Combining GANs and Reinforcement Learning for Goal-Directed Sequence Generation
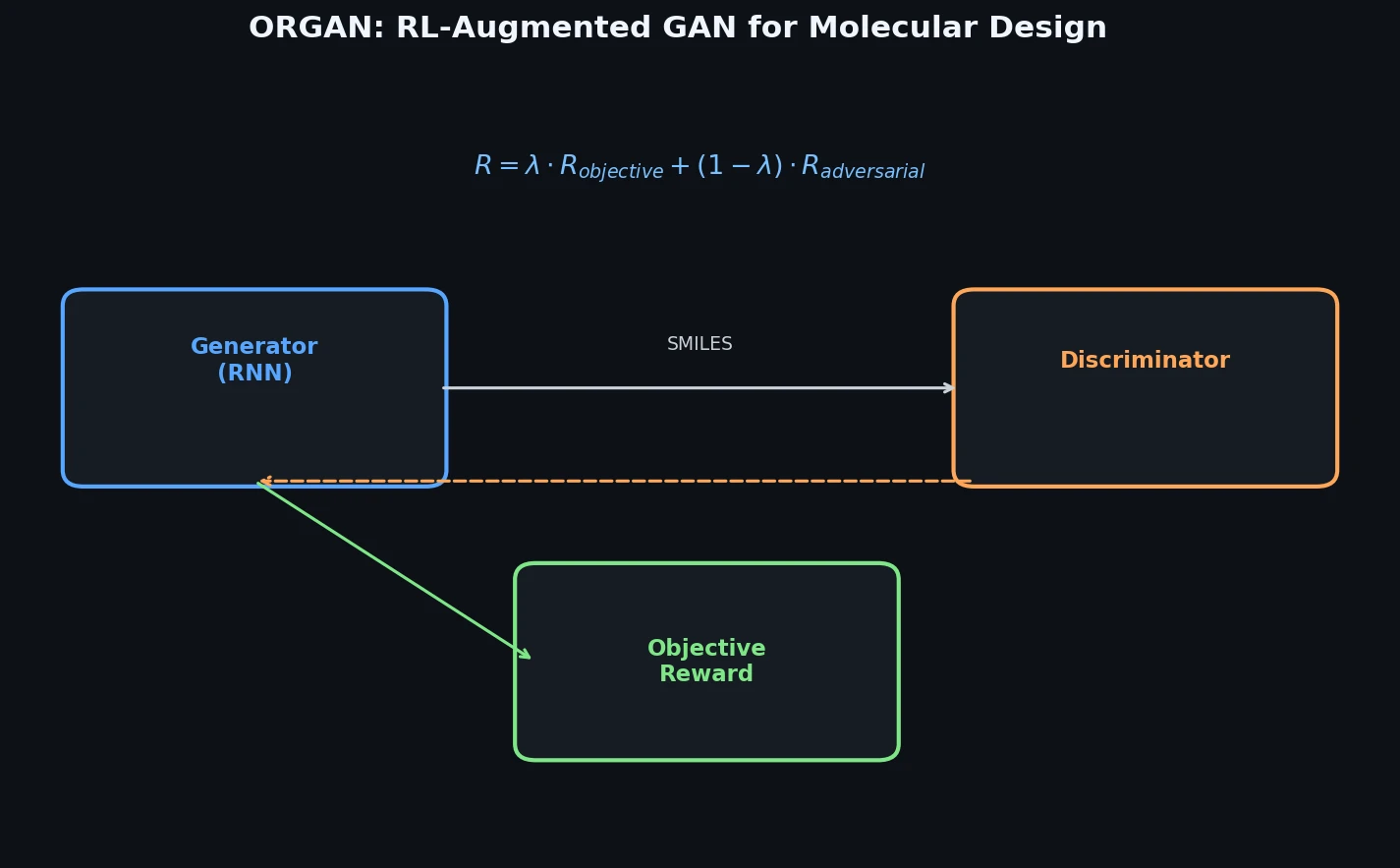
This is a Method paper that introduces ORGAN (Objective-Reinforced Generative Adversarial Network), a framework for generating sequences that are both realistic (close to the training distribution) and optimized for domain-specific objectives. ORGAN extends SeqGAN by adding external reward functions to the reinforcement learning signal, with a tunable parameter $\lambda$ controlling the balance between adversarial (discriminator) and objective-based rewards. The authors demonstrate ORGAN on two domains: molecular generation using SMILES strings (optimizing druglikeness, solubility, and synthesizability) and musical melody generation (optimizing tonality and step ratios).
