A Method for Target-Conditioned Molecular Generation
This is a Method paper that introduces TamGen (Target-aware molecular generation), a three-module architecture for generating drug-like compounds conditioned on protein binding pocket structures. The primary contribution is a GPT-like chemical language model pre-trained on 10 million SMILES from PubChem, combined with a Transformer-based protein encoder and a VAE-based contextual encoder for compound refinement. The authors validate TamGen on the CrossDocked2020 benchmark and apply it through a Design-Refine-Test pipeline to discover 14 novel inhibitors of the Mycobacterium tuberculosis ClpP protease, with $\text{IC}_{50}$ values ranging from 1.88 to 35.2 $\mu$M.
Bridging Generative AI and Practical Drug Discovery
Target-based generative drug design aims to create novel compounds with desired pharmacological properties from scratch, exploring the estimated $10^{60}$ feasible compounds in chemical space rather than screening existing libraries of $10^{4}$ to $10^{8}$ molecules. Prior approaches using diffusion models, GANs, VAEs, and autoregressive models have demonstrated the feasibility of generating compounds conditioned on target proteins. However, most generated compounds lack satisfactory physicochemical properties for drug-likeness, and validations with biophysical or biochemical assays are largely missing.
The key limitations of existing 3D generation methods (TargetDiff, Pocket2Mol, ResGen, 3D-AR) include:
- Generated compounds frequently contain multiple fused rings, leading to poor synthetic accessibility
- High cellular toxicity and decreased developability associated with excessive fused ring counts
- Slow generation speeds (tens of minutes to hours per 100 compounds)
- Limited real-world experimental validation of generated candidates
TamGen addresses these issues by operating in 1D SMILES space rather than 3D coordinate space, leveraging pre-training on natural compound distributions to produce more drug-like molecules.
Three-Module Architecture with Pre-Training and Refinement
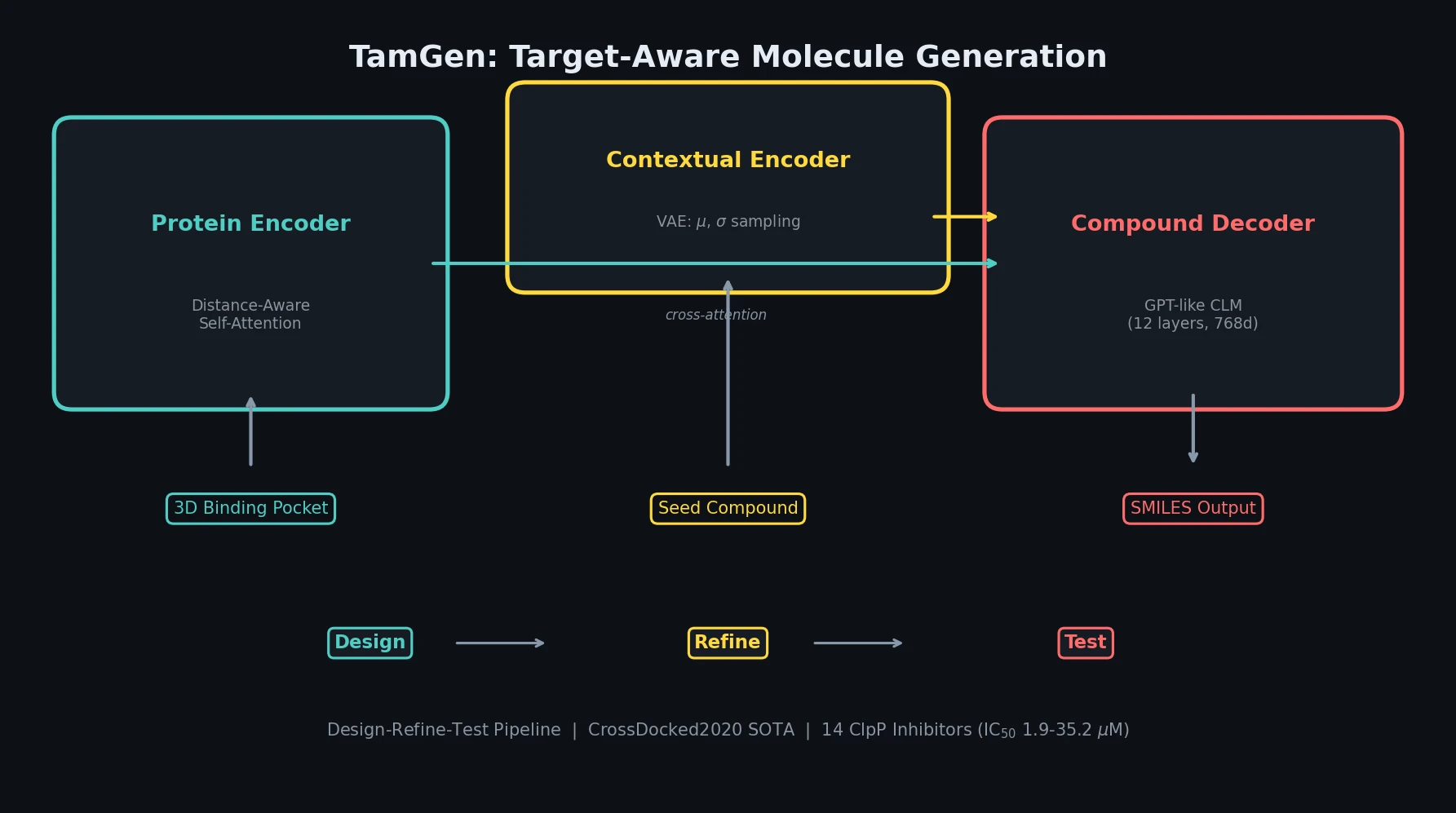
TamGen consists of three components: a compound decoder, a protein encoder, and a contextual encoder.
Compound Decoder (Chemical Language Model)
