A Language Model for Joint 3D Molecular Graph and Conformation Generation
BindGPT is a Method paper that introduces a GPT-based language model for generating 3D molecular structures. The primary contribution is a unified framework that jointly produces molecular graphs (via SMILES) and 3D coordinates (via XYZ tokens) within a single autoregressive model. This eliminates the need for external graph reconstruction tools like OpenBabel, which are error-prone when applied to noisy atom positions. The same pre-trained model serves as a 3D molecular generative model, a conformer generator conditioned on molecular graphs, and a pocket-conditioned 3D molecule generator.
The Graph Reconstruction Problem in 3D Molecular Generation
Most existing 3D molecular generators focus on predicting atom types and positions, relying on supplementary software (e.g., OpenBabel or RDKit) to reconstruct molecular bonds from predicted coordinates. This introduces a fragile dependency: small positional errors can drastically change the reconstructed molecular graph or produce disconnected structures. Additionally, while diffusion models and equivariant GNNs have shown strong results for 3D molecular generation, they often depend on SE(3) equivariance inductive biases and are computationally expensive at sampling time (up to $10^6$ seconds for 1000 valid molecules for EDM). The pocket-conditioned generation task is further limited by the small size of available 3D binding pose datasets (e.g., CrossDocked), making it difficult for specialized models to generalize without large-scale pre-training.
SMILES+XYZ Tokenization: Jointly Encoding Graphs and Coordinates
The core innovation in BindGPT is coupling SMILES notation with XYZ coordinate format in a single token sequence. The sequence starts with a <LIGAND> token, followed by character-level SMILES tokens encoding the molecular graph, then an <XYZ> token marking the transition to coordinate data. Each 3D atom position is encoded using 6 tokens (integer and fractional parts for each of the three coordinates). The atom ordering is synchronized between SMILES and XYZ, so atom symbols from SMILES are not repeated in the coordinate section.
For protein pockets, sequences begin with a <POCKET> token followed by atom names and coordinates. Following AlphaFold’s approach, only alpha-carbon coordinates are retained to keep pocket representations compact.
The model uses the GPT-NeoX architecture with rotary position embeddings (RoPE), which enables length generalization between pre-training and fine-tuning where sequence lengths differ substantially. The pre-trained model has 108M parameters with 15 layers, 12 attention heads, and a hidden dimension of 768.
Pre-training on Large-Scale 3D Data
Pre-training uses the Uni-Mol dataset containing 208M conformations for 12M molecules and 3.2M protein pocket structures. Each training batch contains either ligand sequences or pocket sequences (not mixed within a sequence). Since pockets are far fewer than ligands, the training schedule runs 5 pocket epochs per ligand epoch, resulting in roughly 8% pocket tokens overall. Training uses large batches of 1.6M tokens per step with Flash Attention and DeepSpeed optimizations.
Supervised Fine-Tuning with Augmentation
For pocket-conditioned generation, BindGPT is fine-tuned on CrossDocked 2020, which contains aligned pocket-ligand pairs. Unlike prior work that subsamples less than 1% of the best pairs, BindGPT uses all intermediate ligand poses (including lower-quality ones), yielding approximately 27M pocket-ligand pairs. To combat overfitting on the limited diversity (14k unique molecules, 3k pockets), two augmentation strategies are applied:
- SMILES randomization: Each molecule can yield 100-1000 different valid SMILES strings
- Random 3D rotation: The same rotation matrix is applied to both pocket and ligand coordinates
During fine-tuning, the pocket token sequence is concatenated before the ligand token sequence. An optional variant conditions on binding energy scores from the CrossDocked dataset, enabling contrastive learning between good and bad binding examples.
Reinforcement Learning with Docking Feedback
BindGPT applies REINFORCE (not PPO or REINVENT, which were found less stable) to further optimize pocket-conditioned generation. On each RL step, the model generates 3D ligand structures for a batch of random protein pockets, computes binding energy rewards using QVINA docking software, and updates model parameters. A KL-penalty between the current model and the SFT initialization stabilizes training.
