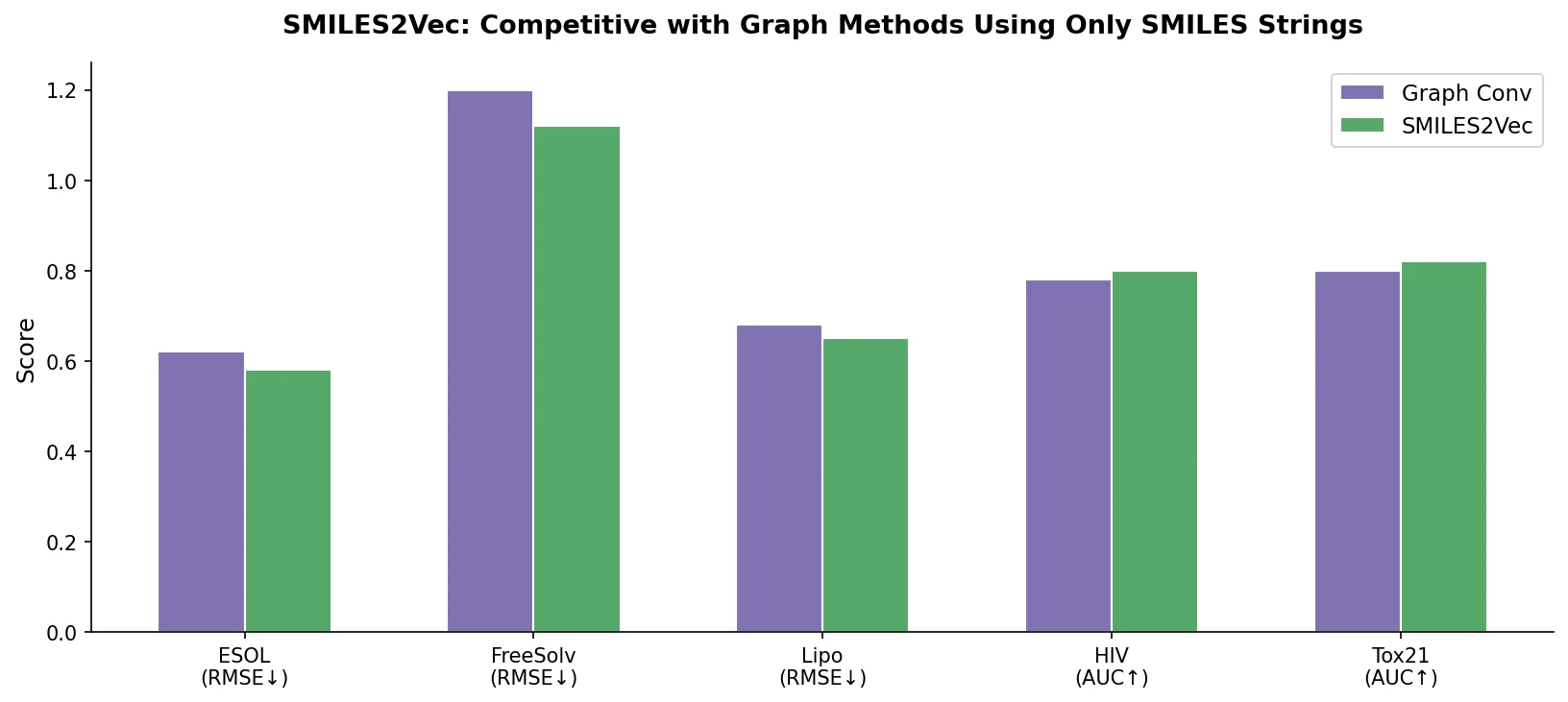
| SMILES2Vec | 2017 | CNN-GRU | Interpretable property prediction from raw SMILES embeddings |
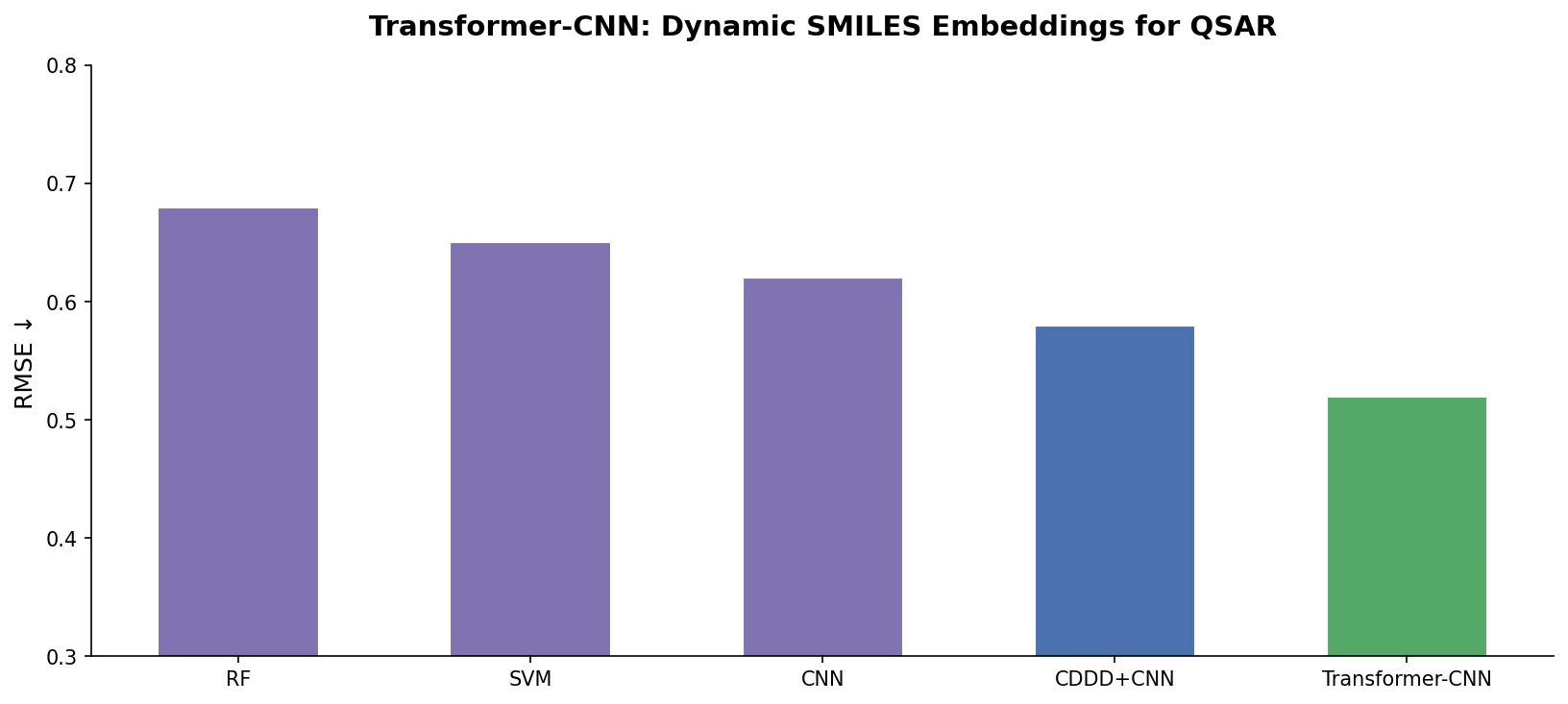