Exposure Bias and Mode Collapse in Discrete Sequence Generation
Generating discrete sequences with desirable properties presents two intertwined challenges. First, RNNs trained via maximum likelihood estimation (MLE) suffer from exposure bias, where the model sees only ground-truth prefixes during training but must condition on its own (potentially erroneous) outputs at generation time. Second, while GANs can address some of these issues through adversarial training, they were not initially applicable to discrete data due to non-differentiability of the sampling step. SeqGAN resolved this by framing the generator as an RL agent, but it optimizes only for distributional fidelity (fooling the discriminator) without any mechanism to steer generation toward specific property targets.
In drug discovery, simply generating valid, drug-like molecules is insufficient. Practitioners need to optimize for particular pharmaceutical properties (e.g., solubility, synthesizability, druglikeness) while maintaining structural diversity. Naive RL approaches can optimize properties effectively but tend to collapse onto trivial solutions (e.g., repeating “CCCCCCC” to maximize solubility). The challenge is to combine the distributional regularization of adversarial training with the goal-directedness of RL.
Mixed Reward: Interpolating Between Adversarial and Objective Signals
ORGAN’s core innovation is a reward function that linearly interpolates between the discriminator score and domain-specific objectives:
$$R(Y_{1:T}) = \lambda \cdot D_{\phi}(Y_{1:T}) + (1 - \lambda) \cdot O_{i}(Y_{1:T})$$
When $\lambda = 1$, the model reduces to SeqGAN (pure adversarial training). When $\lambda = 0$, it becomes naive RL optimizing only the objective. Intermediate values allow the adversarial component to regularize the generator, keeping samples within the distribution while the objective component steers toward desired properties.
The generator $G_{\theta}$ is an LSTM-based RNN that produces sequences token-by-token. Training follows the REINFORCE algorithm, where the expected long-term reward is:
$$J(\theta) = \mathbb{E}\left[R(Y_{1:T}) \mid s_{0}, \theta\right] = \sum_{y_{1} \in Y} G_{\theta}(y_{1} \mid s_{0}) \cdot Q(s_{0}, y_{1})$$
For intermediate timesteps (partial sequences), the action-value function $Q$ is estimated via $N$-time Monte Carlo rollouts:
$$Q(Y_{1:t-1}, y_{t}) = \begin{cases} \frac{1}{N} \sum_{n=1}^{N} R(Y_{1:T}^{n}), & \text{if } t < T \\ R(Y_{1:T}), & \text{if } t = T \end{cases}$$
where $Y_{1:T}^{n}$ are completions sampled by rolling out the current policy $G_{\theta}$ from state $Y_{1:t}$.
The policy gradient is:
$$\nabla_{\theta} J(\theta) \simeq \frac{1}{T} \sum_{t=1}^{T} \mathbb{E}_{y_{t} \sim G_{\theta}(y_{t} \mid Y_{1:t-1})} \left[\nabla_{\theta} \log G_{\theta}(y_{t} \mid Y_{1:t-1}) \cdot Q(Y_{1:t-1}, y_{t})\right]$$
Two additional mechanisms improve training:
- Diversity penalty: Repeated sequences have their reward divided by their copy count, providing diminishing returns for non-unique outputs.
- Wasserstein distance: The authors also implement a variant (OR(W)GAN) that replaces the standard GAN discriminator loss with the Wasserstein-1 distance via Kantorovich-Rubinstein duality, which can improve training stability and diversity.
