A Memory Module for Diverse Molecular Generation via RL
This is a Method paper that introduces a memory unit for reinforcement learning (RL)-based molecular generation. The primary contribution is a hash-table-based memory mechanism that integrates into the REINVENT framework’s scoring function. By tracking previously generated high-scoring molecules and penalizing the reward when new molecules are too similar to those already stored, the memory unit forces the generative model to explore different regions of chemical space rather than collapsing onto a single scaffold family.
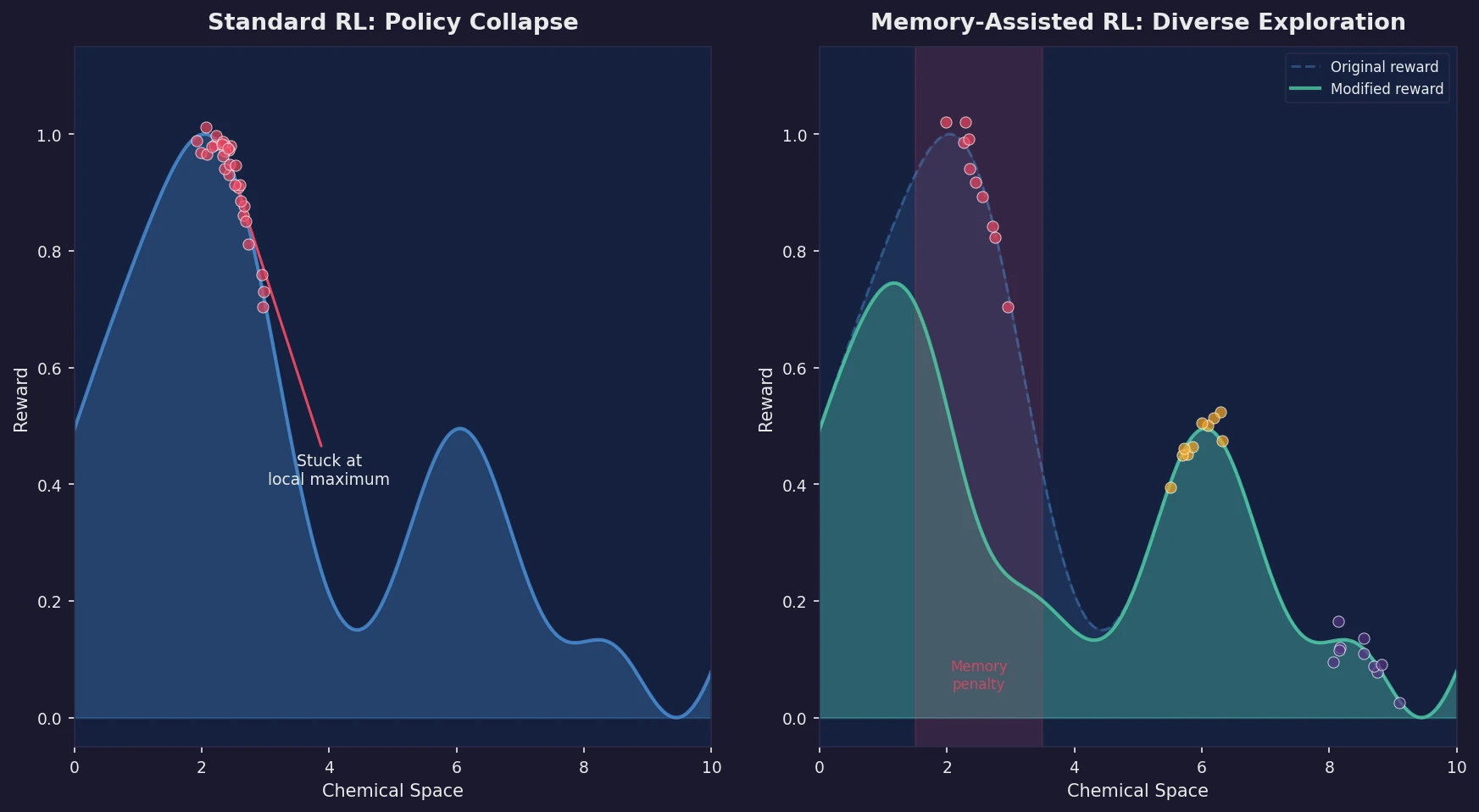
Policy Collapse Limits RL-Based De Novo Design
Recurrent neural networks (RNNs) trained with reinforcement learning can generate novel molecules optimized for desired properties. The REINVENT algorithm and related approaches (ORGANIC, GENTRL) demonstrated the viability of coupling a pretrained SMILES-based generative model with a scoring function via RL. However, a persistent problem is policy collapse (also called mode collapse): once the model discovers a high-scoring region of chemical space, it continues to exploit that region, producing structurally similar compounds with minor substitution differences. This severely limits the practical utility of RL-based generation in drug design, where medicinal chemists need diverse scaffolds to explore structure-activity relationships and manage intellectual property concerns.
