This group covers generative models that condition molecule design on protein target information. The goal is to generate molecules complementary to a specific protein, whether through sequence translation, pocket encoding, or 3D-aware coordinate generation.
| Paper | Year | Approach | Key Idea |
|---|---|---|---|
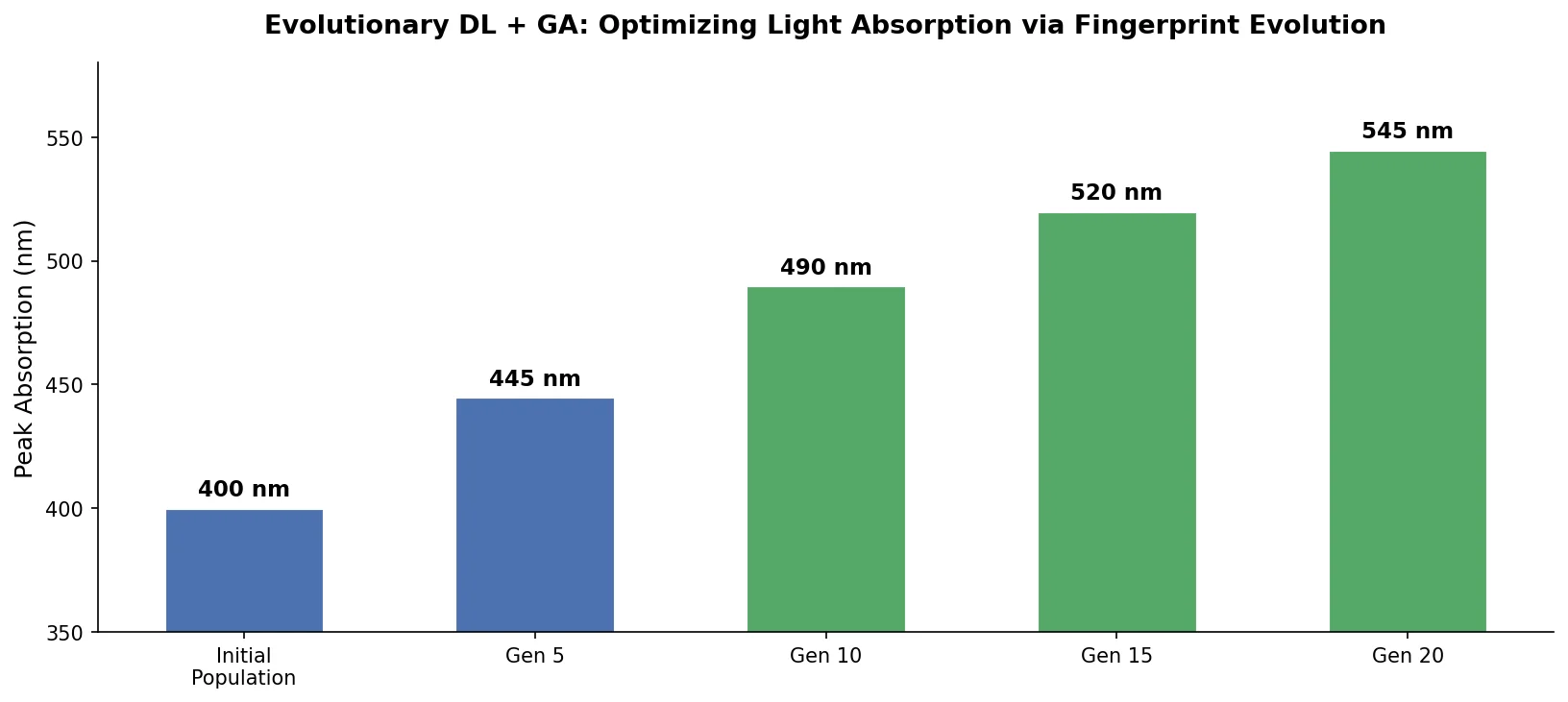
| Evolutionary Design | 2021 | GA + RNN