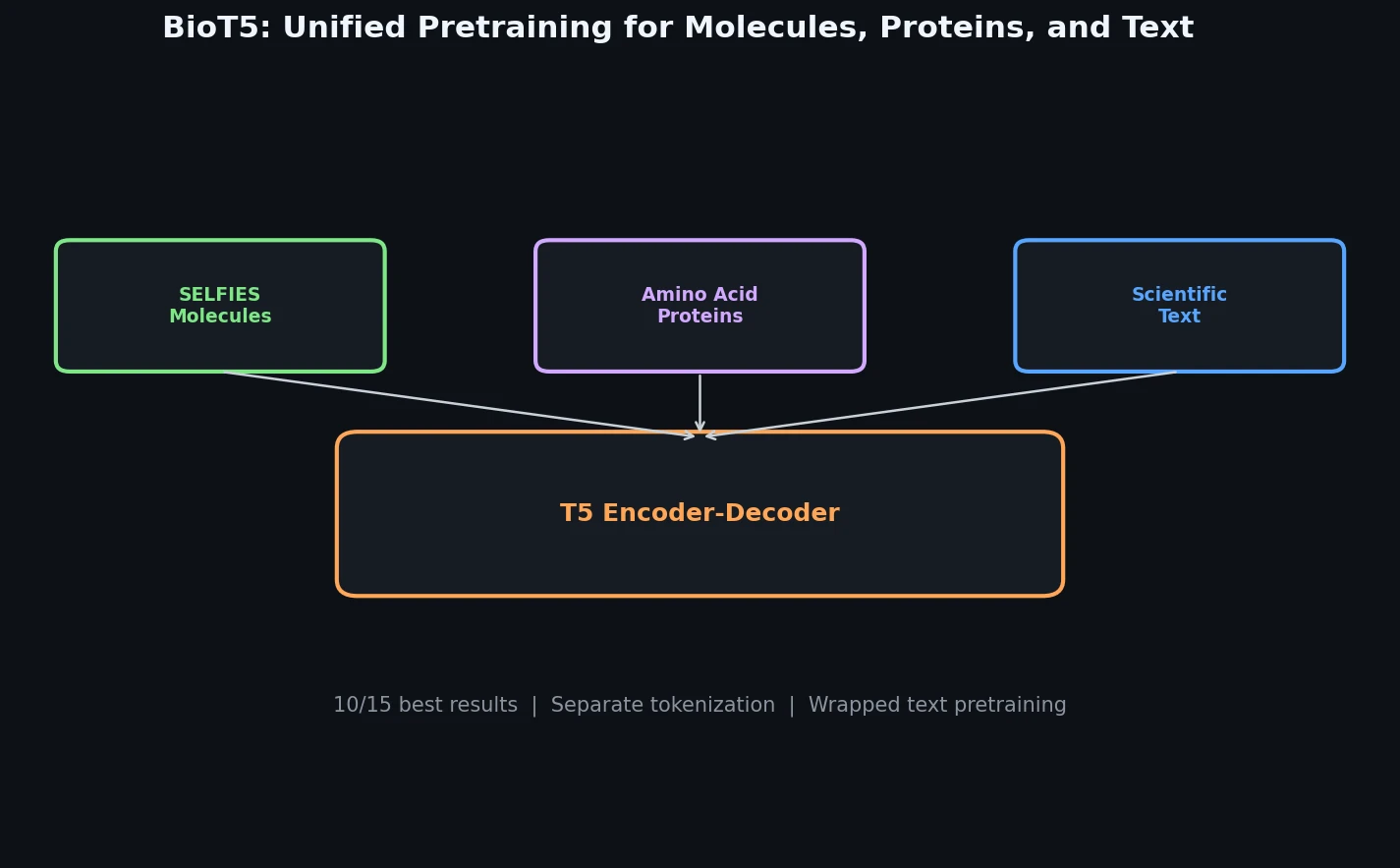
A Unified Pretraining Framework for Molecules, Proteins, and Text
BioT5 is a Method paper that introduces a comprehensive T5-based pretraining framework for cross-modal integration of molecules, proteins, and natural language. The primary contribution is a multi-task pretraining approach that uses SELFIES (instead of SMILES) for 100% valid molecular representations, separate tokenization for each modality, and a combination of masked language modeling and translation objectives to connect structured biological data with unstructured scientific text. After fine-tuning, BioT5 (252M parameters) achieves state-of-the-art performance on 10 out of 15 downstream tasks spanning molecule property prediction, protein property prediction, drug-target interaction, protein-protein interaction, molecule captioning, and text-based molecule generation.
