Fingerprint-Based Evolutionary Molecular Design
This is a Method paper that introduces an evolutionary design methodology (EDM) for goal-directed molecular optimization. The primary contribution is a three-component framework where (1) molecules are encoded as extended-connectivity fingerprint (ECFP) vectors, (2) a genetic algorithm evolves these fingerprint vectors through mutation and crossover, (3) a recurrent neural network (RNN) decodes the evolved fingerprints back into valid SMILES strings, and (4) a deep neural network (DNN) evaluates molecular fitness. The key advantage over prior evolutionary approaches is that no hand-crafted chemical rules or fragment libraries are needed, as the RNN learns valid molecular reconstruction from data.
Challenges in Evolutionary Molecular Optimization
Evolutionary algorithms for molecular design face two core challenges. First, maintaining chemical validity of evolved molecules is difficult when operating on graph or string representations directly. Prior methods rely on predefined chemical rules and fragment libraries to constrain structural modifications (atom/bond additions, deletions, substitutions), but these introduce bias and risk convergence to local optima. Each new application domain requires specifying new chemical rules, which may not exist for emerging areas. Second, fitness evaluation must be both efficient and accurate. Simple evaluation methods like structural similarity indices or semi-empirical quantum chemistry calculations reduce computational cost but may not capture complex property relationships.
High-throughput computational screening (HTCS) is a common alternative, but it depends on the quality of predefined virtual chemical libraries and often requires multiple iterative enumerations, limiting its ability to explore novel chemical space.
Core Innovation: Evolving Fingerprints with Neural Decoding
The key insight is to perform genetic operations in fingerprint space rather than in molecular graph or SMILES string space. The framework comprises three learned functions:
Encoding function $e(\cdot)$: Converts a SMILES string $\mathbf{m}$ into a 5000-dimensional ECFP vector $\mathbf{x}$ using Morgan fingerprints with a neighborhood radius of 6. This is a deterministic hash-based encoding (not learned).
Decoding function $d(\cdot)$: An RNN with three hidden layers of 500 LSTM units that reconstructs a SMILES string from an ECFP vector. The RNN generates SMILES as a sequence of three-character substrings, conditioning each prediction on the current substring and the input ECFP vector:
$$d(\mathbf{x}) = \mathbf{m}, \quad \text{where } p(\mathbf{m}_{t+1} | \mathbf{m}_{t}, \mathbf{x})$$
The three-character substring approach reduces the ratio of invalid SMILES by imposing additional constraints on subsequent characters.
Property prediction function $f(\cdot)$: A five-layer DNN with 250 hidden units per layer that predicts molecular properties from ECFP vectors:
$$\mathbf{t} = f(e(\mathbf{m}))$$
The RNN is trained by minimizing cross-entropy loss between the softmax output and the target SMILES string $\mathbf{m}_{i}$, learning the relationship $d(e(\mathbf{m}_{i})) = \mathbf{m}_{i}$. The DNN is trained by minimizing mean squared error between predicted and computed property values. Both use the Adam optimizer with mini-batch size 100, 500 training epochs, and dropout rate 0.5.
Genetic Algorithm Operations
The GA evolves ECFP vectors using the DEAP library with the following parameters:
- Population size: 50
- Crossover rate: 0.7 (uniform crossover, mixing ratio 0.2)
- Mutation rate: 0.3 (Gaussian mutation, $N(0, 0.2^{2})$, applied to 1% of elements)
- Selection: Tournament selection with size 3, top 3 individuals as parents
- Termination: 500 generations or 30 consecutive generations without fitness improvement
The evolutionary loop proceeds as follows: a seed molecule $\mathbf{m}_{0}$ is encoded to $\mathbf{x}_{0}$, mutated to generate a population $\mathbf{P}^{0} = {\mathbf{z}_{1}, \mathbf{z}_{2}, \ldots, \mathbf{z}_{L}}$, each vector is decoded via the RNN, validity is checked with RDKit, fitness is evaluated via the DNN, and the top parents produce the next generation through crossover and mutation.