decoder | Genetic algorithm on ECFP fingerprints with DNN property predictor |
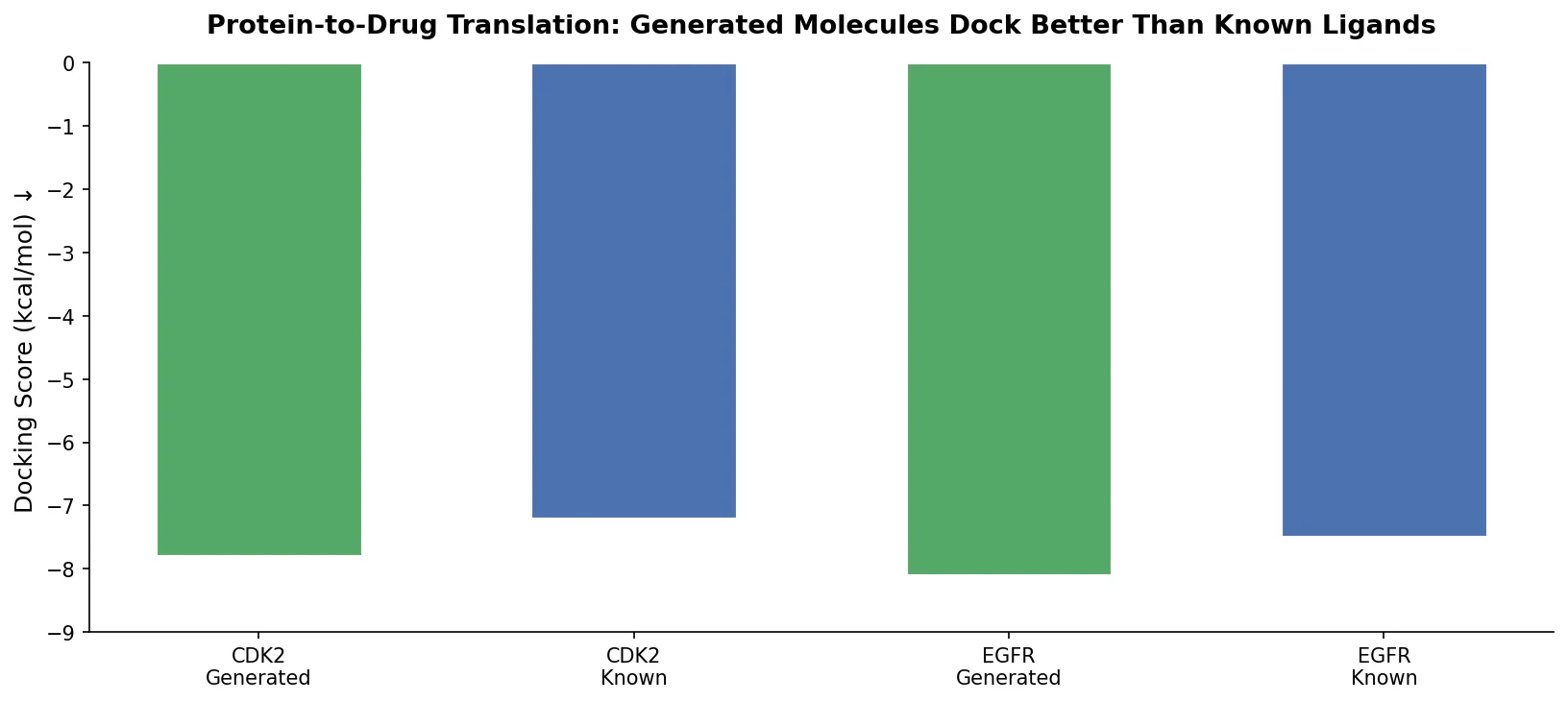
| Protein-to-Drug | 2021 | Seq2seq Transformer | Amino acid sequence to SMILES translation |
