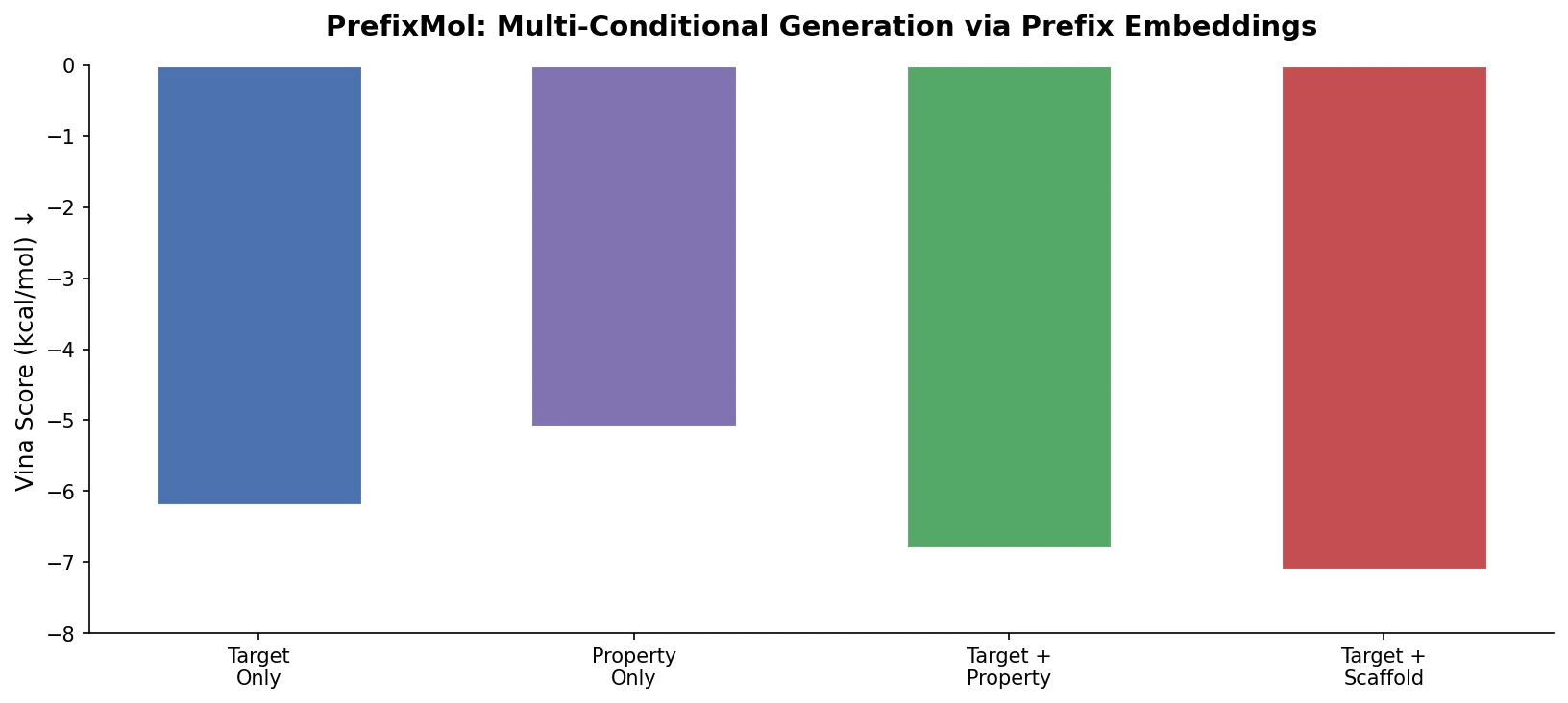
Unified Multi-Conditional Molecular Generation
PrefixMol is a Method paper that introduces a unified generative model for structure-based drug design that simultaneously conditions on protein binding pockets and multiple chemical properties. The primary contribution is a prefix-embedding mechanism, borrowed from NLP multi-task learning, that represents each condition (pocket geometry, Vina score, QED, SA, LogP, Lipinski) as a learnable feature vector prepended to the input sequence of a GPT-based SMILES generator. This allows a single model to handle customized multi-conditional generation without the negative transfer that typically arises from merging separate task-specific models.
Bridging Target-Aware and Chemistry-Aware Molecular Design
Prior structure-based drug design methods (e.g., Pocket2Mol, GraphBP) generate molecules conditioned on protein binding pockets but impose no constraints on the chemical properties of the output. Conversely, controllable molecule generation methods (e.g., REINVENT, RetMol, CMG) can steer chemical properties but ignore protein-ligand interactions. Merging these two objectives into a single model is difficult for two reasons:
- Data scarcity: Few datasets contain both protein-ligand binding affinity data and comprehensive molecular property annotations.