| Transformer-CNN | 2020 | Transformer + CNN | Transformer SMILES embeddings with CNN for interpretable QSAR |
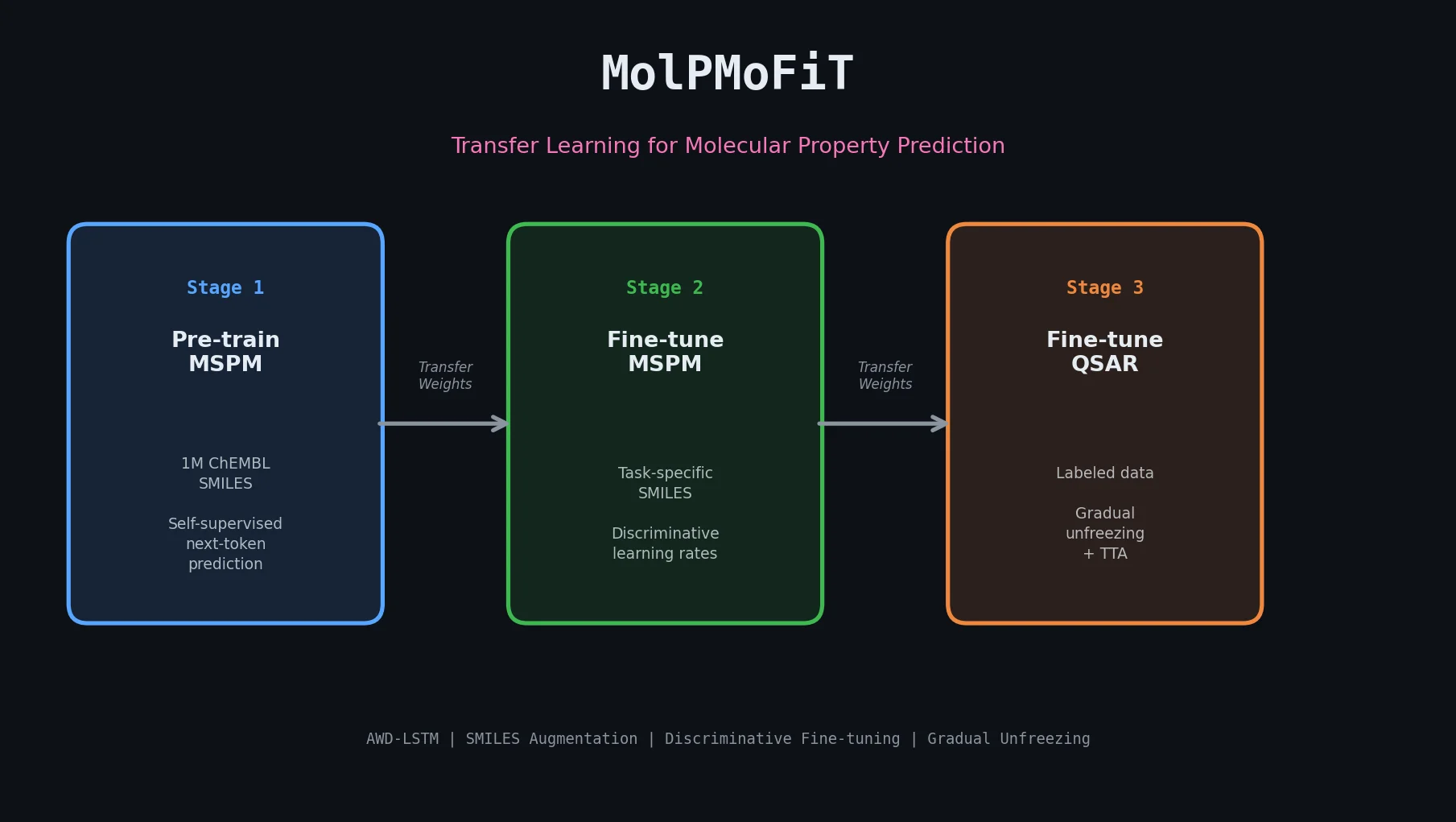