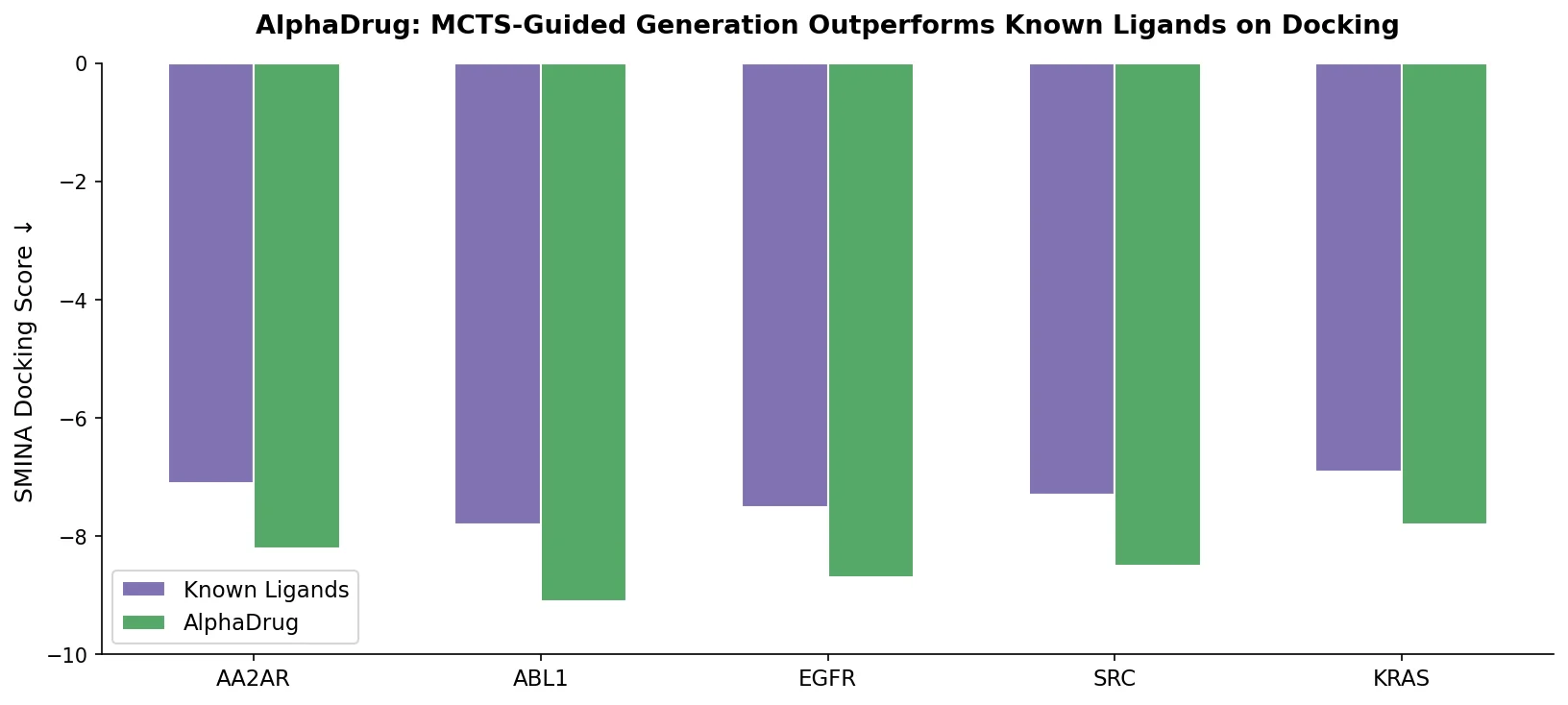
| AlphaDrug | 2022 | Transformer + MCTS | Monte Carlo tree search with docking rollouts for target-specific design |
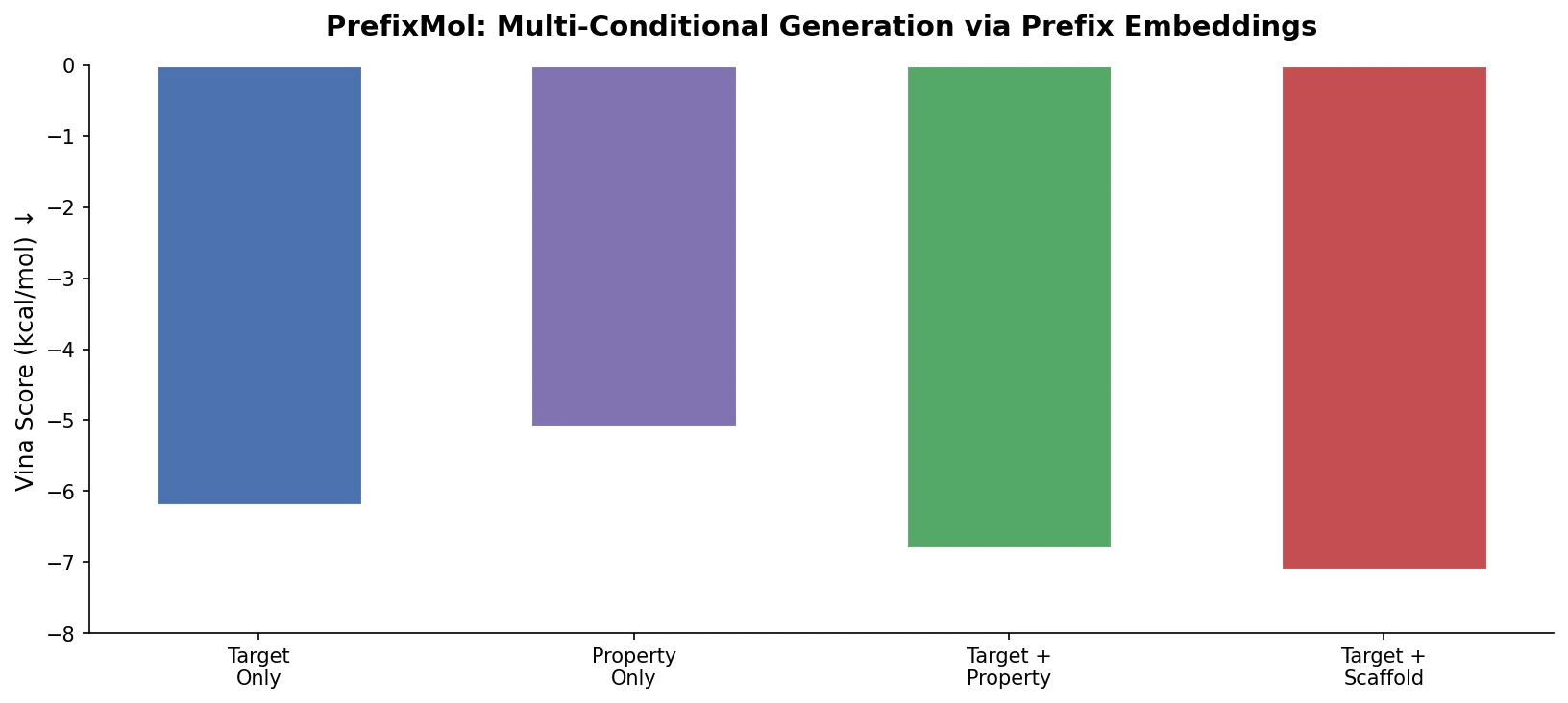