Prior work by Liu et al. [31] attempted to address this by engineering an explorative RNN alongside the standard generative RNN, but it did not substantially increase diversity compared to standard REINVENT. Other approaches like Generative Examination Networks (GEN) performed statistical analysis during training but were not evaluated in optimization scenarios.
Core Innovation: Hash-Table Memory Unit for Reward Modification
The key insight is to dynamically modify the reward surface during RL by maintaining a memory of previously explored chemical space. The memory unit is a hash table of index-bucket pairs. Each bucket stores up to a fixed number of high-scoring molecules (default: 25) that are chemically similar to a seed molecule (the index).
Integration with REINVENT
The memory unit modifies the augmented likelihood used in REINVENT. For a generated compound $c$, the augmented log-likelihood becomes:
$$ \log P(c)_{Aug} = \log P(c)_{PriorNetwork} + \sigma \times S(c) \times M(c) $$
where $\sigma$ is a scalar coefficient, $S(c)$ is the scoring function output, and $M(c)$ is the memory unit output (either 0 or 1). The reward is:
$$ R(c) = \left(\log P(c)_{Aug} - \log P(c)_{AgentNetwork}\right)^2 $$
and the loss is $\text{loss} = -R(c)$.
Memory Unit Operation
When a high-scoring molecule is generated:
- Its fingerprint or scaffold is compared against all index structures in the memory
- If it is similar to an index (above a Tanimoto cutoff, default 0.6) and the corresponding bucket is not full, $M(c) = 1$ and the molecule is added to the bucket
- If the bucket is full, $M(c) = 0$, effectively zeroing the reward contribution and discouraging the model from generating similar molecules
- If no similar index exists, a new index-bucket pair is created
Four Similarity Criteria
The authors evaluate four criteria for grouping molecules in the memory:
- Compound similarity: ECFP4 Tanimoto similarity at the whole-molecule level
- Identical Bemis-Murcko (BM) scaffold: exact match of Bemis-Murcko frameworks
- Identical carbon skeleton: exact match of carbon skeletons (BM scaffolds with all heteroatoms replaced by carbon and bonds set to single)
- Scaffold similarity: atom pair fingerprint Tanimoto similarity between carbon skeletons (fuzzy matching)
Alternative Output Modes
Beyond the binary output ($M(c) \in {0, 1}$), the authors also explored smooth output functions. The linear mode:
$$ M(c) = 1 - \frac{\text{compounds in bucket}}{\text{bucket size}} $$
And the sigmoid mode:
$$ M(c) = 1 - \frac{1}{1 + e^{-\left(\frac{\frac{\text{compounds in bucket}}{\text{bucket size}} \times 2 - 1}{0.15}\right)}} $$
Both smooth modes yielded slightly fewer analogs than the binary mode and were not pursued further.
