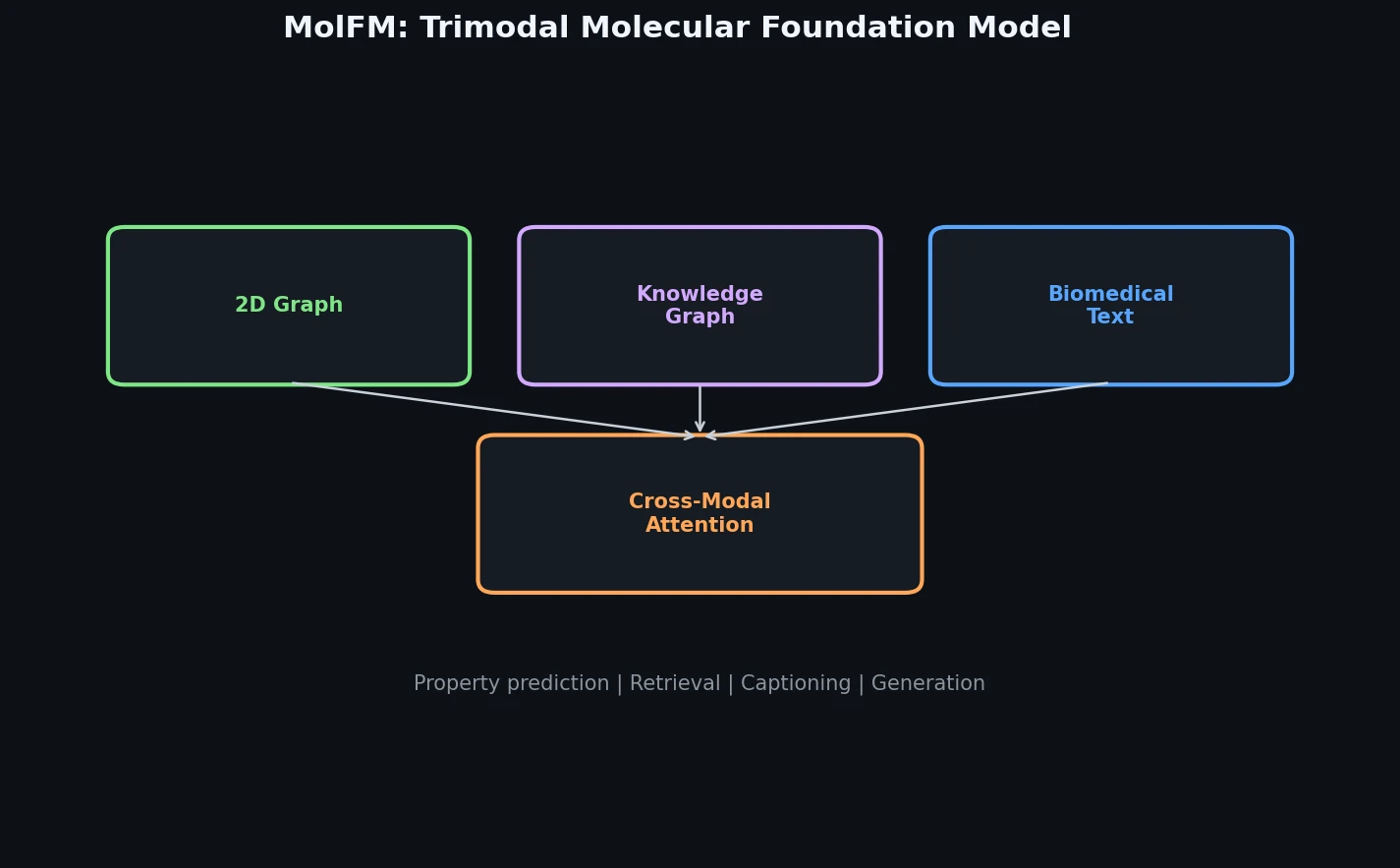
Trimodal Pre-training for Molecular Understanding
MolFM is a Method paper that introduces a multimodal molecular foundation model integrating three distinct sources of molecular knowledge: 2D molecular graphs, biomedical text, and knowledge graphs. The primary contribution is a pre-training framework that uses fine-grained cross-modal attention to fuse information across all three modalities, combined with theoretical justification from a deep metric learning perspective. MolFM achieves the best reported results (at time of publication) on cross-modal retrieval, molecule captioning, text-based molecule generation, and molecular property prediction.
