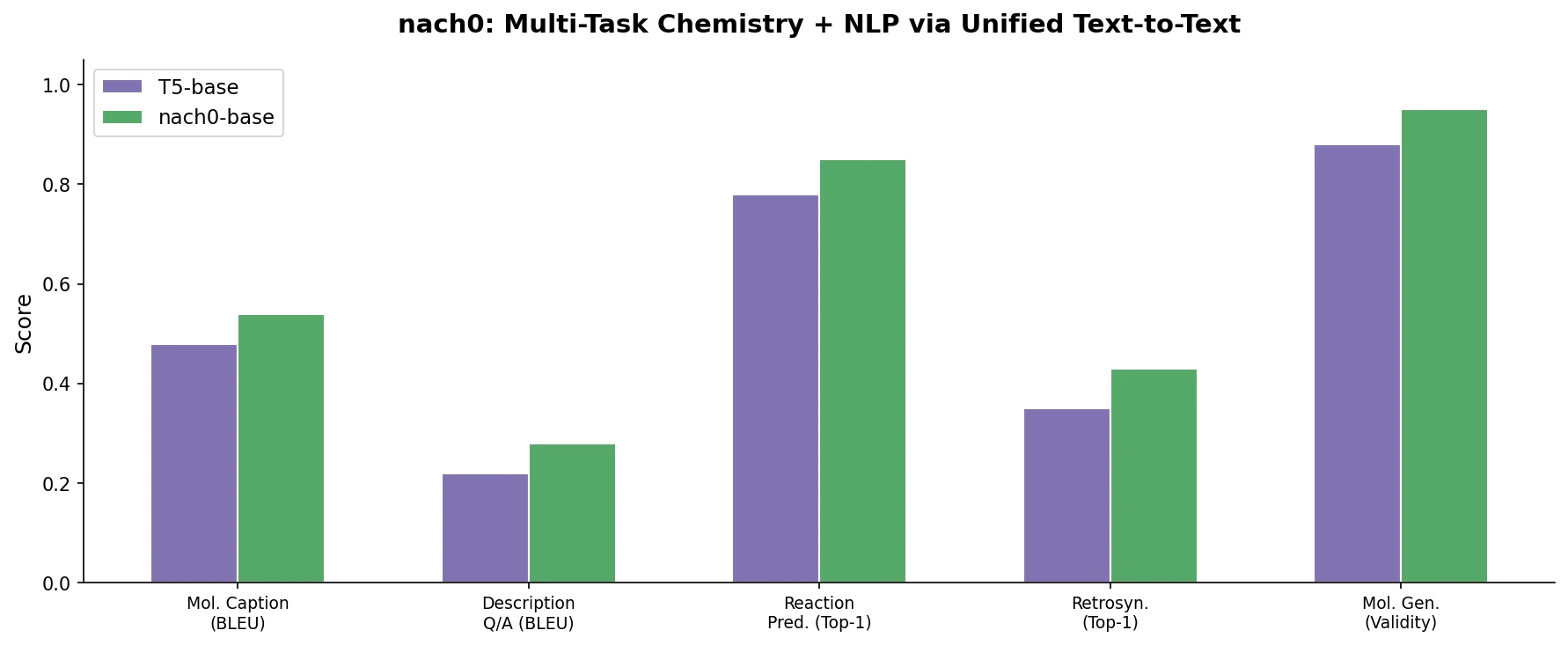
A Multi-Domain Encoder-Decoder for Chemistry and NLP
nach0 is a Method paper that introduces a unified encoder-decoder foundation model capable of handling both natural language processing (NLP) tasks and chemistry tasks within a single architecture. The primary contribution is demonstrating that a T5-based model pre-trained on scientific text, patents, and SMILES molecular strings can be instruction-tuned to perform molecular property prediction, reaction prediction, molecular generation, named entity recognition, question answering, and cross-domain translation (text-to-molecule and molecule-to-text) simultaneously. The model is available in base (250M parameters) and large (780M parameters) configurations.
Bridging Chemical and Linguistic Representations
Most existing biomedical language models (BioBERT, SciFive, BioMegatron) are trained exclusively on natural language text from sources like PubMed, omitting chemical structure information encoded in SMILES strings. Conversely, chemistry-specific models trained on SMILES data often lack the ability to process natural language instructions or perform NLP tasks. Models like Galactica and MolT5 attempted to bridge this gap by training on both natural language and chemical data, but they were not fine-tuned on a diverse set of chemical tasks using instruction tuning in a multi-task fashion.
nach0 addresses this by creating a shared representation space for both modalities and fine-tuning across a comprehensive set of tasks spanning three domains: NLP-only tasks, chemistry-only tasks, and cross-domain tasks that require translating between natural language and molecular representations.
Unified Text-to-Text Framework with SMILES Tokenization
The core innovation in nach0 is formulating all chemical and linguistic tasks as text-to-text problems within a single encoder-decoder transformer, combined with a specialized SMILES tokenization strategy.