| MolPMoFiT | 2020 | Transfer learning | ULMFiT-style inductive transfer for QSAR on small datasets |
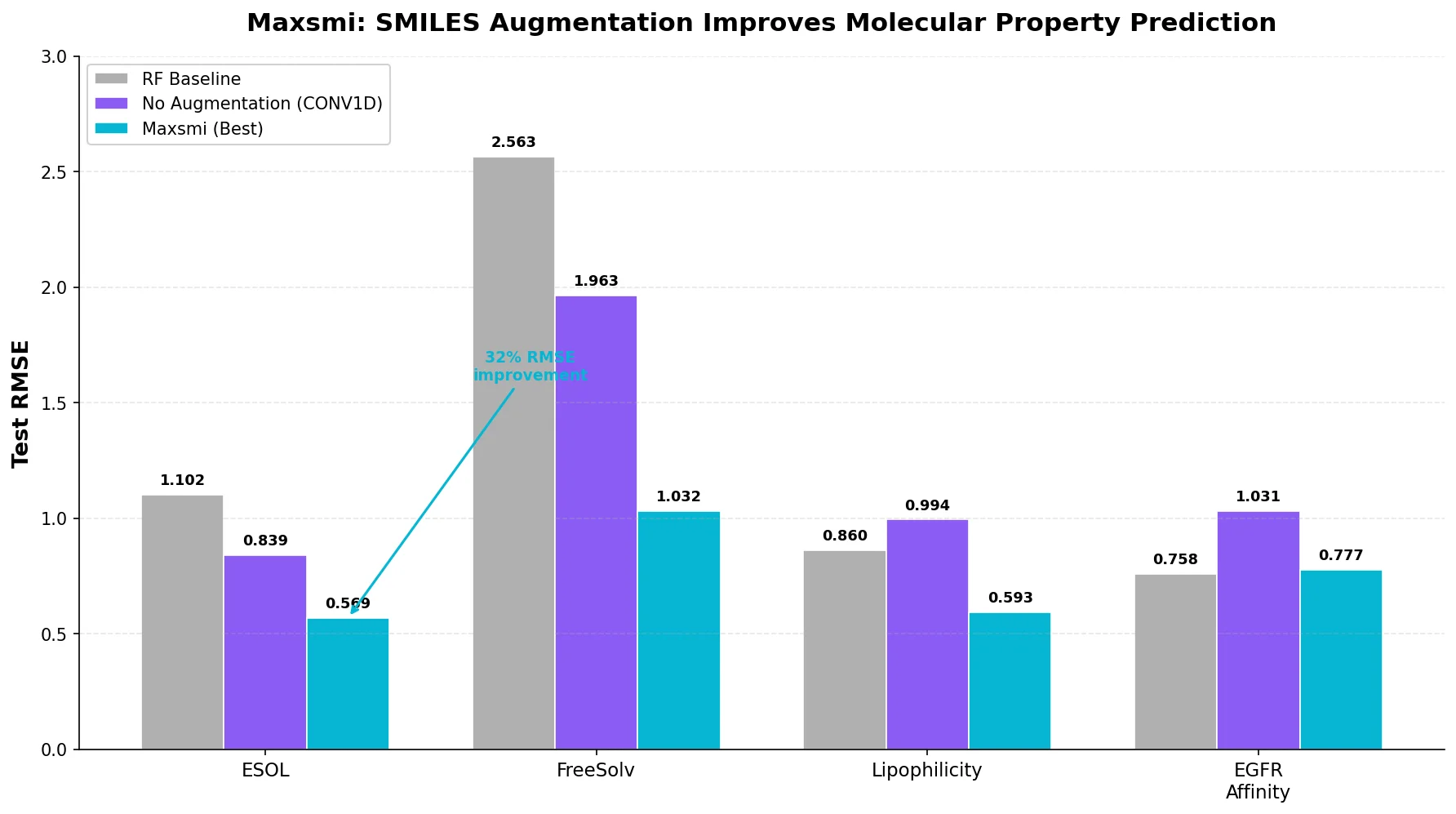