Why Existing Molecular Models Fall Short
Prior multimodal molecular foundation models operate on at most two modalities (structures and text) and suffer from two key limitations. First, generative approaches like KV-PLM and MolT5 rely on 1D SMILES strings, which cannot capture complex topological and spatial molecular properties such as macrocycles. Contrastive approaches like MoMu and MoleculeSTM learn global alignment between molecule graphs and text but overlook fine-grained connections between specific substructures and textual descriptions.
Second, and more fundamentally, no prior model incorporates knowledge graphs as a third modality. Knowledge graphs encode global-level relationships among molecules, target ligands, diseases, and other biomedical entities. These relationships capture functional and structural similarity patterns that cannot be learned from individual molecule-text pairs alone. MolFM addresses both gaps by introducing cross-modal attention across all three modalities and providing theoretical guarantees about what the pre-training objectives learn.
Cross-Modal Attention and Metric Learning Guarantees
Architecture
MolFM uses three pre-trained single-modal encoders:
- Molecular graph encoder: A 5-layer GIN (1.8M parameters) initialized from GraphMVP, producing atom-level features $h_{SA}$ and a graph-level feature $h_{SM}$
- Text encoder: A 6-layer transformer (61.8M parameters) initialized from KV-PLM’s first 6 layers, producing token features $h_T$
- Knowledge graph encoder: A TransE model (12.6M parameters) trained on the knowledge graph for 500 epochs, producing entity features $h_K$
A multimodal encoder (61.8M parameters, 6 transformer layers with cross-attention) fuses the three modalities. The cross-attention uses text token features as queries and the concatenation of atom features and knowledge graph neighbor features as keys and values. For each molecule, the knowledge graph input is the molecule’s entity and $N=4$ randomly sampled one-hop neighbors.
Pre-training Objectives
MolFM combines four losses:
Structure-text contrastive (STC) aligns the global feature spaces of structure and text encoders using a symmetric InfoNCE loss:
$$\mathcal{L}_{stc} = -\frac{1}{2} \left[ \log \frac{\exp(s(z_S, z_T) / \tau)}{\sum_{S’ \in B} \exp(s(z_{S’}, z_T) / \tau)} + \log \frac{\exp(s(z_S, z_T) / \tau)}{\sum_{T’ \in B} \exp(s(z_S, z_{T’}) / \tau)} \right]$$
where $s(\cdot, \cdot)$ is cosine similarity and $\tau = 0.1$ is a temperature parameter.
Cross-modal matching (CMM) predicts whether a structure-text-knowledge triplet corresponds to the same molecule, using cross-entropy over the multimodal encoder’s CLS token:
$$\mathcal{L}_{cmm} = \sum_{(\tilde{S}, \tilde{T}, \tilde{K}) \in \tilde{B}} H\left[y_{cmm}(\tilde{S}, \tilde{T}, \tilde{K}),; p_{cmm}\left(\mathcal{M}_\theta(h_{\tilde{S}}, h_{\tilde{T}}, h_{\tilde{K}})\right)\right]$$
Masked language modeling (MLM) predicts masked text tokens conditioned on all three modalities:
$$\mathcal{L}_{mlm} = H\left[y_{mlm}(\hat{T}),; p_{mlm}\left(\mathcal{M}_\theta(h_S, h_{\hat{T}}, h_K)\right)\right]$$
Knowledge graph embedding (KGE) regularizes entity embeddings with a max-margin TransE loss:
$$\mathcal{L}_{kge} = \sum_{h \in K} \left[\max(0, d(h,r,t) - d(h,r,\tilde{t}) + \Delta) + \max(0, d(h,r,t) - d(\tilde{h},r,t) + \Delta)\right]$$
where $d(h,r,t) = | f(h) + g(r) - f(t) |_2$ and $\Delta = 0.2$.
The total pre-training loss is:
$$\mathcal{L} = \mathbb{E}_{(S,T,K)}\left[\mathcal{L}_{stc} + \mathcal{L}_{cmm} + \mathcal{L}_{mlm} + \mathcal{L}_{kge}\right]$$
Theoretical Justifications
The authors provide metric learning interpretations for each objective. For CMM, they show that the loss is proportional to assigning higher scores to matched triplets and lower scores to unmatched ones, aligning the feature space across all three modalities.