The compound decoder is a GPT-style autoregressive model pre-trained on 10 million SMILES randomly sampled from PubChem. The pre-training objective follows standard next-token prediction:
$$ \min -\sum_{y \in \mathcal{D}_0} \frac{1}{M_y} \sum_{i=1}^{M_y} \log P(y_i \mid y_{i-1}, y_{i-2}, \ldots, y_1) $$
where $M_y$ is the SMILES sequence length. This enables both unconditional and conditional generation. The decoder uses 12 Transformer layers with hidden dimension 768.
Protein Encoder with Distance-Aware Attention
The protein encoder processes binding pocket residues using both sequential and geometric information. Given amino acids $\mathbf{a} = (a_1, \ldots, a_N)$ with 3D coordinates $\mathbf{r} = (r_1, \ldots, r_N)$, the input representation combines amino acid embeddings with coordinate embeddings:
$$ h_i^{(0)} = E_a a_i + E_r \rho\left(r_i - \frac{1}{N}\sum_{j=1}^{N} r_j\right) $$
where $\rho$ denotes a random roto-translation operation applied as data augmentation, and coordinates are centered to the origin.
The encoder uses a distance-aware self-attention mechanism that weights attention scores by spatial proximity:
$$ \begin{aligned} \hat{\alpha}_j &= \exp\left(-\frac{|r_i - r_j|^2}{\tau}\right)(h_i^{(l)\top} W h_j^{(l)}) \\ \alpha_j &= \frac{\exp \hat{\alpha}_j}{\sum_{k=1}^{N} \exp \hat{\alpha}_k} \\ \hat{\boldsymbol{h}}_i^{(l+1)} &= \sum_{j=1}^{N} \alpha_j (W_v h_j^{(l)}) \end{aligned} $$
where $\tau$ is a temperature hyperparameter and $W$, $W_v$ are learnable parameters. The encoder uses 4 layers with hidden dimension 256. Outputs are passed to the compound decoder via cross-attention.
VAE-Based Contextual Encoder
A VAE-based contextual encoder determines the mean $\mu$ and standard deviation $\sigma$ for any (compound, protein) pair. During training, the model recovers the input compound. During application, a seed compound enables compound refinement. The full training objective combines reconstruction loss with KL regularization:
$$ \min_{\Theta, q} \frac{1}{|\mathcal{D}|} \sum_{(\mathbf{x}, \mathbf{y}) \in \mathcal{D}} -\log P(\mathbf{y} \mid \mathbf{x}, z; \Theta) + \beta \mathcal{D}_{\text{KL}}(q(z \mid \mathbf{x}, \mathbf{y}) | p(z)) $$
where $\beta$ is a hyperparameter controlling the KL divergence weight, and $p(z)$ is a standard Gaussian prior.
