A Dual-Branch Pre-training Method for Molecular Property Prediction
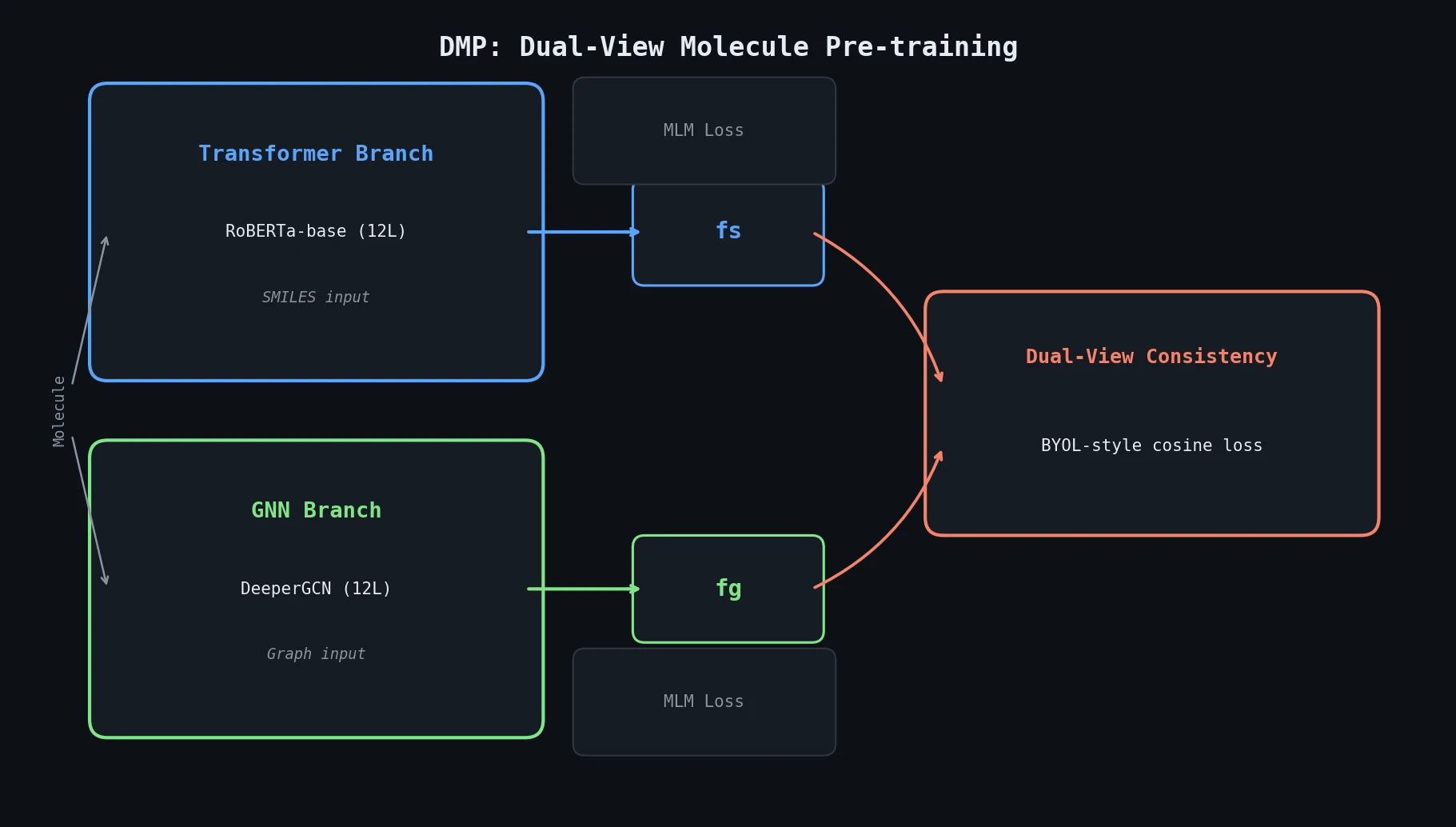
DMP (Dual-view Molecule Pre-training) is a Method paper that introduces a pre-training framework combining two complementary molecular encoders: a Transformer operating on SMILES strings and a Graph Neural Network (GNN) operating on molecular graphs. The two branches are trained jointly with masked language modeling (MLM) objectives plus a BYOL-style dual-view consistency loss. After pre-training on 10M PubChem molecules, either branch (or both) can be fine-tuned for downstream tasks. The authors recommend the Transformer branch based on empirical results. DMP achieves the best reported performance on 7 of 9 MoleculeNet classification tasks and 3 retrosynthesis benchmarks (at the time of the 2021 arXiv version).
Why Combine SMILES and Graph Views for Molecules
Prior molecule pre-training methods used either graph representations with GNNs or SMILES representations with Transformers, but not both. The authors observe that the two views are complementary: Transformers handle molecules with large atom distances (long chains) well, while GNNs handle molecules with many concatenated rings better. Neither model alone captures the full range of molecular structures effectively.