Bridging the Gap Between Molecular Sequences and Scientific Knowledge
Prior cross-modal models in computational biology face three recurring challenges. First, models like MolT5 and MolXPT rely on SMILES to represent molecules, but SMILES strings are syntactically fragile: random perturbations or model-generated sequences frequently produce invalid molecular structures. Edwards et al. (2022) and Li et al. (2023) both highlight this validity problem as a bottleneck for text-to-molecule generation. Second, the contextual information surrounding molecular and protein names in scientific literature (e.g., mentions in PubMed abstracts that describe properties, interactions, and experimental results) remains underutilized. Most models either ignore this context or treat it identically to structured database entries. Third, existing approaches like MolT5 and Galactica share a single tokenizer and embedding space across molecules, proteins, and text. This leads to chemically incorrect tokenization: the bromine atom “Br” in SMILES gets split into “B” (boron) and “r”, producing erroneous downstream predictions.
BioT5 addresses all three issues simultaneously by adopting SELFIES for molecular representation, extracting entity-linked contextual knowledge from PubMed, and employing separate vocabularies for each modality.
SELFIES, Separate Tokenization, and Multi-Task Pretraining
The core innovations of BioT5 center on three design decisions:
SELFIES for Robust Molecular Representation
BioT5 replaces SMILES with SELFIES (Self-referencing Embedded Strings) for all molecular representations. Every permutation of symbols within the SELFIES alphabet generates a chemically valid molecular structure, guaranteeing 100% validity in generation tasks. Molecules from ZINC20 are converted from SMILES to SELFIES during data preprocessing.
Modality-Specific Tokenization
Rather than sharing a single SentencePiece vocabulary across modalities, BioT5 maintains three separate dictionaries:
- Molecules: Each SELFIES token corresponds to a chemically meaningful atom group enclosed in brackets (e.g.,
[C],[=C],[Br]). - Proteins: Amino acids are prefixed with a special
<p>token to distinguish them from text characters (e.g.,<p>M,<p>K,<p>R). - Text: The standard T5 vocabulary is retained.
This prevents semantic conflation across modalities. The total vocabulary size is 35,073, and the model comprises 252M parameters using the T5-v1.1-base architecture.
Multi-Task Pretraining Objectives
BioT5 uses six pretraining tasks organized into three categories:
- Single-modal T5 objective: Standard span corruption and recovery applied independently to molecule SELFIES (task 1), protein FASTA (task 2), and general text from C4 (task 3).
- Wrapped text T5 objective (task 4): Applied to PubMed articles where molecular names are replaced with corresponding SELFIES strings and gene names are appended with protein FASTA sequences, using BERN2 for named entity recognition and entity linking.
- Bidirectional translation (tasks 5 and 6): Molecule SELFIES to text description and vice versa (using 339K pairs from PubChem), and protein FASTA to text description and vice versa (using 569K pairs from Swiss-Prot).
The translation direction is randomly sampled with probability 0.5 for each example. For downstream tasks, BioT5 uses prompt-based fine-tuning to cast all tasks into a sequence generation format, reducing the gap between pretraining and fine-tuning.