| PrefixMol | 2023 | GPT + prefix tuning | Learnable prefix embeddings for joint pocket and property conditioning |
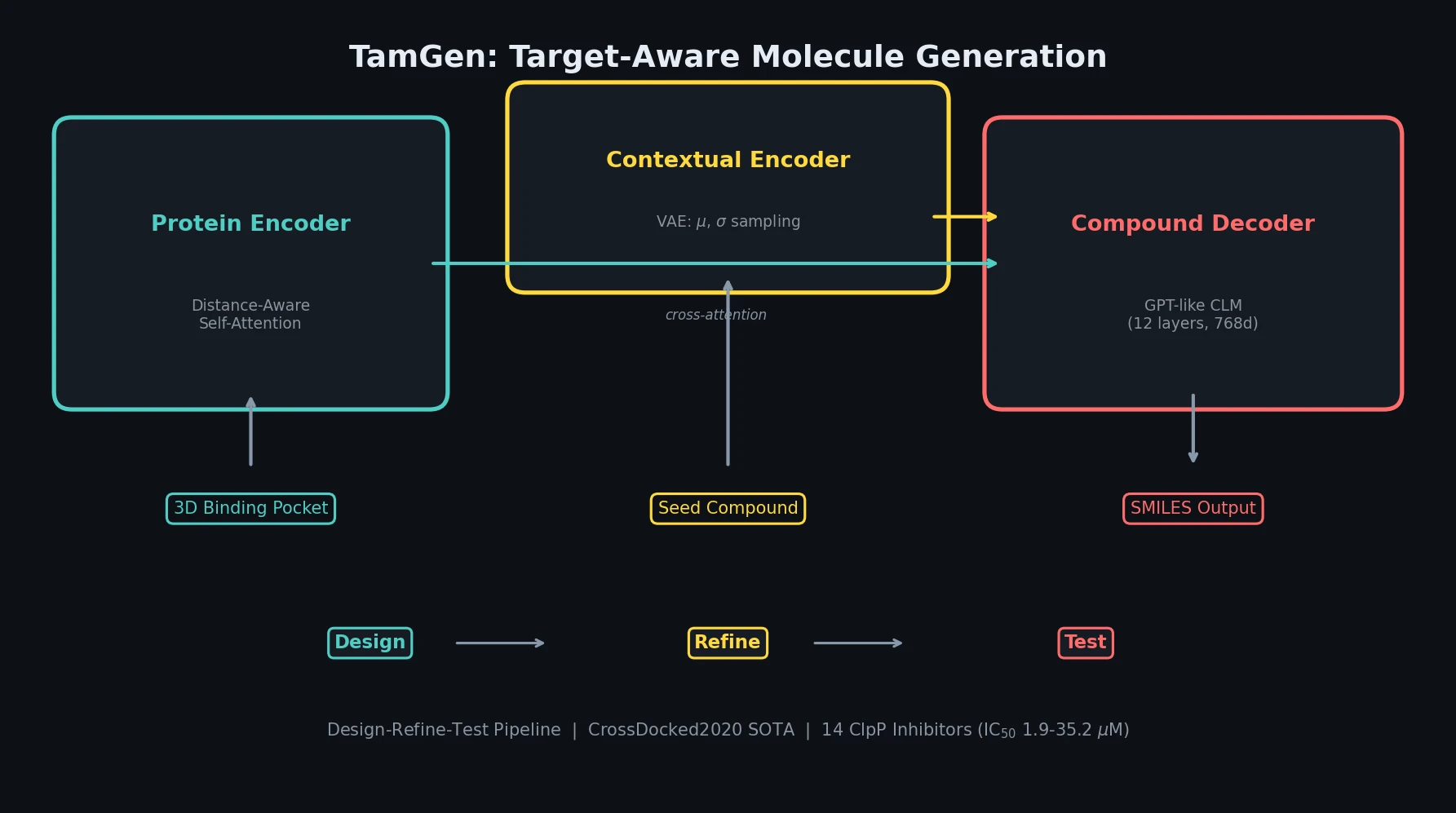