SMILES Token Integration
Rather than treating SMILES as plain text, nach0 extends the T5 vocabulary with dedicated SMILES tokens. Each SMILES token is annotated with special symbols in the format <sm_{token}>, creating a distinct vocabulary space for molecular representations while preserving the natural language vocabulary from FLAN-T5. The embedding matrix is initialized by reusing learned embeddings from the pre-trained model for original tokens, with new chemical tokens initialized from the first embeddings.
Architecture
Both model sizes use the standard T5 encoder-decoder architecture:
| Configuration | Parameters | Layers | Hidden Size | FFN Size | Attention Heads |
|---|---|---|---|---|---|
| Base | 250M | 12 | 768 | 3072 | 12 |
| Large | 780M | 24 | 1024 | 4096 | 16 |
Pre-training Data
The model is pre-trained with a language modeling objective on three data sources:
| Source | Documents | Tokens |
|---|---|---|
| PubMed abstracts (chemistry-filtered) | 13M | 355M |
| USPTO patent descriptions | 119K | 2.9B |
| ZINC molecular database | ~100M | 4.7B |
Instruction Tuning
Following the approach of Raffel et al. and Chung et al., nach0 uses natural language prompts to formulate each task. For example, a retrosynthesis task might be phrased as “What reactants could be used to synthesize [SMILES]?” and a property prediction task as “Can [SMILES] penetrate the BBB?” This enables multi-task training across all domains with a single loss function and shared hyperparameters.
Training uses a batch size of 1024, learning rate of $1 \times 10^{-4}$, and weight decay of 0.01. Pre-training runs for one epoch, and fine-tuning for 10 epochs. Data mixing follows the examples-proportional mixing strategy from T5.
Multi-Task Evaluation Across NLP and Chemistry Benchmarks
nach0 is evaluated on a comprehensive set of benchmarks spanning three task categories.
Task Categories
NLP tasks: Named entity recognition (BC5CDR-Chemical, BC5CDR-Disease, NCBI-Disease, BC2GM, JNLPBA), PICO extraction (EBM PICO), textual entailment (MedNLI, SciTail), relation extraction (ChemProt, DDI, GAD), sentence similarity (BIOSSES), document classification (HoC), and question answering (PubMedQA, BioASQ, MedMCQA, MMLU).
Chemistry tasks: Molecular property prediction (ESOL, FreeSolv, Lipophilicity, BBBP, HIV, BACE from MoleculeNet; QM9 from Mol-Instructions), molecular generation (MOSES), forward reaction prediction, reagent prediction, and retrosynthesis (from Mol-Instructions/USPTO).
Cross-domain tasks: Description-guided molecule design and molecular description generation (from Mol-Instructions).
Baselines
nach0 is compared against FLAN-T5 (250M), SciFive (220M), and MolT5 (220M), all trained in multi-task fashion.
Key Results
On chemistry and cross-domain tasks, nach0 base consistently outperforms all base-sized baselines. Selected highlights from Table 3:
| Task | Metric | MolT5 | SciFive | FLAN | nach0 Base | nach0 Large |
|---|---|---|---|---|---|---|
| Forward reaction | Acc@1 | 27.0% | 60.0% | 59.0% | 88.0% | 89.9% |
| Retrosynthesis | Acc@1 | 15.0% | 31.0% | 31.0% | 53.0% | 56.3% |
| Reagent prediction | Acc@1 | 1.1% | 3.8% | 4.0% | 6.3% | 13.1% |
| BACE | BA | 0.58 | 0.65 | 0.65 | 0.74 | 0.71 |
| BBBP | BA | 0.55 | 0.66 | 0.60 | 0.67 | 0.68 |
| HFE (FreeSolv) | R2 | -0.36 | 0.51 | 0.55 | 0.77 | 0.78 |
| MOSES (FCD) | FCD/Test | 0.521 | 0.578 | 0.529 | 0.311 | 0.304 |
| Description-guided mol. design | BLEU-2 | 30.3% | 44.2% | 43.6% | 49.0% | 48.8% |
| Mol. description gen. | BLEU-2 | 35.6% | 39.6% | 38.6% | 43.9% | 41.7% |
On NLP tasks, nach0 base performs comparably to FLAN base, with the two models trading wins across different tasks. nach0 large improves substantially over nach0 base on most tasks.
Ablation Study
The ablation study (Table 4) examines the impact of multi-task training across chemical task groups. Key findings:
- nach0 trained on all chemical tasks jointly outperforms models trained on individual task groups (prediction-only, reaction-only, or generation-only) on the total set of metrics
- The joint model shows lower novelty scores on MOSES compared to the generation-only model, but this reflects less overfitting to training data rather than worse performance
- nach0 consistently outperforms MolT5 across all chemical task configurations, demonstrating the benefit of pre-training on both natural language and chemical data with specialized SMILES tokens
Case Studies
Two applied case studies demonstrate nach0 in drug discovery scenarios:
End-to-end drug discovery for diabetes mellitus: Using a sequence of prompts, nach0 identifies biological targets, analyzes mechanisms of action, generates molecular structures, proposes synthesis routes, and predicts molecular properties.
