A Method for Generative Linker Design with Reinforcement Learning
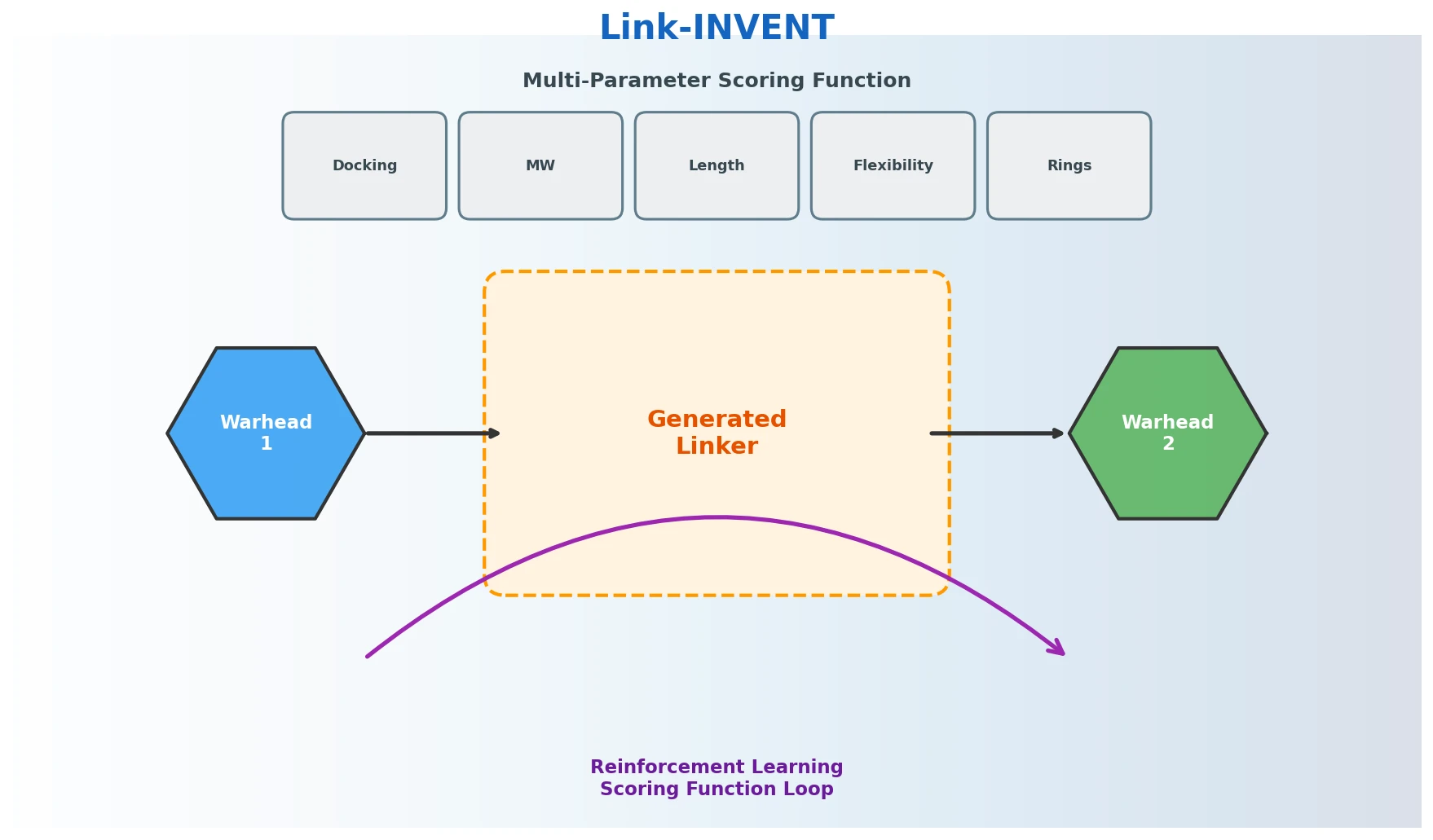
Link-INVENT is a Method paper that introduces a generative model for molecular linker design built on the REINVENT de novo design platform. The primary contribution is an encoder-decoder recurrent neural network (RNN) architecture that generates SMILES-based linkers connecting two molecular subunits, combined with a flexible multi-parameter optimization (MPO) scoring function and reinforcement learning (RL) to steer generation toward desired properties. Link-INVENT targets three practical drug discovery tasks: fragment linking, scaffold hopping, and proteolysis targeting chimera (PROTAC) design.