| Maxsmi | 2021 | CNN/RNN | SMILES augmentation improves CNN and RNN property prediction |
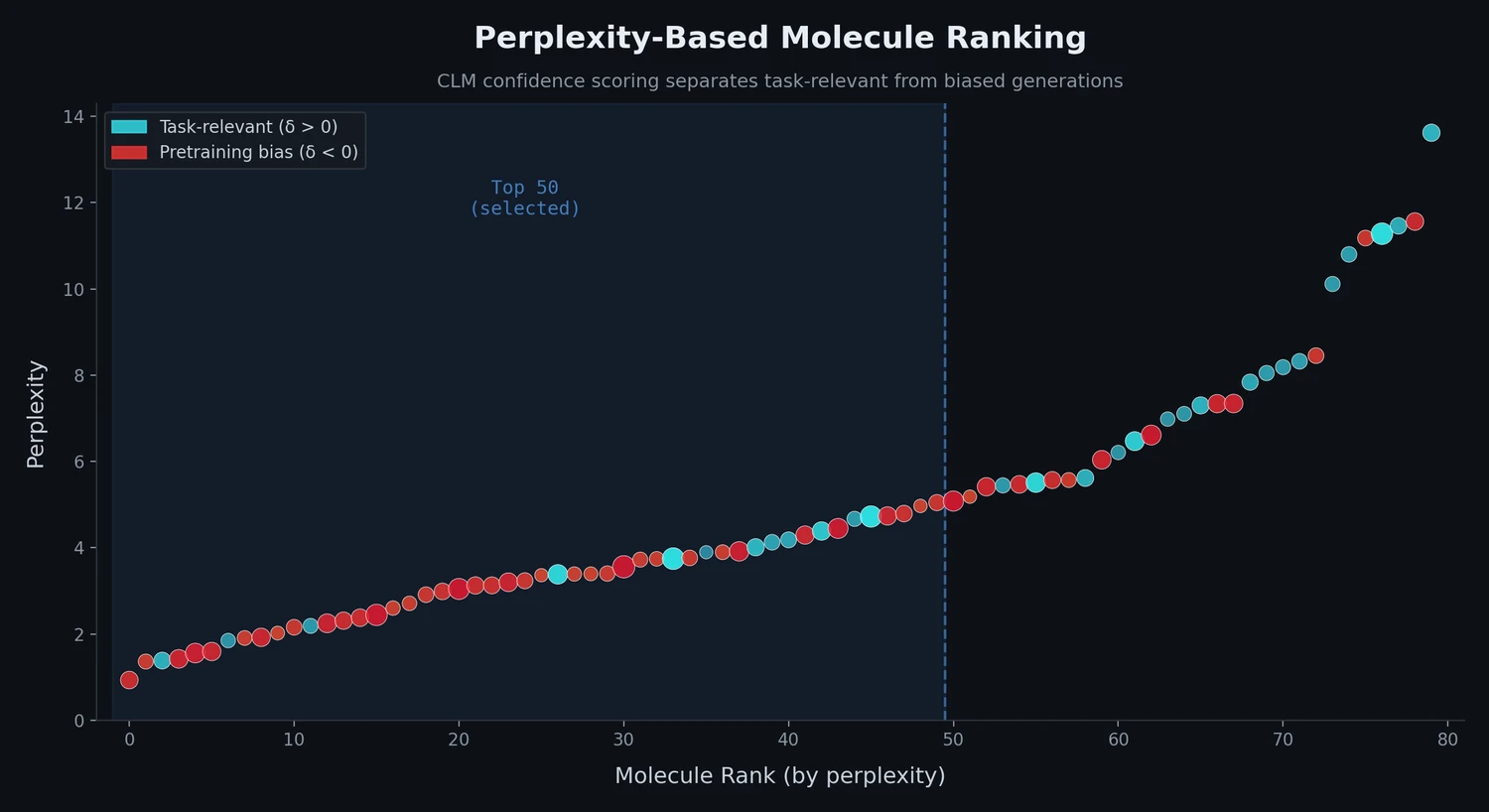
| Perplexity Ranking | 