For KGE, two lemmas provide guarantees about structurally and functionally similar molecules:
Lemma 1 (Structural similarity): For a symmetric structural-similarity relation $r_s$, the KGE loss satisfies:
$$\mathcal{L}_{kge}(h, r_s, t) \propto 2|f(h) - f(t)| - \mathbb{E}_{\tilde{t}}|f(h) - f(\tilde{t})| - \mathbb{E}_{\tilde{h}}|f(\tilde{h}) - f(t)|$$
This shows KGE pulls structurally similar molecules closer while pushing dissimilar ones apart.
Lemma 2 (Functional similarity): For molecules $h$ and $t$ that interact with a common entity $o$, the distance between their embeddings is upper-bounded:
$$|f(h) - f(t)| \leq \alpha,\mathbb{E}_{(e_1, r, e_2) \sim \mathcal{I}}\left[\mathcal{L}_{kge}(e_1, r, e_2)\right] + C$$
where $\alpha \approx 1$ and $C \approx 0$. This guarantees that minimizing KGE also brings functionally similar molecules closer in the embedding space.
Experiments Across Four Downstream Tasks
Pre-training Data
MolFM pre-trains on 15K molecules from PubChem paired with 37M paragraphs from S2ORC. The knowledge graph contains 49K entities and 3.2M relations, constructed from DrugBank, BindingDB, and additional public databases with heuristic augmentation.
Cross-Modal Retrieval
Evaluated on PCdes (paragraph-level) in zero-shot and fine-tuning settings. MolFM uses a re-ranking strategy that linearly combines cosine similarity with CMM logits over the top-$k$ retrieved candidates.
| Mode | Model | S-T MRR | S-T R@1 | S-T R@10 | T-S MRR | T-S R@1 | T-S R@10 |
|---|---|---|---|---|---|---|---|
| Zero-shot | MoMu | 9.89 | 5.08 | 18.93 | 10.33 | 4.90 | 20.69 |
| Zero-shot | MolFM | 21.42 | 13.90 | 36.21 | 23.63 | 16.14 | 39.54 |
| Fine-tune | MoMu | 34.29 | 24.47 | 53.84 | 34.53 | 24.87 | 54.25 |
| Fine-tune | MolFM | 39.56 | 29.76 | 58.63 | 39.34 | 29.39 | 58.49 |
MolFM achieves 12.13% and 5.04% absolute gains over MoMu under zero-shot and fine-tuning settings, respectively.
Molecule Captioning
Evaluated on ChEBI-20 using MolT5 decoders. MolFM’s structure encoder features are concatenated with the MolT5 encoder outputs.
| Decoder | Encoder | BLEU-4 | ROUGE-L | METEOR | Text2Mol |
|---|---|---|---|---|---|
| MolT5-base | MolT5-base | 0.457 | 0.578 | 0.569 | 0.547 |
| MolT5-base | MoMu | 0.462 | 0.575 | 0.576 | 0.558 |
| MolT5-base | GraphMVP | 0.491 | 0.592 | 0.599 | 0.570 |
| MolT5-base | MolFM | 0.498 | 0.594 | 0.607 | 0.576 |
Text-Based Molecule Generation
Also on ChEBI-20 with MolT5 decoders. MolFM’s text features are projected and fed to the decoder.
| Decoder | Encoder | Exact | Valid | Morgan FTS | Text2Mol |
|---|---|---|---|---|---|
| MolT5-base | MolT5-base | 0.082 | 0.786 | 0.601 | 0.543 |
| MolT5-base | MoMu | 0.183 | 0.863 | 0.678 | 0.580 |
| MolT5-base | MolFM | 0.210 | 0.892 | 0.697 | 0.583 |
Molecular Property Prediction
On MoleculeNet (8 classification datasets), MolFM concatenates the structure feature and the multimodal encoder’s CLS feature to predict properties.
| Model | BBBP | Tox21 | ClinTox | HIV | BACE | Avg |
|---|---|---|---|---|---|---|
| GraphMVP | 72.4 | 74.4 | 77.5 | 77.0 | 81.2 | 73.07 |
| DeepEIK | 72.1 | 72.4 | 89.7 | 75.0 | 80.5 | 73.27 |
| MolFM (w/o T+K) | 72.2 | 76.6 | 78.6 | 78.2 | 82.6 | 73.95 |
| MolFM (w/ T+K) | 72.9 | 77.2 | 79.7 | 78.8 | 83.9 | 74.62 |
With multimodal inputs, MolFM averages 74.62% ROC-AUC, a 1.55% absolute gain over GraphMVP.