Existing GNN-based pre-training methods (Hu et al. 2020, MolCLR, GROVER) and SMILES-based methods (ChemBERTa, SMILES-BERT) each have blind spots dictated by their input representation. DMP addresses this by pre-training both views simultaneously and enforcing representation consistency between them, so each branch benefits from the structural knowledge of the other.
Dual-View Consistency with BYOL-Style Training
The core innovation is the dual-view consistency objective, inspired by Bootstrap Your Own Latent (BYOL). Given a molecule $M$ with SMILES representation $M_s$ and graph representation $M_g$, DMP obtains high-level features from each branch:
- Transformer branch: A RoBERTa-base model encodes the SMILES sequence. The [CLS] token output serves as the molecule representation $f_s$.
- GNN branch: A DeeperGCN network encodes the molecular graph. Mean+max pooling over atom representations yields $f_g$.
The dual-view consistency loss uses nonlinear projection heads $\psi_g, \psi_s$ and prediction heads $\rho_g, \rho_s$:
$$ p_g = \psi_g(f_g), \quad q_g = \rho_g(p_g); \quad p_s = \psi_s(f_s), \quad q_s = \rho_s(p_s) $$
The consistency loss maximizes cross-view cosine similarity with stop-gradient (SG) on the target:
$$ \ell_{\text{dual}}(\tilde{M}_g, \tilde{M}_s) = -\cos(q_s, \text{SG}(p_g)) - \cos(q_g, \text{SG}(p_s)) $$
where $\cos(p, q) = \frac{p^\top q}{|p|_2 |q|_2}$ and $\tilde{M}_g, \tilde{M}_s$ are the masked versions of the inputs. The stop-gradient prevents representation collapse without requiring negative samples or a momentum encoder.
The full training objective combines three losses:
- MLM on Transformer: Recover masked tokens in SMILES sequences
- MLM on GNN: Recover masked atoms in molecular graphs
- Dual-view consistency: The BYOL-style loss above
Both MLM objectives and the consistency loss are necessary. Ablations show that removing MLM (using only dual-view loss) degrades performance, and using two branches of the same type (two Transformers or two GNNs) is less effective than the heterogeneous Transformer+GNN combination.
Experiments on MoleculeNet and Retrosynthesis
Pre-training Setup
DMP is pre-trained on 10M molecules from PubChem (matching prior work). The Transformer branch uses RoBERTa-base (12 layers, hidden dim 768, 87M parameters). The GNN branch uses DeeperGCN (12 layers, hidden dim 384, 7.4M parameters). Combined, DMP has 104.1M parameters. Training runs for 200K iterations on 8 V100 GPUs over 3.8 days with Adam optimizer (lr = 5e-4, weight decay 0.01).
Molecular Property Prediction (MoleculeNet)
DMP is evaluated on 6 binary classification tasks (BBBP, Tox21, ClinTox, HIV, BACE, SIDER) using official DeepChem splits, and on 3 additional tasks (BBBP, SIDER, ClinTox classification + ESOL, QM7, QM8 regression) using scaffold splits from GROVER.
Key results on DeepChem splits (ROC-AUC %):
| Dataset | MolCLR | TF (MLM) | DMP_TF | DMP_TF+GNN |
|---|---|---|---|---|
| BBBP | 73.6 | 74.9 | 78.1 | 77.8 |
| Tox21 | 79.8 | 77.6 | 78.8 | 79.1 |
| ClinTox | 93.2 | 92.9 | 95.0 | 95.6 |
| HIV | 80.6 | 80.2 | 81.0 | 81.4 |
| BACE | 89.0 | 88.0 | 89.3 | 89.4 |
| SIDER | 68.0 | 68.4 | 69.2 | 69.8 |
On scaffold splits (comparison with GROVER and MPG):
| Dataset | GROVER | MPG | DMP_TF |
|---|---|---|---|
| BBBP (AUC) | 0.940 | 0.922 | 0.945 |
| SIDER (AUC) | 0.658 | 0.661 | 0.695 |
| ClinTox (AUC) | 0.944 | 0.963 | 0.968 |
| ESOL (RMSE) | 0.831 | 0.741 | 0.700 |
| QM7 (MAE) | 72.6 | - | 69.6 |
| QM8 (MAE) | 0.0125 | - | 0.0124 |
Retrosynthesis
DMP is tested on USPTO-50K (reaction type known/unknown) and USPTO-full. Using a “DMP fusion” approach (fusing pre-trained representations into a Transformer encoder-decoder for retrosynthesis), DMP improves top-1 accuracy by 2-3 points over the baseline Transformer across all settings:
| Setting | Transformer | ChemBERTa fusion | DMP fusion |
|---|---|---|---|
| USPTO-50K (unknown) | 42.3 | 43.9 | 46.1 |
| USPTO-50K (known) | 54.2 | 56.4 | 57.5 |
| USPTO-full | 42.9 | - | 45.0 |
For GNN-based retrosynthesis, replacing GLN’s GNN modules with DMP’s pre-trained GNN branch improves top-1 accuracy from 52.5% to 54.2% (unknown type) and from 64.2% to 66.5% (known type).