2022 | LM scoring | Perplexity scores rank molecules and detect pretraining bias |
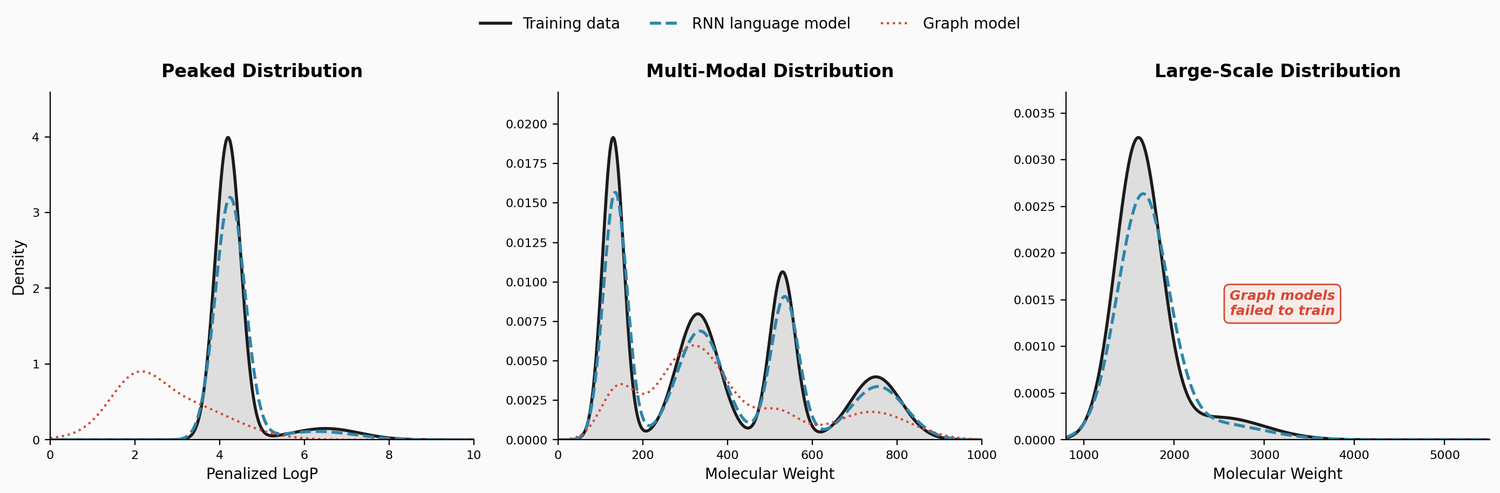
| LM Distributions | 2022 | RNN LM | RNN language models capture complex molecular distributions |
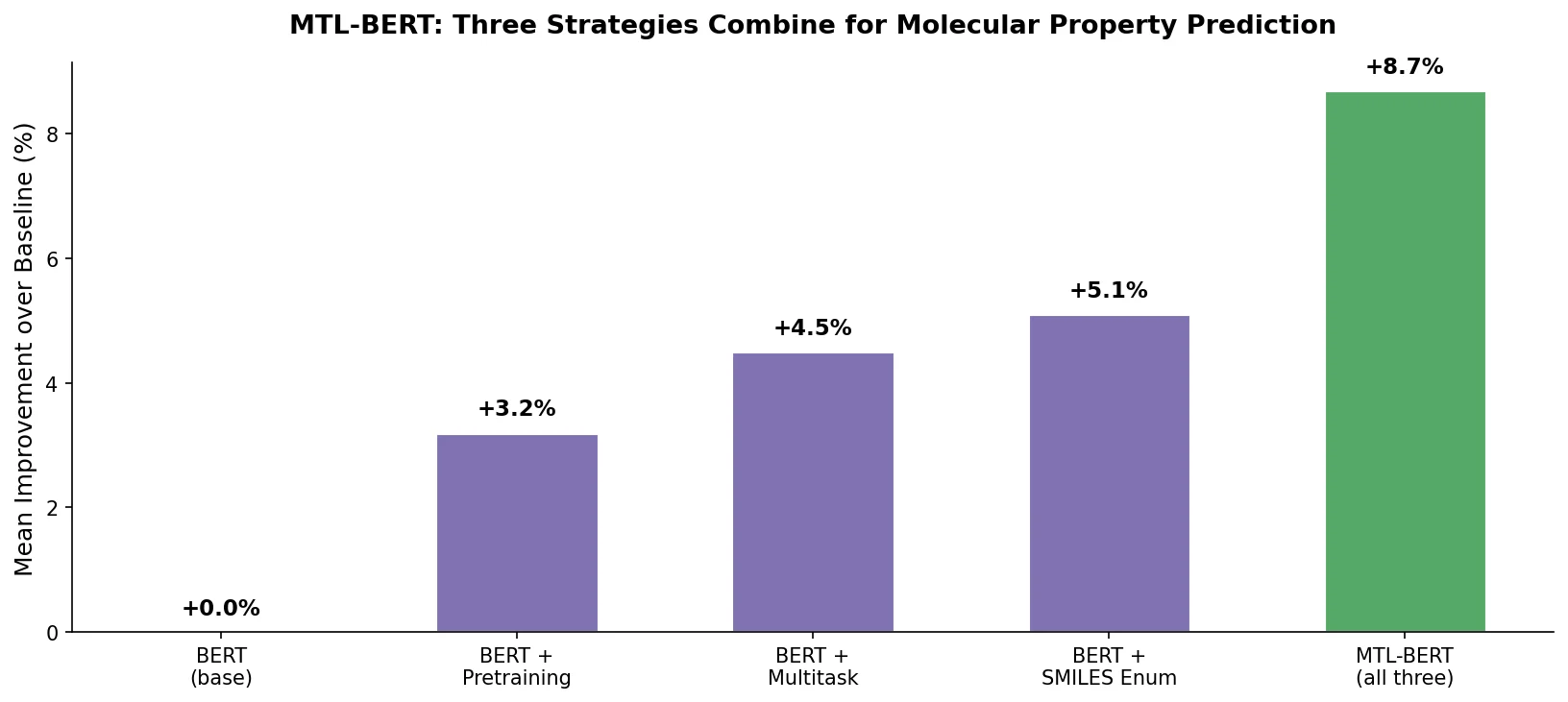
| MTL-BERT | 2022 | BERT | Multitask pretraining