Evaluation Across 15 Downstream Tasks
BioT5 is evaluated on 15 tasks organized into three categories: single-instance prediction, multi-instance prediction, and cross-modal generation.
Molecule Property Prediction (MoleculeNet)
BioT5 is evaluated on six binary classification tasks from MoleculeNet using scaffold splitting: BBBP, Tox21, ClinTox, HIV, BACE, and SIDER. Results are averaged over three random runs.
| Dataset | GEM | MolXPT | BioT5 |
|---|---|---|---|
| BBBP | 72.4 | 80.0 | 77.7 |
| Tox21 | 78.1 | 77.1 | 77.9 |
| ClinTox | 90.1 | 95.3 | 95.4 |
| HIV | 80.6 | 78.1 | 81.0 |
| BACE | 85.6 | 88.4 | 89.4 |
| SIDER | 67.2 | 71.7 | 73.2 |
| Avg | 79.0 | 81.9 | 82.4 |
BioT5 achieves the best average AUROC (82.4) across all six datasets, surpassing both GNN-based methods (GEM) and language model baselines (MolXPT).
Protein Property Prediction (PEER Benchmark)
On the PEER benchmark, BioT5 is evaluated on protein solubility and subcellular localization prediction:
| Model | Params | Solubility (Acc) | Localization (Acc) |
|---|---|---|---|
| ESM-1b | 652.4M | 70.23 | 92.40 |
| ProtBert | 419.9M | 68.15 | 91.32 |
| BioT5 | 252.1M | 74.65 | 91.69 |
BioT5 achieves the best solubility prediction accuracy (74.65%) despite being 2-3x smaller than dedicated protein language models like ESM-1b and ProtBert.
Drug-Target Interaction Prediction
BioT5 is evaluated on three DTI datasets (BioSNAP, Human, BindingDB) with five random runs:
| Method | BioSNAP AUROC | Human AUROC | BindingDB AUROC |
|---|---|---|---|
| DrugBAN | 0.903 | 0.982 | 0.960 |
| BioT5 | 0.937 | 0.989 | 0.963 |
BioT5 consistently outperforms DrugBAN and other specialized DTI models across all three datasets.
Molecule Captioning and Text-Based Molecule Generation
On the ChEBI-20 dataset, BioT5 outperforms all baselines in molecule captioning:
| Model | Params | BLEU-4 | METEOR | Text2Mol |
|---|---|---|---|---|
| MolT5-large | 783M | 0.508 | 0.614 | 0.582 |
| MolXPT | 350M | 0.505 | 0.626 | 0.594 |
| BioT5 | 252M | 0.556 | 0.656 | 0.603 |
For text-based molecule generation, BioT5 achieves an exact match score of 0.413 (vs. 0.311 for MolT5-large) while maintaining 100% validity, compared to 90.5% for MolT5-large. This demonstrates the direct benefit of SELFIES: every generated sequence is a valid molecule.
Protein-Protein Interaction Prediction
On the PEER PPI benchmarks (Yeast and Human), BioT5 achieves competitive results, outperforming fully fine-tuned ProtBert and ESM-1b on the Yeast dataset (64.89% vs. 63.72% for ProtBert) and placing second on Human (86.22% vs. 88.06% for ESM-1b with frozen weights).
Key Findings, Limitations, and Future Directions
BioT5 demonstrates that integrating molecular, protein, and textual modalities within a single pretraining framework yields consistent improvements across diverse biological tasks. Three factors drive BioT5’s performance: (1) SELFIES guarantees 100% molecular validity in generation tasks, eliminating a persistent failure mode of SMILES-based models; (2) separate tokenization preserves the semantic integrity of each modality; (3) wrapped text pretraining on PubMed provides contextual biological knowledge that pure sequence models miss.
The authors acknowledge several limitations. BioT5 requires full-parameter fine-tuning for each downstream task because instruction-tuning does not generalize across tasks, and combining datasets via instructions causes data leakage (the authors note overlaps between BindingDB training data and BioSNAP/Human test sets). The model only handles sequence-format bio-entities and does not incorporate 2D or 3D structural information. Additional biological modalities such as DNA/RNA sequences and cell-level data are also left for future work.