- Negative transfer: Treating each condition as a separate task in a multi-task framework can hurt overall performance when tasks conflict.
PrefixMol addresses both problems by extending the CrossDocked dataset with molecular property labels and using a parameter-efficient prefix conditioning strategy that decouples task-specific knowledge from the shared generative backbone.
Prefix Conditioning in Attention Layers
The core innovation adapts prefix-tuning from NLP to molecular generation. Given a GPT transformer that generates SMILES token-by-token, PrefixMol prepends $n_c$ learnable condition vectors $\mathbf{p}_{\phi} \in \mathbb{R}^{n_c \times d}$ to the left of the sequence embedding $\mathbf{x} \in \mathbb{R}^{l \times d}$, forming an extended input $\mathbf{x}’ = [\text{PREFIX}; \mathbf{x}]$.
The output of each position is:
$$ h_i = \begin{cases} p_{\phi,i}, & \text{if } i < n_c \\ \text{LM}_\theta(x_i’, h_{<i}), & \text{otherwise} \end{cases} $$
Because the prefix features always sit to the left, the causal attention mask ensures they influence all subsequent token predictions. The key insight is that the attention mechanism decomposes into a weighted sum of self-attention and prefix attention:
$$ \begin{aligned} \text{head} &= (1 - \lambda(\mathbf{x})) \underbrace{\text{Attn}(\mathbf{x}\mathbf{W}_q, \mathbf{c}\mathbf{W}_k, \mathbf{c}\mathbf{W}_v)}_{\text{self-attention}} \\ &\quad + \lambda(\mathbf{x}) \underbrace{\text{Attn}(\mathbf{x}\mathbf{W}_q, \mathbf{p}_\phi\mathbf{W}_k, \mathbf{p}_\phi\mathbf{W}_v)}_{\text{prefix attention}} \end{aligned} $$
where $\lambda(\mathbf{x})$ is a scalar representing the normalized attention weight on the prefix positions. This decomposition shows that conditions modulate generation through an additive attention pathway, and the activation map $\text{softmax}(\mathbf{x}\mathbf{W}_q \mathbf{W}_k^\top \mathbf{p}_\phi^\top)$ directly reveals how each condition steers model behavior.
Condition correlation is similarly revealed. For the prefix features themselves, the causal mask zeros out the cross-attention to the sequence, leaving only the prefix self-correlation term:
$$ \text{head} = \text{Attn}(\mathbf{p}_\phi \mathbf{W}_q, \mathbf{p}_\phi \mathbf{W}_k, \mathbf{p}_\phi \mathbf{W}_v) $$
The attention map $\mathbf{A}(\mathbf{p}_\phi)$ from this term encodes how conditions relate to one another.
Condition Encoders
Each condition has a dedicated encoder:
- 3D Pocket: A Geometric Vector Transformer (GVF) processes the binding pocket as a 3D graph with SE(3)-equivariant node and edge features. GVF extends GVP-GNN with a global attention module over geometric features. A position-aware attention mechanism with radial basis functions produces the pocket embedding.
- Chemical properties: Separate MLPs embed each scalar property (Vina, QED, SA, LogP, Lipinski) into the shared $d$-dimensional space.
Training Objective
PrefixMol is trained with two losses. The auto-regressive loss is:
$$ \mathcal{L}_{AT} = -\sum_{1 < i \leq t} \log p_{\phi, \theta}(x_i \mid \mathbf{x}_{<i}, \mathbf{p}_\phi) $$
A triplet property prediction loss encourages generated molecules to match desired properties:
$$ \mathcal{L}_{Pred} = \max\left((\hat{\mathbf{c}} - \mathbf{c})^2 - (\hat{\mathbf{c}} - \dot{\mathbf{c}})^2, 0\right) $$
where $\mathbf{c}$ is the input condition, $\hat{\mathbf{c}}$ is predicted by an MLP head, and $\dot{\mathbf{c}}$ is computed by RDKit from the generated SMILES (gradient is propagated through $\hat{\mathbf{c}}$ since RDKit is non-differentiable).
Experimental Setup and Controllability Evaluation
Dataset
The authors use the CrossDocked dataset (22.5 million protein-ligand structures) with chemical properties appended for each ligand. Data splitting and evaluation follow Pocket2Mol and Masuda et al.
Metrics
- Vina score (binding affinity, computed by QVina after UFF refinement)
- QED (quantitative estimate of drug-likeness, 0-1)
- SA (synthetic accessibility, 0-1)
- LogP (octanol-water partition coefficient)
- Lipinski (rule-of-five compliance count)
- High Affinity (fraction of pockets where generated molecules match or exceed test set affinities)
- Diversity (average pairwise Tanimoto distance over Morgan fingerprints)
- Sim.Train (maximum Tanimoto similarity to training set)
Baselines
Unconditional comparison against CVAE, AR (Luo et al. 2021a), and Pocket2Mol.