with SMILES enumeration augmentation |
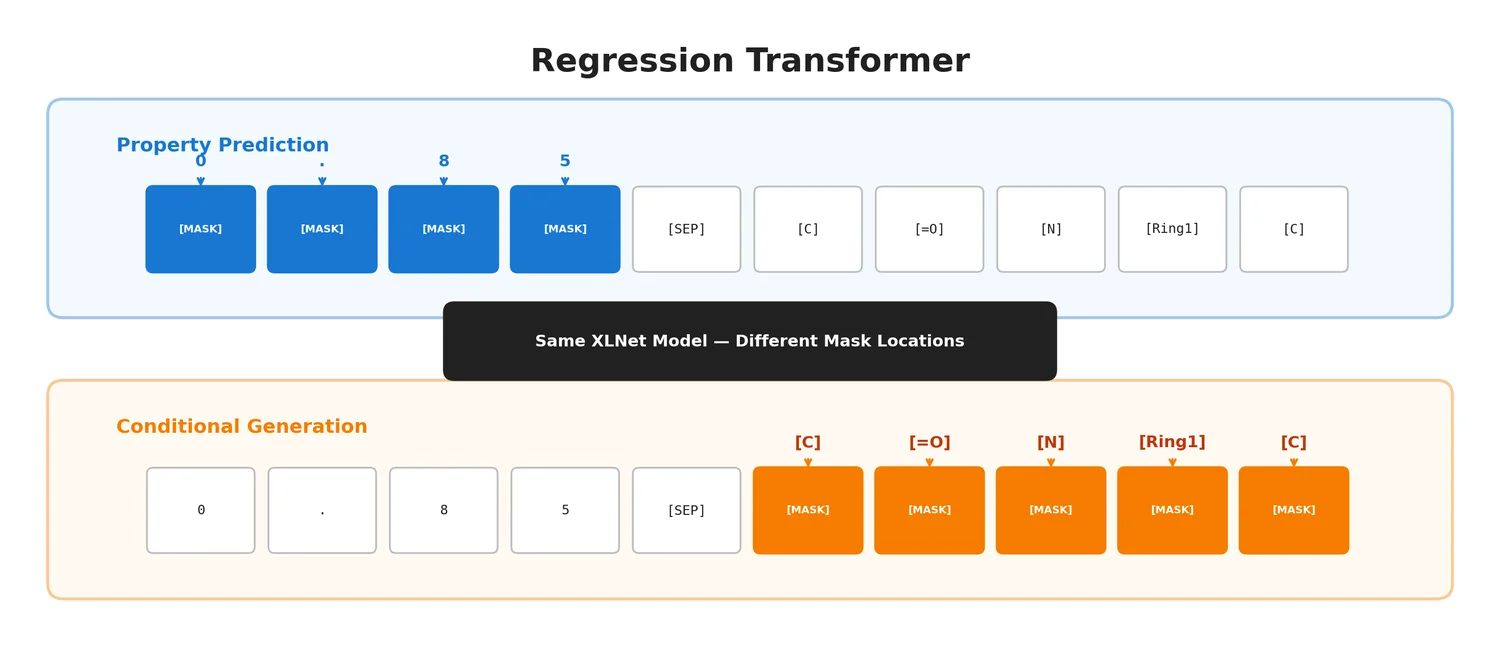
| Regression Transformer | 2023 | Transformer | Unifies property prediction and