The authors also note risks: BioT5 could potentially be misused to generate dangerous molecules, and it may fail to generate effective therapeutic molecules or produce compounds with adverse side effects.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Pretraining (molecules) | ZINC20 | ~300M molecules | Converted from SMILES to SELFIES |
| Pretraining (proteins) | UniRef50 | 27M proteins | Filtered by length |
| Pretraining (text) | C4 | Large | Standard T5 corpus |
| Pretraining (wrapped text) | PubMed | 33M articles | Entity linking via BERN2 |
| Pretraining (molecule-text pairs) | PubChem | 339K pairs | Excludes ChEBI-20 molecules |
| Pretraining (protein-text pairs) | Swiss-Prot | 569K pairs | High-quality annotations |
| Evaluation (molecular properties) | MoleculeNet | 6 datasets | Scaffold splitting |
| Evaluation (protein properties) | PEER | 2 tasks | Solubility and localization |
| Evaluation (DTI) | BioSNAP, Human, BindingDB | 3 datasets | Binary classification |
| Evaluation (PPI) | Yeast, Human | 2 datasets | From PEER benchmark |
| Evaluation (generation) | ChEBI-20 | 33K pairs | Molecule captioning and text-to-molecule |
Algorithms
- Architecture: T5-v1.1-base (encoder-decoder transformer)
- Optimizer: AdamW with RMS scaling
- Learning rate: cosine annealing, base $1 \times 10^{-2}$, minimum $1 \times 10^{-5}$
- Warmup steps: 10,000
- Dropout: 0.0
- Maximum input length: 512 tokens
- Pretraining steps: 350K
- Batch size: 96 per GPU (6 data types per batch)
- Prompt-based fine-tuning for all downstream tasks
Models
| Model | Parameters | Vocabulary Size | Architecture |
|---|---|---|---|
| BioT5 | 252M | 35,073 | T5-v1.1-base |
Evaluation
- Molecule property prediction: AUROC on 6 MoleculeNet tasks (scaffold split, 3 runs)
- Protein property prediction: accuracy on PEER benchmark (3 runs)
- Drug-target interaction: AUROC, AUPRC, accuracy on 3 DTI datasets (5 runs)
- Protein-protein interaction: accuracy on 2 PPI datasets (3 runs)
- Molecule captioning: BLEU, ROUGE, METEOR, Text2Mol on ChEBI-20
- Text-based molecule generation: BLEU, exact match, fingerprint similarities, FCD, validity on ChEBI-20
Hardware
- 8x NVIDIA A100 80GB GPUs for pretraining
- Codebase: nanoT5
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| BioT5 Code | Code | MIT | Official implementation |
Paper Information
Citation: Pei, Q., Zhang, W., Zhu, J., Wu, K., Gao, K., Wu, L., Xia, Y., & Yan, R. (2023). BioT5: Enriching Cross-modal Integration in Biology with Chemical Knowledge and Natural Language Associations. Proceedings of the 2023 Conference on Empirical Methods in Natural Language Processing, 1102-1123. https://doi.org/10.18653/v1/2023.emnlp-main.70
@inproceedings{pei2023biot5,
title={BioT5: Enriching Cross-modal Integration in Biology with Chemical Knowledge and Natural Language Associations},
author={Pei, Qizhi and Zhang, Wei and Zhu, Jinhua and Wu, Kehan and Gao, Kaiyuan and Wu, Lijun and Xia, Yingce and Yan, Rui},
booktitle={Proceedings of the 2023 Conference on Empirical Methods in Natural Language Processing},
pages={1102--1123},
year={2023},
publisher={Association for Computational Linguistics},
doi={10.18653/v1/2023.emnlp-main.70}
}