| TamGen | 2024 | GPT + pocket encoder | Pocket-conditioned generation with VAE refinement and experimental validation |
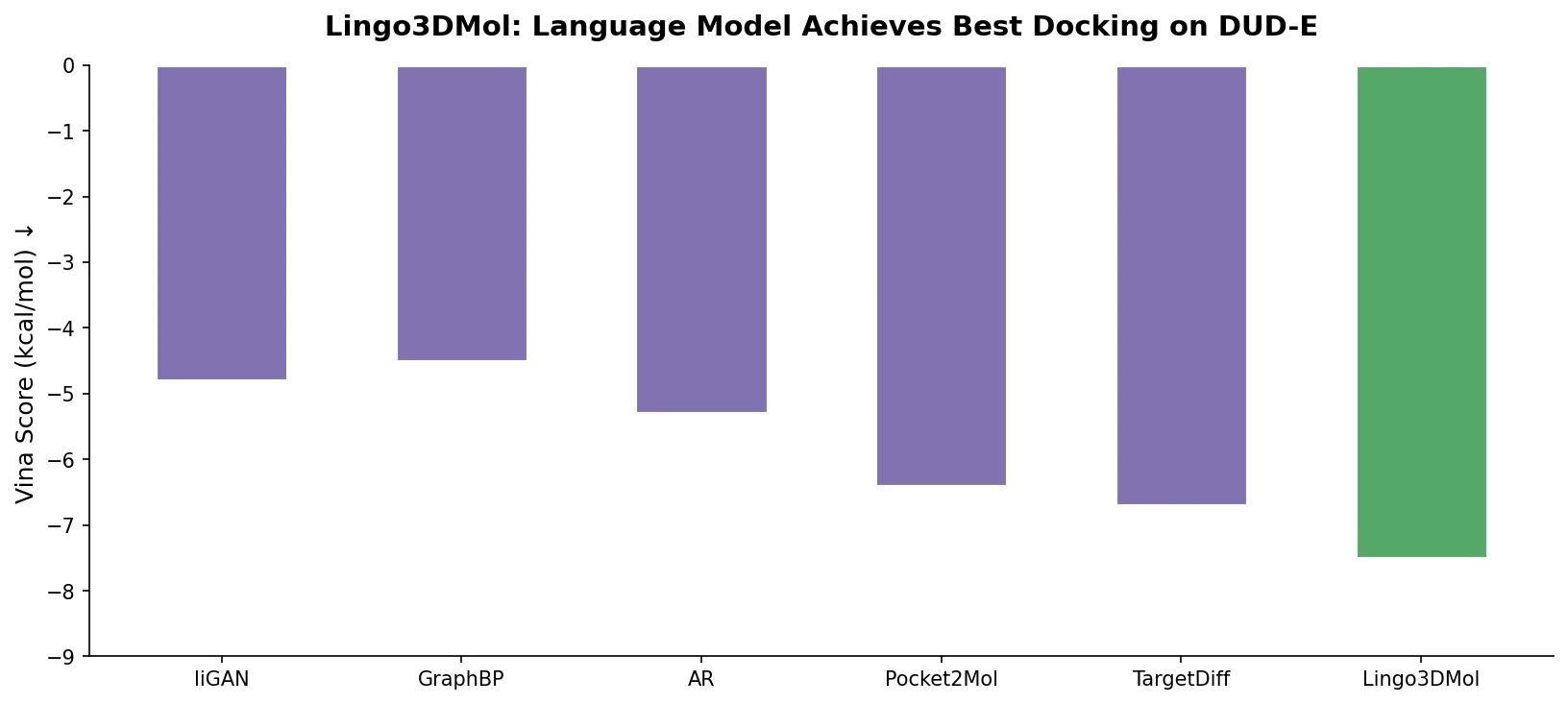
| Lingo3DMol | 2024 | LM + geometric DL | Fragment-based