Benchmark Evaluation and Tuberculosis Drug Discovery
CrossDocked2020 Benchmark
TamGen was evaluated against five baselines (liGAN, 3D-AR, Pocket2Mol, ResGen, TargetDiff) on the CrossDocked2020 dataset (~100k drug-target pairs for training, 100 test binding pockets). For each target, 100 compounds were generated per method. Evaluation metrics included:
- Docking score (AutoDock-Vina): binding affinity estimate
- QED: quantitative estimate of drug-likeness
- Lipinski’s Rule of Five: physicochemical property compliance
- SAS: synthetic accessibility score
- LogP: lipophilicity (optimal range 0-5 for oral administration)
- Molecular diversity: Tanimoto similarity between Morgan fingerprints
TamGen ranked first or second on 5 of 6 metrics and achieved the best overall score using mean reciprocal rank (MRR) across all metrics. On synthetic accessibility for high-affinity compounds, TamGen performed best. The generated compounds averaged 1.78 fused rings, closely matching FDA-approved drugs, while competing 3D methods produced compounds with significantly more fused rings.
TamGen was also 85x to 394x faster than competing methods: generating 100 compounds per target in an average of 9 seconds on a single A6000 GPU, compared to tens of minutes or hours for the baselines.
Design-Refine-Test Pipeline for ClpP Inhibitors
The practical application targeted ClpP protease of Mycobacterium tuberculosis, an emerging antibiotic target with no documented advanced inhibitors beyond Bortezomib.
Design stage: Using the ClpP binding pocket from PDB structure 5DZK, TamGen generated 2,612 unique compounds. Compounds were filtered by molecular docking (retaining those with better scores than Bortezomib) and Ligandformer phenotypic activity prediction. Peptidomimetic compounds were excluded for poor ADME properties. Four seed compounds were selected.
Refine stage: Using the 4 seed compounds plus 3 weakly active compounds ($\text{IC}_{50}$ 100-200 $\mu$M) from prior experiments, TamGen generated 8,635 unique compounds conditioned on both the target and seeds. After filtering, 296 compounds were selected for testing.
Test stage: From a 446k commercial compound library, 159 analogs (MCS similarity > 0.55) were identified. Five analogs showed significant inhibitory effects. Dose-response experiments revealed $\text{IC}_{50}$ values below 20 $\mu$M for all five, with Analog-005 achieving $\text{IC}_{50}$ of 1.9 $\mu$M. Three additional novel compounds were synthesized for SAR analysis:
| Compound | Series | Source | $\text{IC}_{50}$ ($\mu$M) | Key Feature |
|---|---|---|---|---|
| Analog-005 | II | Commercial library | 1.9 | Most potent analog |
| Analog-003 | I | Commercial library | < 20 | Strongest single-dose inhibition |
| Syn-A003-01 | I | TamGen (synthesized) | < 20 | Diphenylurea scaffold |
Both compound series (diphenylurea and benzenesulfonamide scaffolds) represent novel ClpP inhibitor chemotypes distinct from Bortezomib. Additionally, 6 out of 8 directly synthesized TamGen compounds demonstrated $\text{IC}_{50}$ below 40 $\mu$M, confirming TamGen’s ability to produce viable hits without the library search step.
Ablation Studies
Four ablation experiments clarified the contributions of TamGen’s components:
- Without pre-training: Significantly worse docking scores and simpler structures. The optimal decoder depth dropped from 12 to 4 layers without pre-training due to overfitting.
- Shuffled pocket-ligand pairs (TamGen-r): Substantially worse docking scores, confirming TamGen learns meaningful pocket-ligand interactions rather than generic compound distributions.
- Without distance-aware attention: Significant decline in docking scores when removing the geometric attention term from Eq. 2.
- Without coordinate augmentation: Performance degradation when removing the roto-translation augmentation $\rho$, highlighting the importance of geometric invariance.
Validated Drug-Like Generation with Practical Limitations
TamGen demonstrates that 1D SMILES-based generation with pre-training on natural compounds produces molecules with better drug-likeness properties than 3D generation methods. The experimental validation against ClpP is a notable strength, as most generative drug design methods lack biochemical assay confirmation.