The RL objective can be written as:
$$\mathcal{L}_{\text{RL}} = -\mathbb{E}_{x \sim \pi_\theta}\left[ R(x) \right] + \beta \cdot D_{\text{KL}}(\pi_\theta | \pi_{\text{SFT}})$$
where $R(x)$ is the docking reward from QVINA and $\beta$ controls the strength of the KL regularization.
Experimental Evaluation Across Three 3D Generation Tasks
Datasets
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Pre-training | Uni-Mol 3D | 208M conformations (12M molecules) + 3.2M pockets | Large-scale 3D molecular dataset |
| Fine-tuning (SFT) | CrossDocked 2020 | ~27M pocket-ligand pairs | 14k molecules x 3k pockets, includes all pose qualities |
| Fine-tuning (conformer) | GEOM-DRUGS | 27M conformations for 300k molecules | Standard benchmark for 3D conformer generation |
| Evaluation (conformer) | Platinum | Experimentally validated conformations | Zero-shot evaluation holdout |
| Evaluation (pocket) | CrossDocked holdout | 100 pockets | Held out from training |
Task 1: 3D Molecule Generation (Pre-training)
Compared against XYZ-Transformer (the only other model capable of large-scale pre-training), BindGPT achieves 98.58% validity (vs. 12.87% for XYZ-TF without hydrogens), higher SA (0.77 vs. 0.21), QED (0.59 vs. 0.30), and Lipinski scores (4.86 vs. 4.79). BindGPT also produces conformations with RMSD of 0.89 (XYZ-TF’s RMSD calculation failed to converge). Generation is 12x faster (13s vs. 165s for 1000 molecules).
Task 2: 3D Molecule Generation (Fine-tuned on GEOM-DRUGS)
Against EDM and MolDiff (diffusion baselines), BindGPT outperforms on nearly all 3D distributional metrics:
| Metric | EDM | MolDiff | BindGPT |
|---|---|---|---|
| JS bond lengths | 0.246 | 0.365 | 0.029 |