JAK3 inhibitor generation with Chemistry42: nach0 replaces 42 specialized generative models in Insilico Medicine’s Chemistry42 platform. In 45 minutes, nach0 generates 8 molecules satisfying all 2D and 3D requirements (hinge binding, active site binding), compared to a 0.04% discovery rate from a combinatorial generator over 24 hours. Chemistry42’s full pipeline (72 hours) still produces better structures since it uses reinforcement learning feedback and explicit structural constraints.
Comparison with ChatGPT
On a subset evaluation, fine-tuned nach0 base outperforms GPT-3.5-turbo on all tested tasks: EBM PICO (F1: 67.6% vs. 64.4%), MedMCQA-Open (BLEU-2: 6.3% vs. 1.7%), and molecular description generation (BLEU-2: 42.8% vs. 2.2%).
Competitive Multi-Task Performance with Clear Limitations
nach0 demonstrates that a single encoder-decoder model can achieve competitive results across both chemical and NLP tasks when pre-trained on mixed-modality data and fine-tuned with instruction tuning. The model’s strongest advantages appear on chemistry tasks (reaction prediction, property prediction, molecular generation), where specialized SMILES tokenization and chemical pre-training provide clear benefits over general-purpose models of similar scale.
Limitations Acknowledged by the Authors
Not at chemist expert level: Human evaluations indicate the model does not match domain expert performance. Key gaps include chemical reasoning, knowledge alignment with domain-specific knowledge graphs, and the ability to learn from expert feedback.
SMILES-only molecular representation: The model lacks 3D geometric information. SMILES notation is not one-to-one with molecular structures, and the model does not incorporate molecular graphs or 3D coordinates. The authors suggest SELFIES as a potential alternative representation.
Prompt sensitivity: Performance depends on prompt quality and specificity. Over-reliance on domain-specific prompts may limit response diversity.
Limited chemical diversity: Cross-domain datasets from Mol-Instructions primarily cover known drugs and chemical probes from PubChem, representing only a fraction of predicted chemical space.
Future Directions
The authors propose extending nach0 with protein sequence modalities (using Group SELFIES), expanding zero-shot evaluation capabilities, and integrating knowledge graph information through self-supervised approaches.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Pre-training (text) | PubMed abstracts | 13M docs, 355M tokens | Filtered for chemistry-related content |
| Pre-training (text) | USPTO patents | 119K docs, 2.9B tokens | Patent descriptions |
| Pre-training (chemical) | ZINC | ~100M docs, 4.7B tokens | Molecular SMILES strings |
| Fine-tuning (NLP) | 17 NLP datasets | Varies | See Table 1 in paper |
| Fine-tuning (chemistry) | MoleculeNet, MOSES, Mol-Instructions | Varies | Predefined or random splits |
Algorithms
- Architecture: T5 encoder-decoder (base: 250M, large: 780M parameters)
- Pre-training objective: Language modeling (masked span prediction)
- Fine-tuning: Multi-task instruction tuning with examples-proportional mixing
- Hyperparameters: batch size 1024, learning rate $1 \times 10^{-4}$, weight decay 0.01
- Pre-training: 1 epoch; fine-tuning: 10 epochs
Models
| Artifact | Type | License | Notes |
|---|---|---|---|
| nach0 Base (HuggingFace) | Model | CC-BY-NC-4.0 | 250M parameter encoder-decoder |
| nach0 Large (HuggingFace) | Model | CC-BY-NC-4.0 | 780M parameter encoder-decoder |
| nach0 GitHub Repository | Code | Not specified | Training and inference code |
Evaluation
Evaluation spans 17+ NLP benchmarks and 10+ chemistry benchmarks. Metrics include F1 (NER, RE, classification), accuracy (QA, entailment, reaction prediction), balanced accuracy (molecular property classification), R2/RMSE (regression), BLEU-2 (generation), and FCD/SNN/validity/novelty (molecular generation via MOSES).
Hardware
- Base models: NVIDIA A4000 and A5000 GPUs
- Large models: NVIDIA DGX cloud platform
- Training used tensor and pipeline parallelism via NeMo toolkit
- Specific GPU counts and training times not reported
Paper Information
Citation: Livne, M., Miftahutdinov, Z., Tutubalina, E., Kuznetsov, M., Polykovskiy, D., Brundyn, A., Jhunjhunwala, A., Costa, A., Aliper, A., Aspuru-Guzik, A., & Zhavoronkov, A. (2024). nach0: Multimodal Natural and Chemical Languages Foundation Model. Chemical Science, 15(22), 8380-8389. https://doi.org/10.1039/D4SC00966E
@article{livne2024nach0,
title={nach0: multimodal natural and chemical languages foundation model},
author={Livne, Micha and Miftahutdinov, Zulfat and Tutubalina, Elena and Kuznetsov, Maksim and Polykovskiy, Daniil and Brundyn, Annika and Jhunjhunwala, Aastha and Costa, Anthony and Aliper, Alex and Aspuru-Guzik, Al{\'a}n and Zhavoronkov, Alex},
journal={Chemical Science},
volume={15},
number={22},
pages={8380--8389},
year={2024},
publisher={Royal Society of Chemistry},
doi={10.1039/D4SC00966E}
}