Experimental Setup: LogP Optimization and Target Activity Prediction
Case Study 1: LogP Optimization
As a proof of concept, the authors optimized LogP values for known DRD2 inhibitors. Starting from 487 DRD2 compounds with LogP >= 5 (from ExCAPE-DB), they applied transfer learning to the prior model for 20 epochs, then ran RL for 150 iterations (100 compounds per iteration, 15,000 total). The scoring function was:
$$ S = 1 - \tanh\left(\min\left(|2 - \text{AlogP}|, |3 - \text{AlogP}|\right)\right) $$
targeting LogP values between 2.0 and 3.0.
Case Study 2: HTR1A and DRD2 Activity Prediction
For a more complex scenario, the authors trained SVM classifiers (with Platt scaling for probabilistic output) on bioactivity data from ExCAPE-DB to predict activity against two neurotransmitter receptors:
- HTR1A: 3,599 actives (pIC50 >= 7) and 66,684 inactives
- DRD2: 2,981 actives (pIC50 >= 7) and 346,206 inactives (100,000 sampled)
Data was split using Butina clustering on ECFP6 at a 0.4 Tanimoto cutoff (60/20/20 train/val/test). The SVM models achieved excellent performance:
| Target | Set | Balanced Accuracy | ROC AUC | F1 | MCC |
|---|---|---|---|---|---|
| HTR1A | Test | 0.96 | 0.99 | 0.75 | 0.75 |
| DRD2 | Test | 0.95 | 0.99 | 0.71 | 0.72 |
RL was run for 300 iterations (100 compounds each, 30,000 total). Compounds with predicted activity >= 0.7 were considered active.
Generative Model Architecture
The RNN prior model followed the REINVENT architecture: an embedding layer, three GRU layers with 256 dimensions, and a linear output layer. It was pretrained on ~1.5 million ChEMBL 25 compounds (filtered to remove known HTR1A actives and DRD2 analogs) for 10 epochs using Adam with a learning rate of 0.01.
Comparisons
The authors compared memory-assisted RL against:
- Standard REINVENT RL (no memory)
- Experience replay (re-presenting 8 high-scoring compounds per iteration)
- Temperature scaling (values from 1.0 to 10.0)
- Memory + experience replay combined
Results: Up to Fourfold Increase in Diverse Active Compounds
LogP Optimization Results
Memory-assisted RL increased the number of optimized compounds (LogP 2-3) by roughly threefold:
| Memory Type | Optimized Compounds | Unique BM Scaffolds | Unique Carbon Skeletons |
|---|---|---|---|
| No memory | 938 | 727 | 396 |
| Compound similarity | 3,451 | 2,963 | 1,472 |
| Identical BM Scaffold | 3,428 | 2,865 | 1,398 |
| Identical Carbon Skeleton | 3,315 | 3,002 | 1,799 |
| Scaffold Similarity | 3,591 | 3,056 | 1,538 |
The memory unit also increased the generation of relevant analogs. ECFP6 analogs (Tanimoto >= 0.4 to training set) increased from 145 to up to 549, and shared MMP cores increased from 5 to up to 19, confirming that the memory unit promoted exploration of chemically relevant space rather than random drift.
HTR1A and DRD2 Activity Optimization Results
The improvements were even more pronounced for target activity optimization:
| Target | Memory Type | Active Compounds | Unique BM Scaffolds | Unique Carbon Skeletons |
|---|---|---|---|---|
| HTR1A | No memory | 9,323 | 7,312 | 5,446 |
| HTR1A | Compound similarity | 16,779 | 13,304 | 9,887 |
| HTR1A | Identical Carbon Skeleton | 17,597 | 15,531 | 12,408 |
| DRD2 | No memory | 5,143 | 2,635 | 1,949 |
| DRD2 | Compound similarity | 21,486 | 17,844 | 12,749 |
| DRD2 | Scaffold Similarity | 22,784 | 20,712 | 16,434 |
For DRD2, the effect was particularly striking: standard RL showed clear policy collapse with only 576 ECFP6 analogs to the training set, while memory-assisted RL generated up to 6,315. The compound similarity memory unit produced the most MMP analogs (217 to the training set vs. 7 without memory).