Molecular and Musical Melody Generation Experiments
Architecture
The generator $G_{\theta}$ is an RNN with LSTM cells. The discriminator $D_{\phi}$ is a CNN for text classification following Kim (2014), with 75% dropout and L2 regularization. All optimization uses Adam. Molecular metrics are computed with RDKit.
Molecular Generation Setup
Training data consists of 5,000 random molecules from the QM9 dataset (134k stable small molecules with up to 9 heavy atoms), encoded as SMILES strings with maximum sequence length 51 and alphabet size 43. Each generator is pre-trained for 250 MLE epochs, with the discriminator trained for 10 epochs. Adversarial/RL training then proceeds for up to 100 additional epochs. The default $\lambda$ is 0.5.
Three molecular objectives are evaluated:
- Solubility (LogP): water-octanol partition coefficient via RDKit’s Crippen function
- Synthesizability: SA score estimating ease of synthesis (0 = hard, 1 = easy)
- Druglikeness: QED score capturing medicinal chemistry aesthetics
Diversity is measured using average Jaccard distance of molecular fingerprints relative to a random training subset.
Molecular Generation Results
| Objective | Algorithm | Validity (%) | Diversity | Druglikeness | Synthesizability | Solubility |
|---|---|---|---|---|---|---|
| None | MLE | 75.9 | 0.64 | 0.48 (0%) | 0.23 (0%) | 0.30 (0%) |
| None | SeqGAN | 80.3 | 0.61 | 0.49 (+2%) | 0.25 (+6%) | 0.31 (+3%) |
| Druglikeness | ORGAN | 88.2 | 0.55 | 0.52 (+8%) | 0.32 (+38%) | 0.35 (+18%) |
| Druglikeness | OR(W)GAN | 85.0 | 0.95 | 0.60 (+25%) | 0.54 (+130%) | 0.47 (+57%) |
| Druglikeness | Naive RL | 97.1 | 0.80 | 0.57 (+19%) | 0.53 (+126%) | 0.50 (+67%) |
| Synthesizability | ORGAN | 96.5 | 0.92 | 0.51 (+6%) | 0.83 (+255%) | 0.45 (+52%) |
| Synthesizability | OR(W)GAN | 97.6 | 1.00 | 0.20 (-59%) | 0.75 (+223%) | 0.84 (+184%) |
| Solubility | ORGAN | 94.7 | 0.76 | 0.50 (+4%) | 0.63 (+171%) | 0.55 (+85%) |
| Solubility | OR(W)GAN | 94.1 | 0.90 | 0.42 (-12%) | 0.66 (+185%) | 0.54 (+81%) |
| Solubility | Naive RL | 92.7 | 0.75 | 0.49 (+3%) | 0.70 (+200%) | 0.78 (+162%) |
| All (alternated) | ORGAN | 96.1 | 92.3 | 0.52 (+9%) | 0.71 (+206%) | 0.53 (+79%) |
Key observations: OR(W)GAN consistently achieves higher diversity than standard ORGAN. Naive RL often achieves higher raw objective scores but at the cost of generating trivial solutions (e.g., simple atom chains for solubility). The Wasserstein variant provides better diversity properties. Multi-objective training via alternating objectives across epochs achieves gains comparable to individually optimized models.
Music Generation Setup
Using 1,000 melodies from the EsAC folk dataset, each encoded as 36-token sequences where tokens represent sixteenth-note events across three octaves (C3-B5). Two metrics are optimized: tonality (proportion of perfect fifths) and ratio of steps (conjunct melodic motion). Diversity is measured as average pairwise edit distance.
Music Results
| Objective | Algorithm | Diversity | Tonality | Ratio of Steps |
|---|---|---|---|---|
| None | MLE | 0.221 | 0.007 | 0.010 |
| None | SeqGAN | 0.187 | 0.005 | 0.010 |
| Tonality | Naive RL | 0.100 | 0.478 | 2.9E-05 |
| Tonality | ORGAN | 0.268 | 0.372 | 1.78E-04 |
| Tonality | OR(W)GAN | 0.268 | 0.177 | 2.4E-04 |
| Ratio of Steps | Naive RL | 0.321 | 0.001 | 0.829 |
| Ratio of Steps | ORGAN | 0.433 | 0.001 | 0.632 |
| Ratio of Steps | OR(W)GAN | 0.134 | 5.95E-05 | 0.622 |
ORGAN outperforms SeqGAN and MLE on all metrics. Naive RL achieves higher raw scores but with lower diversity, producing simpler, less interesting outputs.