Ablation Studies
Zero-shot retrieval ablations reveal that cross-modal attention to atoms and CMM are the most critical components. Removing either causes a sharp drop (approximately 3% on S-T retrieval). Knowledge graph incorporation yields a 1.5% average improvement, with both attention to neighbors and KGE contributing marginally.
Key Findings and Limitations
MolFM demonstrates that incorporating knowledge graphs as a third modality provides consistent improvements across all evaluated tasks. The theoretical analysis connecting pre-training objectives to deep metric learning provides interpretability for why the model works: STC and CMM align representations of the same molecule across modalities, while KGE pulls structurally and functionally similar molecules closer in the embedding space.
The cross-modal attention visualizations show that MolFM learns to associate specific atom substructures with relevant text tokens and knowledge graph entities. For example, the model correctly attends to functional groups mentioned in textual descriptions.
The authors acknowledge several limitations:
- Data quality: The pre-training dataset (15K molecules) is small and may introduce biases
- Cold-start problem: MolFM provides limited benefit for newly emerged molecules lacking text and knowledge graph information
- Entity scope: The model focuses on molecules and does not incorporate proteins, genes, or cell lines, which could further improve biomedical understanding
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Pre-training (molecules) | PubChem | 15K molecules | Follows MoMu’s pre-training data |
| Pre-training (text) | S2ORC | 37M paragraphs | Biomedical literature paragraphs |
| Knowledge graph | DrugBank, BindingDB, public DBs | 49K entities, 3.2M relations | Constructed with heuristics from MoCL |
| Cross-modal retrieval | PCdes | Paragraph-level | Test split |
| Captioning/Generation | ChEBI-20 | - | Following MolT5 splits |
| Property prediction | MoleculeNet | 8 datasets | Classification tasks, ROC-AUC metric |
Algorithms
- Optimizer: AdamW with weight decay $1 \times 10^{-4}$
- Learning rate: linear warmup to $1 \times 10^{-4}$ over 2,000 iterations, cosine annealing to $1 \times 10^{-5}$
- Batch size: 128
- Pre-training epochs: 300
- Knowledge graph neighbors per molecule: $N = 4$
- Temperature: $\tau = 0.1$
- Margin: $\Delta = 0.2$
Models
| Component | Architecture | Parameters | Initialization |
|---|---|---|---|
| Graph encoder | 5-layer GIN | 1.8M | GraphMVP |
| Text encoder | 6-layer Transformer | 61.8M | KV-PLM (first 6 layers) |
| Knowledge encoder | TransE | 12.6M | Trained 500 epochs on KG |
| Multimodal encoder | 6-layer Transformer + cross-attention | 61.8M | KV-PLM (last 6 layers) |
| Total | ~138M |
Evaluation
| Task | Metrics |
|---|---|
| Cross-modal retrieval | MRR, Recall@1/5/10 |
| Molecule captioning | BLEU-2/4, ROUGE-1/2/L, METEOR, Text2Mol |
| Text-to-molecule generation | BLEU, Exact ratio, Validity, Levenshtein, Fingerprint Tanimoto (MACCS/RDKit/Morgan), Text2Mol |
| Property prediction | ROC-AUC per dataset |
Hardware
- 4 NVIDIA A100 GPUs for pre-training
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| OpenBioMed | Code | MIT | Official implementation including MolFM |
Paper Information
Citation: Luo, Y., Yang, K., Hong, M., Liu, X. Y., & Nie, Z. (2023). MolFM: A Multimodal Molecular Foundation Model. arXiv preprint arXiv:2307.09484.
@article{luo2023molfm,
title={MolFM: A Multimodal Molecular Foundation Model},
author={Luo, Yizhen and Yang, Kai and Hong, Massimo and Liu, Xing Yi and Nie, Zaiqing},
journal={arXiv preprint arXiv:2307.09484},
year={2023}
}