Parameter Sensitivity
Bucket size had a modest effect: larger buckets (allowing more compounds before penalization) slightly increased analog generation. The Tanimoto similarity threshold of 0.6 was near-optimal for the scaffold similarity memory; higher thresholds reduced diversity gains. The compound similarity memory showed increasing analogs with higher thresholds, but BM scaffold and carbon skeleton counts plateaued above 0.6.
Comparison with Experience Replay and Temperature Scaling
- Experience replay alone increased diversity compared to vanilla RL but was less effective than the memory unit alone
- Memory + experience replay achieved the best results overall, as experience replay provided the model with diverse starting points for exploration after the memory unit altered the reward landscape
- Temperature scaling was largely ineffective: only a value of 1.25 showed improvement, and even then it achieved only about 50% of the analogs generated by memory-assisted RL. Temperatures above 2.0 degraded SMILES validity, and above 4.0 prevented valid molecule generation entirely
Limitations
The authors acknowledge several limitations:
- All evaluations are retrospective; no synthesized compounds were experimentally tested
- The SVM activity models, while accurate, may have applicability domain limitations for highly novel scaffolds
- The binary memory output mode was found to work best, but the transition from exploration to exploitation is abrupt
- The method was only tested with two biological targets and one physicochemical property
- Computational overhead of the memory unit is not discussed
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Prior model training | ChEMBL 25 | ~1.5M compounds | Filtered: max 50 heavy atoms, no stereochemistry, removed HTR1A actives and DRD2 analogs |
| HTR1A activity data | ExCAPE-DB | 3,599 actives + 66,684 inactives | pIC50 >= 7 threshold for actives |
| DRD2 activity data | ExCAPE-DB | 2,981 actives + 100,000 inactives (sampled) | pIC50 >= 7 threshold for actives |
Algorithms
- Generative model: RNN with embedding + 3 GRU layers (256 dim) + linear output (REINVENT architecture)
- RL: Augmented likelihood formulation with sigma scaling coefficient
- SVM classifiers: Non-linear SVM with MinMax kernel, Platt scaling, ECFP6 count-based fingerprints (2048 dim)
- Butina clustering: ECFP6 Tanimoto cutoff 0.4 for train/val/test splitting
Evaluation
| Metric | Description |
|---|---|
| Unique compounds | Number of distinct valid SMILES generated |
| Unique BM scaffolds | Bemis-Murcko framework diversity |
| Unique carbon skeletons | Carbon skeleton diversity (stripped BM scaffolds) |
| ECFP6 analogs | Compounds with Tanimoto >= 0.4 to known actives |
| MMP analogs | Matched molecular pair relationships with known actives |
| Shared MMP cores | Scaffold cores shared between generated and known compounds |
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| reinvent-memory | Code | MIT | Official implementation with prepared datasets |
Hardware
Not specified in the paper.
Paper Information
Citation: Blaschke, T., Engkvist, O., Bajorath, J., & Chen, H. (2020). Memory-assisted reinforcement learning for diverse molecular de novo design. Journal of Cheminformatics, 12, 68. https://doi.org/10.1186/s13321-020-00473-0
@article{blaschke2020memory,
title={Memory-assisted reinforcement learning for diverse molecular de novo design},
author={Blaschke, Thomas and Engkvist, Ola and Bajorath, J{\"u}rgen and Chen, Hongming},
journal={Journal of Cheminformatics},
volume={12},
number={1},
pages={68},
year={2020},
publisher={Springer},
doi={10.1186/s13321-020-00473-0}
}