| JS bond angles | 0.282 | 0.155 | 0.075 |
| JS dihedral angles | 0.328 | 0.162 | 0.098 |
| JS freq. bond types | 0.378 | 0.163 | 0.045 |
| JS freq. bond pairs | 0.396 | 0.136 | 0.043 |
| JS freq. bond triplets | 0.449 | 0.125 | 0.042 |
| Time (1000 molecules) | 1.4e6 s | 7500 s | 200 s |
BindGPT is two orders of magnitude faster than diffusion baselines while producing more accurate 3D geometries. MolDiff achieves better drug-likeness scores (QED, SA), but the authors argue 3D distributional metrics are more relevant for evaluating 3D structure fidelity.
Task 3: Pocket-Conditioned Molecule Generation
| Method | Vina Score | SA | QED | Lipinski |
|---|---|---|---|---|
| Pocket2Mol | -7.15 +/- 4.89 | 0.75 | 0.57 | 4.88 |
| TargetDiff | -7.80 +/- 3.61 | 0.58 | 0.48 | 4.51 |
| BindGPT-FT | -5.44 +/- 2.09 | 0.78 | 0.50 | 4.72 |
| BindGPT-RFT | -7.24 +/- 1.68 | 0.74 | 0.48 | 4.32 |
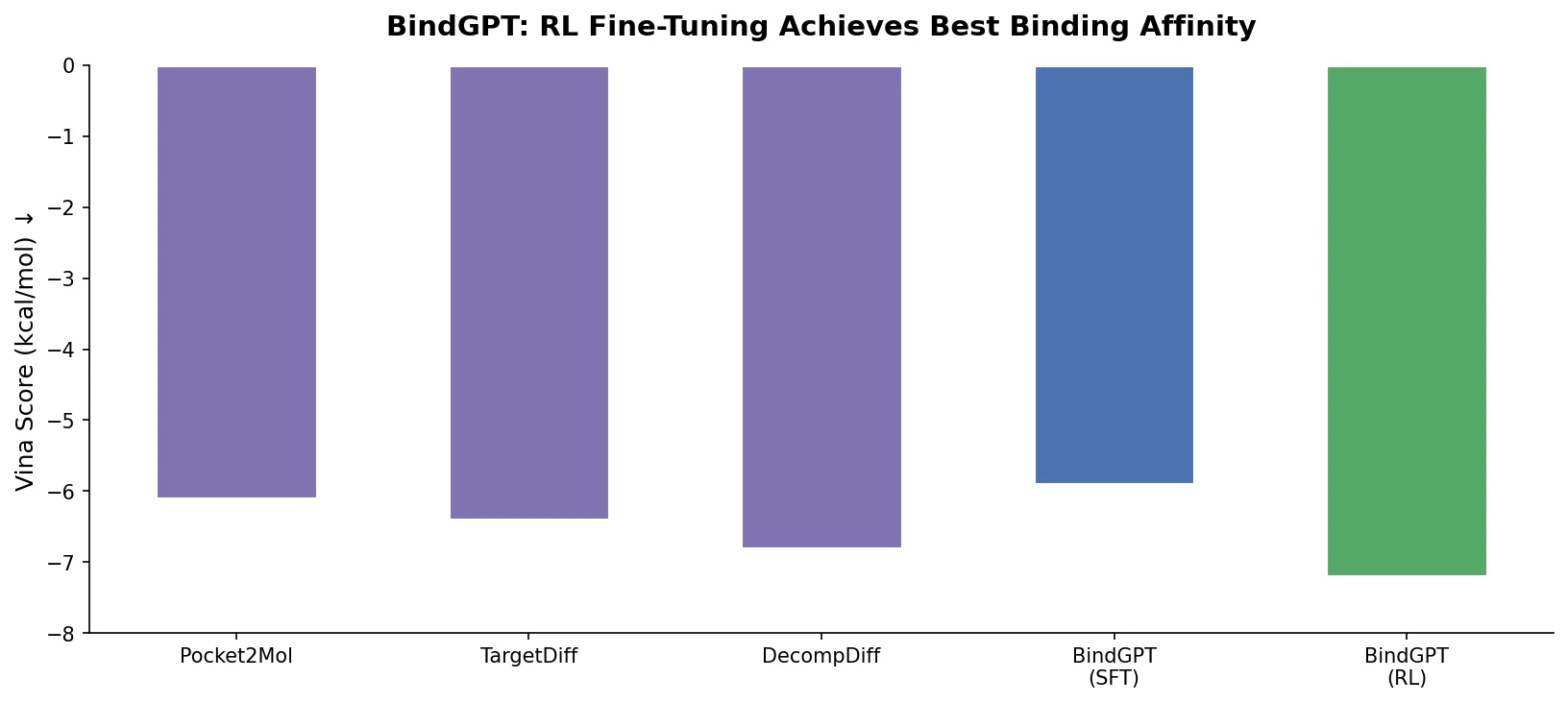
| BindGPT-RL | -8.60 +/- 1.90 | 0.84 | 0.43 | 4.81 |
The RL-fine-tuned model achieves the best Vina binding scores (-8.60 vs. -7.80 for TargetDiff) with lower variance and the highest SA score (0.84). The SFT-only model (BindGPT-FT) underperforms baselines on binding score, demonstrating that RL is essential for strong pocket-conditioned generation. QED is lower for BindGPT-RL, but the authors note that QED could be included in the RL reward and was excluded for fair comparison.
Conformer Generation
On the Platinum dataset (zero-shot), BindGPT matches the performance of Torsional Diffusion (the specialized state-of-the-art) when assisted by RDKit, with a small gap without RDKit assistance. Uni-Mol fails to generalize to this dataset despite pre-training on the same Uni-Mol data.