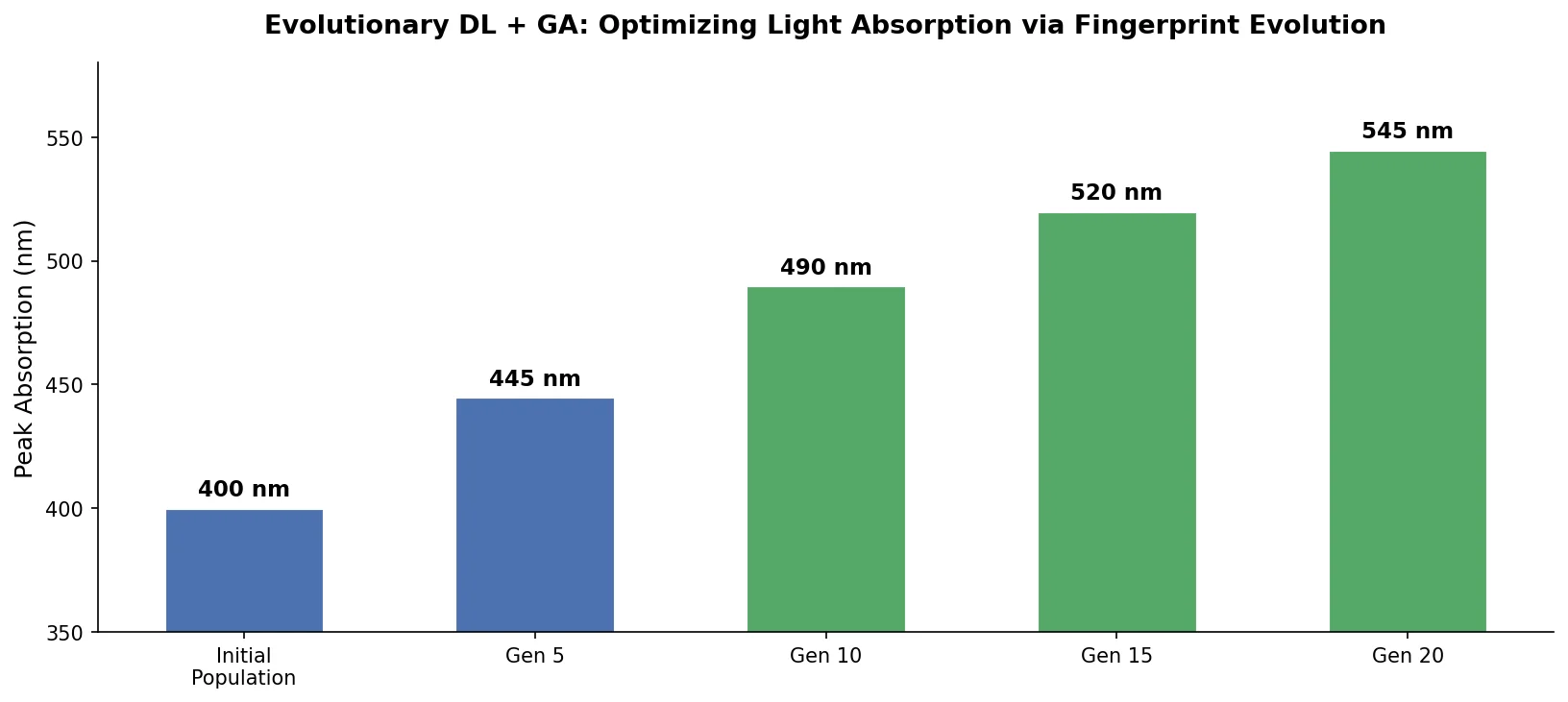
Experimental Setup: Light-Absorbing Wavelength Optimization
Training Data and Deep Learning Performance
The models were trained on 10,000 to 100,000 molecules randomly sampled from PubChem (molecular weight 200-600 g/mol). Each molecule was labeled with DFT-computed excitation energy ($S_{1}$), HOMO, and LUMO energies using B3LYP/6-31G.