Representation Quality
t-SNE visualization of pre-trained representations shows that DMP produces better scaffold-based clustering than either GNN-only or Transformer-only pre-training. The Davies-Bouldin index improves from 3.56 (GNN) and 3.59 (Transformer) to 2.19 (DMP), indicating much tighter within-scaffold clusters.
Key Findings and Limitations
Key findings:
- Combining heterogeneous views (SMILES + graph) during pre-training is more effective than using two branches of the same type. TF(x2) and GNN(x2) variants show smaller gains.
- Both MLM and dual-view consistency loss contribute. Removing MLM (dual-view only) hurts performance, especially on BBBP (71.1 vs 78.1 with both losses).
- The Transformer branch alone is recommended for downstream tasks, as it achieves strong results without adding GNN parameters at inference time.
- Scaling pre-training data from 10M to 100M compounds yields marginal additional improvement.
Limitations acknowledged by the authors:
- Training cost is higher than single-branch methods (3.8 days vs 2.5 days for TF-only on 8 V100s), since both branches must be trained jointly.
- A fixed branch selection strategy is used at inference time. The authors note that a meta-controller for dynamic branch selection per molecule would be preferable.
- The GNN branch uses simple atom masking without bond deletion or subgraph removal, leaving room for stronger graph-level pre-training objectives.
Relation to co-training: The authors clarify that DMP differs from classical co-training (Blum and Mitchell 1998) in that it does not require conditional independence between views and produces a pre-trained model rather than additional labeled data.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Pre-training | PubChem subset | 10M compounds | Same subset as MolCLR and ChemBERTa |
| Pre-training (large) | PubChem subset | 100M compounds | Additional scale experiment |
| Evaluation (classification) | MoleculeNet (BBBP, Tox21, ClinTox, HIV, BACE, SIDER) | 1.5K-41K molecules | Official DeepChem splits |
| Evaluation (regression) | MoleculeNet (ESOL, QM7, QM8) | Varies | Scaffold splits from GROVER |
| Evaluation (retrosynthesis) | USPTO-50K, USPTO-full | 50K / 950K reactions | Splits from Dai et al. (2019) |
Algorithms
- Transformer branch: RoBERTa-base with MLM. SMILES tokenized using regex from Schwaller et al. (2019).
- GNN branch: DeeperGCN with 12 layers, atom masking for MLM.
- Dual-view loss: BYOL-style with 3-layer MLP projection heads and 2-layer MLP prediction heads, stop-gradient on targets.
- Optimizer: Adam (lr=5e-4, beta1=0.9, beta2=0.98, epsilon=1e-6), weight decay 0.01, 10K warmup steps, linear decay.
Models
| Component | Architecture | Parameters |
|---|---|---|
| Transformer branch | RoBERTa-base (12L, 768H, 12 heads) | 87M |
| GNN branch | DeeperGCN (12L, 384H) | 7.4M |
| DMP (total) | Transformer + GNN + projection/prediction heads | 104.1M |
Evaluation
- Classification: ROC-AUC, averaged over 3 random seeds
- Regression: RMSE (ESOL) or MAE (QM7, QM8)
- Retrosynthesis: Top-k exact match accuracy (k=1,3,5,10,20,50)
Hardware
- Pre-training: 8 NVIDIA V100 GPUs, batch size 12288 tokens, gradient accumulation 16x
- Pre-training time: 3.8 days (DMP), 2.5 days (TF-only), 1.7 days (GNN-only)
Artifacts
No public code repository or pre-trained model weights were identified for this paper. The paper references GLN’s code repository (https://github.com/Hanjun-Dai/GLN) for the retrosynthesis baseline but does not release DMP-specific code.
| Artifact | Type | License | Notes |
|---|---|---|---|
| GLN (baseline) | Code | MIT | Retrosynthesis baseline, not DMP code |
Paper Information
Citation: Zhu, J., Xia, Y., Wu, L., Xie, S., Zhou, W., Qin, T., Li, H., & Liu, T.-Y. (2023). Dual-view Molecular Pre-training. In Proceedings of the 29th ACM SIGKDD Conference on Knowledge Discovery and Data Mining (pp. 3615-3627). https://doi.org/10.1145/3580305.3599317
@inproceedings{zhu2023dualview,
title={Dual-view Molecular Pre-training},
author={Zhu, Jinhua and Xia, Yingce and Wu, Lijun and Xie, Shufang and Zhou, Wengang and Qin, Tao and Li, Houqiang and Liu, Tie-Yan},
booktitle={Proceedings of the 29th ACM SIGKDD Conference on Knowledge Discovery and Data Mining},
pages={3615--3627},
year={2023},
doi={10.1145/3580305.3599317}
}