Key Results
Unconditional generation (Table 1): PrefixMol without conditions achieves sub-optimal results on Vina (-6.532), QED (0.551), SA (0.750), and LogP (1.415) compared to Pocket2Mol. However, it substantially outperforms all baselines on diversity (0.856 vs. 0.688 for Pocket2Mol) and novelty (Sim.Train of 0.239 vs. 0.376), indicating it generates genuinely novel molecules rather than memorizing training data.
Single-property control (Table 2): Molecular properties are positively correlated with conditional inputs across VINA, QED, SA, LogP, and Lipinski. With favorable control scales, PrefixMol surpasses Pocket2Mol on QED (0.767 vs. 0.563), SA (0.924 vs. 0.765), and LogP. The Vina score also improves when QED or LogP conditions are increased (e.g., -7.733 at QED control scale +2), revealing coupling between conditions.
Multi-property control (Table 3): Jointly adjusting all five conditions shows consistent positive relationships. For example, at control scale +4, QED reaches 0.722, SA reaches 0.913, and Lipinski saturates at 5.0. Joint QED+SA control at +2.0 achieves Lipinski = 5.0, confirming that certain properties are coupled.
Condition Relation Analysis
By computing partial derivatives of the prefix attention map with respect to each condition, the authors construct a relation matrix $\mathbf{R} = \sum_{i=2}^{6} |\partial \mathbf{A} / \partial c_i|$. Key findings:
- Vina is weakly self-controllable but strongly influenced by QED, LogP, and SA, explaining why multi-condition control improves binding affinity even when Vina alone responds poorly.
- LogP and QED are the most correlated property pair.
- Lipinski is coupled to QED and SA, saturating at 5.0 when both QED and SA control scales reach +2.
Key Findings, Limitations, and Interpretability Insights
PrefixMol demonstrates that prefix embedding is an effective strategy for unifying target-aware and chemistry-aware molecular generation. The main findings are:
- A single prefix-conditioned GPT model can control multiple chemical properties simultaneously while targeting specific protein pockets.
- Multi-conditional generation outperforms unconditional baselines in drug-likeness metrics, and the controllability enables PrefixMol to surpass Pocket2Mol on QED, SA, and LogP.
- The attention mechanism provides interpretable coupling relationships between conditions, offering practical guidance (e.g., improving QED indirectly improves Vina).
Limitations: The paper does not report validity rates for generated SMILES. The unconditional model underperforms Pocket2Mol on binding affinity (Vina), suggesting that generating 2D SMILES strings and relying on post hoc 3D conformer generation may be less effective than direct atom-by-atom 3D generation for binding affinity optimization. The condition relation analysis uses a first-order finite difference approximation ($\Delta = 1$), which may not capture nonlinear interactions. No external validation on prospective drug discovery tasks is provided. Hardware and training time details are not reported.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Training / Evaluation | CrossDocked (extended) | 22.5M protein-ligand structures | Extended with molecular properties (QED, SA, LogP, Lipinski, Vina) |
Algorithms
- GPT-based auto-regressive SMILES generation with prefix conditioning
- GVF (Geometric Vector Transformer) for 3D pocket encoding, extending GVP-GNN with global attention
- Separate MLP encoders for each chemical property
- Triplet property prediction loss with non-differentiable RDKit-computed properties
- QVina for Vina score computation with UFF refinement
Models
- GPT transformer backbone for SMILES generation
- 6 prefix condition vectors ($n_c = 6$): Pocket, Vina, QED, SA, LogP, Lipinski
- Specific architectural hyperparameters (hidden dimension, number of layers, heads) not reported in the paper
Evaluation
| Metric | PrefixMol (unconditional) | Pocket2Mol | Notes |
|---|---|---|---|
| Vina (kcal/mol) | -6.532 | -7.288 | Lower is better |
| QED | 0.551 | 0.563 | Higher is better |
| SA | 0.750 | 0.765 | Higher is better |
| Diversity | 0.856 | 0.688 | Higher is better |
| Sim.Train | 0.239 | 0.376 | Lower is better |
Hardware
Not reported in the paper.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| PrefixMol | Code | Not specified | Official PyTorch implementation |
Paper Information
Citation: Gao, Z., Hu, Y., Tan, C., & Li, S. Z. (2023). PrefixMol: Target- and Chemistry-aware Molecule Design via Prefix Embedding. arXiv preprint arXiv:2302.07120.
@article{gao2023prefixmol,
title={PrefixMol: Target- and Chemistry-aware Molecule Design via Prefix Embedding},
author={Gao, Zhangyang and Hu, Yuqi and Tan, Cheng and Li, Stan Z.},
journal={arXiv preprint arXiv:2302.07120},
year={2023}
}