| Training Data | Validity (%) | Reconstructability (%) | $S_{1}$ (R, MAE) | HOMO (R, MAE) | LUMO (R, MAE) |
|---|---|---|---|---|---|
| 100,000 | 88.8 | 62.4 | 0.977, 0.185 eV | 0.948, 0.168 eV | 0.960, 0.195 eV |
| 50,000 | 86.7 | 60.1 | 0.973, 0.198 eV | 0.945, 0.172 eV | 0.955, 0.209 eV |
| 30,000 | 85.3 | 59.8 | 0.930, 0.228 eV | 0.934, 0.191 eV | 0.945, 0.224 eV |
| 10,000 | 83.2 | 55.7 | 0.913, 0.278 eV | 0.885, 0.244 eV | 0.917, 0.287 eV |
Validity refers to the proportion of chemically valid SMILES after RDKit inspection. Reconstructability measures how often the RNN can reproduce the original molecule from its ECFP (62.4% at 100k training samples by matching canonical SMILES among 10,000 generated strings).
Design Task 1: Unconstrained S1 Modification
Fifty seed molecules with $S_{1}$ values between 3.8 eV and 4.2 eV were evolved in both increasing and decreasing directions. With 50,000 training samples, $S_{1}$ increased by approximately 60% on average in the increasing direction and showed slightly lower rates of change in the decreasing direction. The asymmetry is attributed to the skewed $S_{1}$ distribution of training data (average $S_{1}$ of 4.3-4.4 eV, higher than the seed median of 4.0 eV). Performance saturated at approximately 50,000 training samples.
Design Task 2: S1 Modification with HOMO/LUMO Constraints
The same 50 seeds were evolved with constraints: $-7.0 \text{ eV} < \text{HOMO} < -5.0 \text{ eV}$ and $\text{LUMO} < 0.0 \text{ eV}$. In the increasing $S_{1}$ direction, constraints suppressed the rate of change because both HOMO and LUMO bounds limit the achievable HOMO-LUMO gap. In the decreasing direction, constraints had minimal effect because LUMO could freely decrease while HOMO had sufficient room to rise within the allowed range.
Design Task 3: Extrapolation Beyond Training Data
To generate molecules with $S_{1}$ values below 1.77 eV (outside the training distribution, which had mean $S_{1}$ of 4.91 eV), the authors introduced iterative “phases”: generate molecules, compute their properties via DFT, retrain the models, and repeat. Starting from the 30 lowest-$S_{1}$ seed molecules with 300 generation runs per phase:
- Phase 1: Average $S_{1}$ = 2.20 eV, 12 molecules below 1.77 eV
- Phase 2: Average $S_{1}$ = 2.22 eV, 37 molecules below 1.77 eV
- Phase 3: Average $S_{1}$ = 2.31 eV, 58 molecules below 1.77 eV
While the average $S_{1}$ rose slightly across phases, variance decreased (from 1.40 to 1.36), indicating the model concentrated its outputs closer to the target range. This active-learning-like loop demonstrates the framework can extend beyond the training distribution.
Design Task 4: GuacaMol Benchmarks
The method was evaluated on the GuacaMol goal-directed benchmark suite using the ChEMBL25 training dataset. The RNN model was retrained with three-character substrings.
| Benchmark | Best of Dataset | SMILES LSTM | SMILES GA | Graph GA | Graph MCTS | cRNN | EDM (ours) |
|---|---|---|---|---|---|---|---|
| Celecoxib rediscovery | 0.505 | 1.000 | 0.607 | 1.000 | 0.378 | 1.000 | 1.000 |
| Troglitazone rediscovery | 0.419 | 1.000 | 0.558 | 1.000 | 0.312 | 1.000 | 1.000 |
| Thiothixene rediscovery | 0.456 | 1.000 | 0.495 | 1.000 | 0.308 | 1.000 | 1.000 |
| LogP(-1.0) | 1.000 | 1.000 | 1.000 | 1.000 | 0.980 | 1.000 | 1.000 |
| LogP(8.0) | 1.000 | 1.000 | 1.000 | 1.000 | 0.979 | 1.000 | 1.000 |
| TPSA(150.0) | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 |
| CNS MPO | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 | 1.000 |
| QED | 0.948 | 0.948 | 0.948 | 0.948 | 0.944 | 0.948 | 0.948 |
The EDM achieves maximum scores on all eight tasks, matching the cRNN baseline. The 256 highest-scoring molecules from the ChEMBL25 test set were used as seeds, with 500 SMILES strings generated per seed.