Key Findings, Limitations, and Future Directions
BindGPT demonstrates that a simple autoregressive language model without equivariance inductive biases can match or surpass specialized diffusion models and GNNs across multiple 3D molecular generation tasks. The key findings include:
- Joint SMILES+XYZ generation eliminates graph reconstruction errors, achieving 98.58% validity compared to 12.87% for XYZ-Transformer
- Large-scale pre-training is critical for pocket-conditioned generation, as none of the baselines use pre-training and instead rely on heavy inductive biases
- RL fine-tuning with docking feedback substantially improves binding affinity beyond what SFT alone achieves
- Sampling is two orders of magnitude faster than diffusion baselines (200s vs. 1.4M s for EDM)
Limitations include the relatively modest model size (108M parameters), with the authors finding this sufficient for current tasks but not exploring larger scales. The RL optimization uses only Vina score as reward; multi-objective optimization incorporating SA, QED, and other properties is left as future work. The model also relies on character-level SMILES tokenization rather than more sophisticated chemical tokenizers. BindGPT is the first model to explicitly generate hydrogens at scale, though validity drops from 98.58% to 77.33% when hydrogens are included.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Pre-training | Uni-Mol 3D | 208M conformations, 12M molecules, 3.2M pockets | From Zhou et al. (2023) |
| SFT (pocket) | CrossDocked 2020 | ~27M pocket-ligand pairs | Full version including low-quality poses |
| SFT (conformer) | GEOM-DRUGS | 27M conformations, 300k molecules | Standard benchmark |
| Evaluation | Platinum | Experimentally validated conformations | Zero-shot holdout |
Algorithms
- Architecture: GPT-NeoX with rotary position embeddings (RoPE)
- Pre-training: Causal language modeling with 1.6M tokens per batch
- SFT augmentation: SMILES randomization + random 3D rotation
- RL: REINFORCE with KL-penalty from SFT initialization; QVINA docking as reward
Models
- Size: 108M parameters, 15 layers, 12 heads, hidden size 768
- Vocabulary: Character-level SMILES tokens + special tokens (
<LIGAND>,<POCKET>,<XYZ>) + coordinate tokens (6 per 3D position)
Evaluation
- Validity, SA, QED, Lipinski: Standard drug-likeness metrics
- Jensen-Shannon divergences: Distribution-level 3D structural metrics (bond lengths, angles, dihedrals, bond types)
- RMSD: Alignment quality of generated conformations vs. RDKit reference
- RMSD-Coverage: CDF of RMSD between generated and reference conformers
- Vina score: Binding energy from QVINA docking software
Hardware
- Pre-training and fine-tuning use Flash Attention and DeepSpeed for efficiency
- Specific GPU counts and training times are described in Appendix G (not available in the main text)
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| Project Page | Other | Not specified | Project website with additional details |
No public code repository or pre-trained model weights were identified. The project website exists but no source code has been released as of this writing.
Reproducibility Status: Partially Reproducible. The paper provides detailed architecture specs and hyperparameters, but no public code or model weights are available. All training datasets (Uni-Mol, CrossDocked, GEOM-DRUGS) are publicly accessible.
Paper Information
Citation: Zholus, A., Kuznetsov, M., Schutski, R., Shayakhmetov, R., Polykovskiy, D., Chandar, S., & Zhavoronkov, A. (2025). BindGPT: A Scalable Framework for 3D Molecular Design via Language Modeling and Reinforcement Learning. Proceedings of the AAAI Conference on Artificial Intelligence, 39(24), 26083-26091. https://doi.org/10.1609/aaai.v39i24.34804
@inproceedings{zholus2025bindgpt,
title={BindGPT: A Scalable Framework for 3D Molecular Design via Language Modeling and Reinforcement Learning},
author={Zholus, Artem and Kuznetsov, Maksim and Schutski, Roman and Shayakhmetov, Rim and Polykovskiy, Daniil and Chandar, Sarath and Zhavoronkov, Alex},
booktitle={Proceedings of the AAAI Conference on Artificial Intelligence},
volume={39},
number={24},
pages={26083--26091},
year={2025},
doi={10.1609/aaai.v39i24.34804}
}