Key limitations acknowledged by the authors include:
- Insufficient sensitivity to minor target differences: TamGen cannot reliably distinguish targets with point mutations or protein isoforms, limiting applicability for cancer-related proteins
- Requires known structure and pocket: As a structure-based method, TamGen needs the 3D structure of the target protein and binding pocket information
- Limited cellular validation: The study focuses on hit identification; cellular activities and toxicities of proposed compounds were not extensively tested
- 1D generation trade-off: SMILES-based generation does not fully exploit 3D protein-ligand geometric interactions available in coordinate space
Future directions include integrating insights from 3D autoregressive methods, using Monte Carlo Tree Search or reinforcement learning to guide generation for better docking scores and ADME/T properties, and property-guided generation as explored in PrefixMol.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Pre-training | PubChem (random sample) | 10M SMILES | Compound decoder pre-training |
| Fine-tuning | CrossDocked2020 | ~100k pairs | Filtered pocket-ligand pairs |
| Extended fine-tuning | CrossDocked + PDB | ~300k pairs | Used for TB compound generation |
| Evaluation | CrossDocked2020 test | 100 pockets | Same split as TargetDiff/Pocket2Mol |
Algorithms
- Compound decoder: 12-layer GPT with hidden dimension 768, pre-trained for 200k steps
- Protein encoder: 4-layer Transformer with hidden dimension 256, distance-aware attention
- VAE encoder: 4-layer standard Transformer encoder with hidden dimension 256
- Optimizer: Adam with initial learning rate $3 \times 10^{-5}$
- VAE $\beta$: 0.1 or 1.0 depending on generation stage
- Beam search: beam sizes of 4, 10, or 20 depending on stage
- Pocket definition: residues within 10 or 15 Angstrom distance cutoff from ligand center
Models
Pre-trained model weights are available via Zenodo at https://doi.org/10.5281/zenodo.13751391.
Evaluation
| Metric | TamGen | Best Baseline | Notes |
|---|---|---|---|
| Overall MRR | Best | TargetDiff (2nd) | Ranked across 6 metrics |
| Fused rings (avg) | 1.78 | ~3-5 (others) | Matches FDA-approved drug average |
| Generation speed | 9 sec/100 compounds | ~13 min (ResGen) | Single A6000 GPU |
| ClpP hit rate | 6/8 synthesized | N/A | $\text{IC}_{50}$ < 40 $\mu$M |
Hardware
- Pre-training: 8x V100 GPUs for 200k steps
- Inference benchmarking: 1x A6000 GPU
- Generation time: ~9 seconds per 100 compounds per target
| Artifact | Type | License | Notes |
|---|---|---|---|
| TamGen code | Code | MIT | Official implementation |
| Model weights and data | Model + Data | CC-BY-4.0 | Pre-trained weights, source data |
Paper Information
Citation: Wu, K., Xia, Y., Deng, P., Liu, R., Zhang, Y., Guo, H., Cui, Y., Pei, Q., Wu, L., Xie, S., Chen, S., Lu, X., Hu, S., Wu, J., Chan, C.-K., Chen, S., Zhou, L., Yu, N., Chen, E., Liu, H., Guo, J., Qin, T., & Liu, T.-Y. (2024). TamGen: drug design with target-aware molecule generation through a chemical language model. Nature Communications, 15, 9360. https://doi.org/10.1038/s41467-024-53632-4
@article{wu2024tamgen,
title={TamGen: drug design with target-aware molecule generation through a chemical language model},
author={Wu, Kehan and Xia, Yingce and Deng, Pan and Liu, Renhe and Zhang, Yuan and Guo, Han and Cui, Yumeng and Pei, Qizhi and Wu, Lijun and Xie, Shufang and Chen, Si and Lu, Xi and Hu, Song and Wu, Jinzhi and Chan, Chi-Kin and Chen, Shawn and Zhou, Liangliang and Yu, Nenghai and Chen, Enhong and Liu, Haiguang and Guo, Jinjiang and Qin, Tao and Liu, Tie-Yan},
journal={Nature Communications},
volume={15},
number={1},
pages={9360},
year={2024},
publisher={Nature Publishing Group},
doi={10.1038/s41467-024-53632-4}
}