Key Findings and Limitations
Results
The evolutionary design framework successfully evolved seed molecules toward target properties across all four design tasks. The RNN decoder maintained 88.8% chemical validity at 100k training samples, and the DNN property predictor achieved correlation coefficients above 0.94 for $S_{1}$, HOMO, and LUMO prediction. The iterative retraining procedure enabled exploration outside the training data distribution, generating 58 molecules with $S_{1}$ below 1.77 eV after three phases. On GuacaMol benchmarks, the method achieved maximum scores on all eight tasks, matching SMILES LSTM, Graph GA, and cRNN baselines.
Limitations
Several limitations are worth noting:
- Reconstructability ceiling: Only 62.4% of molecules could be reconstructed from their ECFP vectors, meaning the RNN decoder fails to recover the original molecule approximately 38% of the time. This information loss in the ECFP encoding is a fundamental bottleneck.
- Data dependence: Performance is sensitive to the training data distribution. The asymmetric evolution rates for increasing vs. decreasing $S_{1}$ directly reflect the skewed training data.
- Structural constraints: Three heuristic constraints (fused ring sizes, number of fused rings, alkyl chain lengths) were still needed to maintain reasonable molecular structures, partially undermining the claim of a fully data-driven approach.
- DFT reliance: The extrapolation experiment requires DFT calculations in the loop, which are computationally expensive and may limit scalability.
- Limited benchmark scope: Only 8 GuacaMol tasks were tested, and all achieved perfect scores, making it difficult to differentiate from competing methods. The paper does not report on harder multi-objective benchmarks.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Training/Evaluation | PubChem random sample | 10,000-100,000 molecules | MW 200-600 g/mol, labeled with DFT-computed $S_{1}$, HOMO, LUMO |
| GuacaMol Benchmark | ChEMBL25 | Standard split | Used for retraining RNN; 256 top-scoring seeds |
Algorithms
- Genetic algorithm: DEAP library; population 50, crossover rate 0.7, mutation rate 0.3, tournament size 3
- RNN decoder: 3 hidden layers, 500 LSTM units each, three-character substring generation
- DNN predictor: 5 layers, 250 hidden units, sigmoid activations, linear output
- Training: Adam optimizer, mini-batch 100, 500 epochs, dropout 0.5
Models
All neural networks were implemented using Keras with the Theano backend (GPU-accelerated). No pre-trained model weights are publicly available.
Evaluation
- RNN validity: Proportion of chemically valid SMILES (RDKit check)
- Reconstructability: Fraction of seed molecules recoverable from ECFP (canonical SMILES match in 10,000 generated strings)
- DNN accuracy: Correlation coefficient (R) and MAE via 10-fold cross-validation
- Evolutionary performance: Average rate of $S_{1}$ change across 50 seeds; molecule count in target range
- GuacaMol: Standard rediscovery and property satisfaction benchmarks
Hardware
The paper does not specify GPU models, training times, or computational requirements for the evolutionary runs. DFT calculations used the Gaussian 09 program suite with B3LYP/6-31G.
Artifacts
No public code repository or pre-trained models are available. The paper is published under a CC-BY 4.0 license as open access in Scientific Reports.
| Artifact | Type | License | Notes |
|---|---|---|---|
| Paper (Nature) | Paper | CC-BY 4.0 | Open access |
Reproducibility classification: Partially Reproducible. The method is described in sufficient detail for reimplementation, but no code, trained models, or preprocessed datasets are released. The DFT calculations require Gaussian 09, a commercial software package.
Paper Information
Citation: Kwon, Y., Kang, S., Choi, Y.-S., & Kim, I. (2021). Evolutionary design of molecules based on deep learning and a genetic algorithm. Scientific Reports, 11, 17304. https://doi.org/10.1038/s41598-021-96812-8
@article{kwon2021evolutionary,
title={Evolutionary design of molecules based on deep learning and a genetic algorithm},
author={Kwon, Youngchun and Kang, Seokho and Choi, Youn-Suk and Kim, Inkoo},
journal={Scientific Reports},
volume={11},
number={1},
pages={17304},
year={2021},
publisher={Nature Publishing Group},
doi={10.1038/s41598-021-96812-8}
}