Why Linker Design Needs Flexible Multi-Parameter Optimization
Generating suitable chemical linkers between molecular subunits is a central challenge in fragment-based drug discovery (FBDD), scaffold hopping, and PROTAC design. Traditional computational approaches rely on database searches, inherently limiting the generalizability of proposed linkers to the pre-defined collection. Recent deep learning methods (DeLinker, SyntaLinker, 3DLinker, DiffLinker) can generate novel linkers but offer limited support for optimizing specific physicochemical properties. Users can typically control only linker length and a few properties like hydrogen-bond donor count.
The key gaps that Link-INVENT addresses are:
- Conditioning on both subunits: Prior RNN-based approaches (SAMOA) generate linkers conditioned only on the SMILES sequence seen so far, which may not account for the second molecular subunit. Link-INVENT conditions on both warheads simultaneously.
- Flexible scoring: Existing DL-based linker design tools lack the ability to define tailored MPO objectives. Link-INVENT inherits REINVENT 4’s full scoring infrastructure and adds linker-specific properties.
- Generalizability: A single trained prior handles fragment linking, scaffold hopping, and PROTAC tasks without retraining.
Core Innovation: Conditional Linker Generation with Augmented Likelihood RL
Link-INVENT’s architecture is an encoder-decoder RNN adapted from the Lib-INVENT library design model. The encoder processes a pair of warheads (molecular subunits with defined exit vectors), and the decoder generates a linker token by token, yielding a connected molecule in SMILES format. The model uses three hidden layers of 512 LSTM cells with an embedding size of 256.
Training
The prior is trained on ChEMBL v27 data processed through reaction-based slicing to generate (linker, warheads pair, full molecule) tuples. SMILES randomization augments the training data at each epoch, improving chemical space generalizability. The prior is trained by maximizing the likelihood of generating a linker conditioned on the input warhead pair, with teacher forcing for stability.
Multi-Parameter Optimization via RL
The scoring function $S(x)$ is a weighted geometric mean of individual component scores:
$$ S(x) = \left(\prod_{i=1}^{n} C_{i}(x)^{w_{i}}\right)^{\frac{1}{\sum_{i=1}^{n} w_{i}}} $$
where $x$ is a sampled linked molecule, $C_{i}(x)$ is the score for the $i$-th component, and $w_{i}$ is its weight.
The agent (initialized as a copy of the prior) is updated via the Difference of Augmented and Posterior likelihoods (DAP) loss. The augmented log likelihood is:
$$ \log \pi_{\text{augmented}} = \log \pi_{\text{prior}} + \sigma \cdot S(x) $$
where $\pi$ denotes a policy (token sampling probabilities conditioned on the sequence so far) and $\sigma$ is a scalar factor. The loss function is:
$$ J(\theta) = \left(\log \pi_{\text{augmented}} - \log \pi_{\text{agent}}\right)^{2} $$
Minimizing $J(\theta)$ steers the agent to generate molecules that satisfy the scoring function while remaining anchored to the prior’s chemical space.
Diversity Filters
Link-INVENT uses Diversity Filters (DFs) to balance exploration and exploitation. Buckets of limited size track unique Bemis-Murcko scaffolds. When a bucket is full, further sampling of that scaffold receives a score of zero, encouraging the agent to explore diverse chemical space regions.
Linker-Specific Scoring Components
New scoring components provide direct control over linker properties:
- Linker effective length: number of bonds between attachment atoms
- Linker maximum graph length: bonds in the longest graph traversal path
- Linker length ratio: effective length divided by maximum graph length (controls branching)
- Linker ratio of rotatable bonds: rotatable bonds over total bonds (controls flexibility)
- Linker number of rings: controls linearity vs. cyclicity
- Linker number of HBDs: hydrogen-bond donors in the linker itself
Experimental Evaluation Across Three Drug Discovery Tasks
Link-INVENT was evaluated through four experiments across three drug discovery applications, all using the same pre-trained prior.
Illustrative Example: Two Benzene Rings
A simple experiment linked two benzene rings with the objectives of limiting HBDs and requiring exactly one ring in the linker. Over 20 epochs, the agent learned to satisfy both objectives, demonstrating the basic RL-guided generation process.