conditional generation in one model |
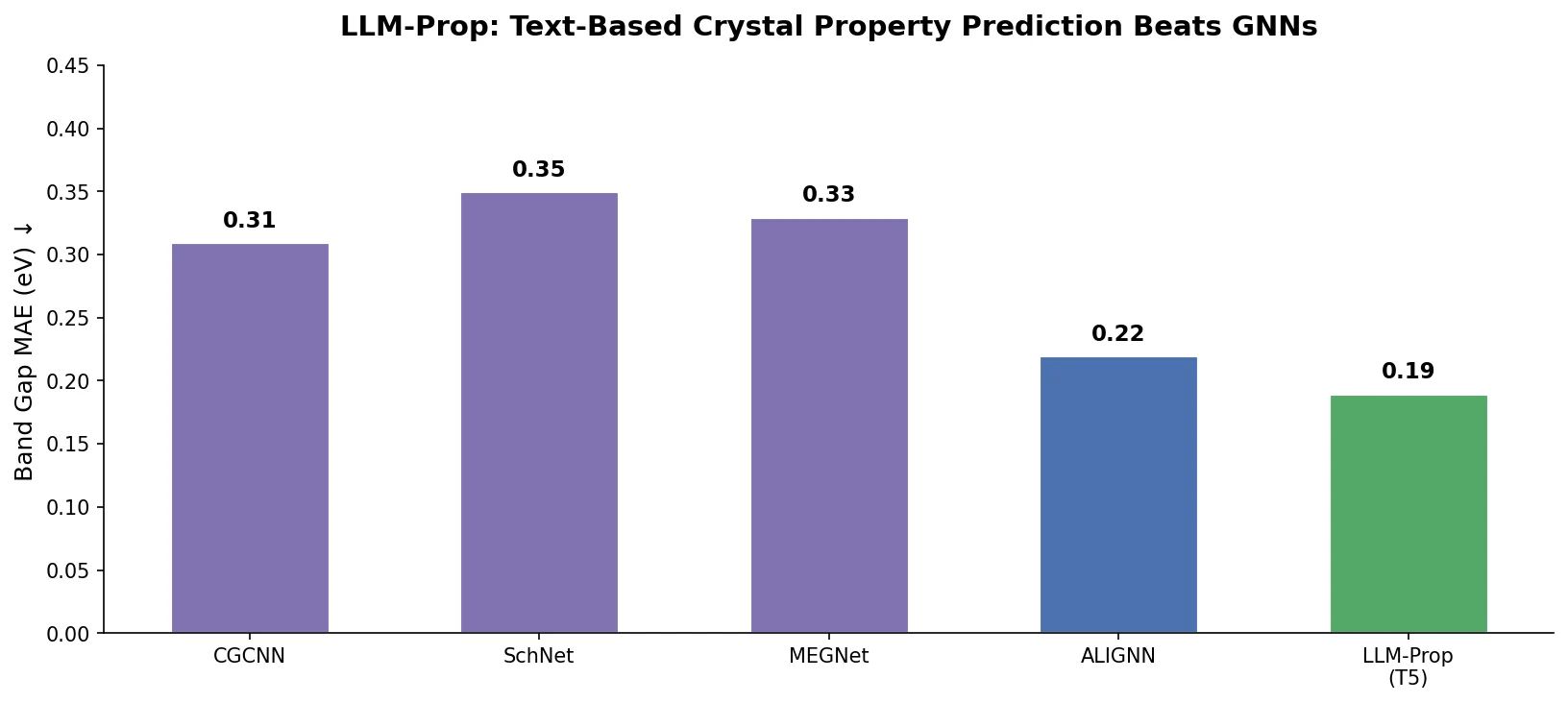
| LLM-Prop | 2025 | T5 | Crystal property prediction from text descriptions |