SMILES with 3D coordinates for pocket-conditioned design |
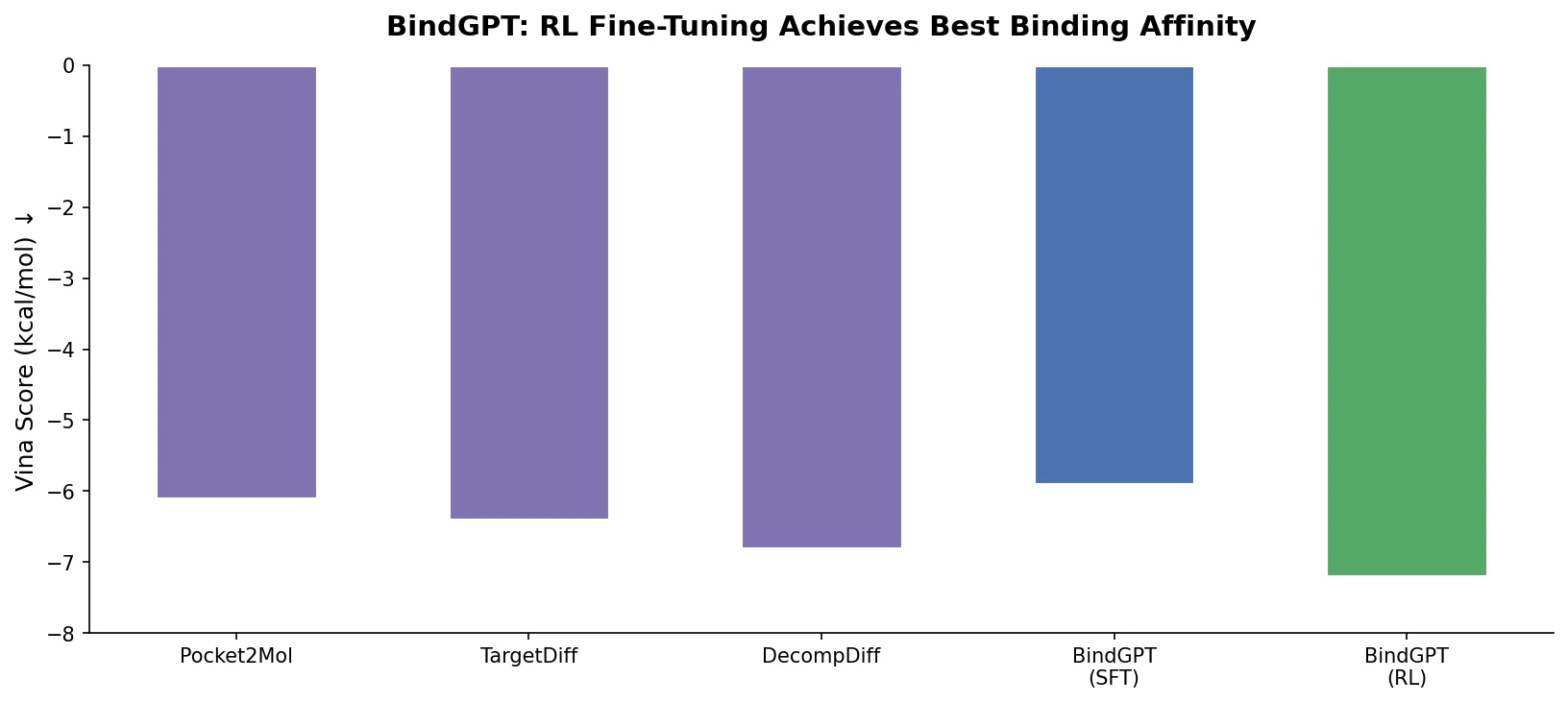
| BindGPT | 2025 | GPT | Autoregressive SMILES+XYZ generation with RL-based docking optimization |