Capacity Ceilings, Trade-offs, and Future Directions
The authors identify several limitations and findings:
Capacity ceiling: GAN-based models tend to generate sequences matching the training set’s average length (15.42 characters). RL-only approaches can break this constraint, generating shorter (9.4) or longer (21.3) sequences depending on the objective. The upper bound of optimized properties also matches the training data’s maximum, suggesting dataset-dependent limits.
Lambda trade-off: Varying $\lambda$ reveals an optimal balance between objective optimization and distributional fidelity. This optimum depends on the model, dataset, and metric, suggesting that hyperparameter search over $\lambda$ is important in practice.
Tonality vs. steps inverse relationship: In the music task, optimizing for tonality (perfect fifths) inherently conflicts with optimizing for step ratios (consecutive notes), since consecutive scale notes do not form perfect fifths.
Limitations: The paper evaluates on relatively small datasets (5k molecules, 1k melodies) and short sequences. The molecular experiments use QM9 (small molecules with up to 9 heavy atoms), which limits the scope of conclusions for drug-like chemical space. The Wasserstein variant sometimes lags behind the standard GAN loss in raw metric scores, though it offers better diversity.
Future directions: The authors propose extending ORGAN to non-sequential data (images, audio) by framing GANs as RL problems more broadly, and investigating how different heuristic choices affect performance. They also suggest exploring other discrete GAN formulations (MaliGAN, BGAN) with RL extensions.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Molecular training | QM9 subset | 5,000 molecules | Random subset from 134k stable small molecules with up to 9 heavy atoms |
| Music training | EsAC folk dataset | 1,000 melodies | 36-token sequences, processed following Chen et al. (2017) |
Algorithms
- Generator pre-trained for 250 epochs via MLE; discriminator for 10 epochs
- Adversarial/RL training for up to 100 epochs
- Default $\lambda = 0.5$ for reward mixing
- Monte Carlo rollouts for intermediate reward estimation
- Duplicate penalty: reward divided by copy count
Models
- Generator: RNN with LSTM cells
- Discriminator: CNN for text classification (Kim, 2014) with 75% dropout, L2 regularization
- Optimizer: Adam for all gradient descent steps
Evaluation
| Metric | Description | Domain |
|---|---|---|
| Validity (%) | Fraction of generated SMILES that decode to valid molecules | Molecules |
| Diversity | Average Jaccard distance of fingerprints to training subset | Molecules |
| Druglikeness (QED) | Quantitative Estimate of Drug-likeness | Molecules |
| Synthesizability (SA) | Synthetic accessibility score | Molecules |
| Solubility (LogP) | Water-octanol partition coefficient | Molecules |
| Tonality | Proportion of perfect fifths | Music |
| Ratio of Steps | Proportion of conjunct melodic intervals | Music |
| Diversity (edit) | Average pairwise edit distance | Music |
Hardware
Not specified in the paper.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| ORGAN | Code | GPL-2.0 | Official implementation including metrics for molecules and music |
Paper Information
Citation: Guimaraes, G. L., Sánchez-Lengeling, B., Outeiral, C., Farias, P. L. C., & Aspuru-Guzik, A. (2017). Objective-Reinforced Generative Adversarial Networks (ORGAN) for Sequence Generation Models. arXiv preprint arXiv:1705.10843.
@article{guimaraes2017organ,
title={Objective-Reinforced Generative Adversarial Networks (ORGAN) for Sequence Generation Models},
author={Guimaraes, Gabriel Lima and Sanchez-Lengeling, Benjamin and Outeiral, Carlos and Farias, Pedro Luis Cunha and Aspuru-Guzik, Al{\'a}n},
journal={arXiv preprint arXiv:1705.10843},
year={2017}
}