Experiment 1a: Fragment Linking (CK2 alpha Inhibitors)
Based on the casein kinase 2 (CK2 alpha) fragment linking campaign by Fusco and Brear et al., Link-INVENT was tasked with linking two fragment hits while retaining the Lys68 hydrogen-bond interaction via a DockStream docking constraint (Glide/LigPrep backend). The scoring function also enforced linker length ratio >= 70 and linker MW <= 200 Da.
Over 100 epochs in triplicate, the agent generated molecules with gradually improving docking scores. Key results:
- Docking score distributions across triplicates were nearly identical, demonstrating reproducibility
- Some generated molecules achieved more favorable docking scores than the reference ligand CAM4066 (-15.20 kcal/mol)
- More than 5000 unique Bemis-Murcko scaffolds were generated, with minimal overlap across replicates
- Binding pose analysis showed the generated linker closely resembled the ground-truth linker, retaining the Lys68 interaction
Experiment 1b: Comparison Fragment Linking (IMPDH Inhibitors)
Using the IMPDH inhibitor fragment linking case study from Trapero et al., this experiment applied core constrained docking (fragment pose within 0.3 A of reference) and compared results to DeLinker and SyntaLinker. The scoring function enforced linker effective length in [3, 5], length ratio >= 70, and linker MW <= 150 Da.
Link-INVENT generated 8960 SMILES across 70 epochs (comparable to DeLinker’s 9000 molecular graphs). Results:
- Link-INVENT generated molecules with more favorable docking scores than the reference ligand across triplicate runs
- Of 20 DeLinker and 3 SyntaLinker example molecules, none and one (the recovered reference) docked better than or equal to the reference
- Approximately 3000 unique Bemis-Murcko scaffolds were generated from 5000 total molecules
- Link-INVENT’s advantage comes from including docking explicitly as a learning objective rather than applying it post hoc
Experiment 2: Scaffold Hopping (DLK Inhibitor CNS Optimization)
Based on Patel et al.’s dual leucine zipper kinase (DLK) inhibitor campaign, Link-INVENT generated new scaffold ideas to improve CNS penetration while retaining potency. The scoring function included a Cys193 docking constraint plus CNS-compatible properties (HBDs < 2, tPSA <= 90 A squared, 3 <= SlogP <= 4, MW <= 450 Da, 1-2 aromatic rings in linker).
The solution space was significantly narrower than fragment linking. The agent still generated diverse scaffolds with favorable docking scores, though fewer exceeded the reference ligand’s score. Binding pose analysis confirmed retained Cys193 interactions and predicted additional Gln195 hydrogen bonds.
Experiment 3: PROTAC Design (Bcl-2/Mcl-1 Dual Degradation)
Three sub-experiments demonstrated linker-specific scoring components for PROTAC design based on Wang et al.’s Bcl-2/Mcl-1 dual degradation strategy:
| Sub-Experiment | Objective | Key Finding |
|---|---|---|
| Sub-Exp 1: Linker length | Generate linkers within specified length intervals [4,6], [7,9], [10,12], [13,15] | Clear enrichment within target intervals vs. baseline broad distribution |
| Sub-Exp 2: Linearity | Control linear vs. cyclic linkers at fixed length [7,9] | Baseline ratio ~1:2 linear:cyclic; enforcing linearity or cyclicity achieved strong enrichment |
| Sub-Exp 3: Flexibility | Generate linkers with Low [0,30], Moderate [40,60], or High [70,100] rotatable bond ratios | Agent learned that rings and sp2 atoms yield rigidity; linear sp3 chains yield flexibility |
Key Findings and Practical Implications for Drug Discovery
Link-INVENT demonstrates several practical advantages for molecular linker design:
- Single prior, multiple tasks: The same pre-trained model handles fragment linking, scaffold hopping, and PROTAC design without retraining.
- Docking as a learning signal: Including molecular docking explicitly in the scoring function (via DockStream) during RL yields molecules with more favorable docking scores than approaches that apply docking post hoc.
- Implicit 3D awareness: The docking constraint guides the agent toward 3D structural awareness without explicit 3D coordinate inputs, as demonstrated by the overlap between generated and reference binding poses.
- Diverse and reproducible output: Diversity filters ensure exploration of multiple chemical space regions, and triplicate experiments show consistent docking score distributions with minimal scaffold overlap.
Limitations acknowledged by the authors include:
- The linker flexibility metric (ratio of rotatable bonds) is agnostic to intra-molecular hydrogen bonds and does not account for all rigidity factors
- Molecular docking is an approximation that can be exploited (e.g., excessive HBDs achieving favorable scores at the expense of permeability)
- Experiments 1a and 1b require a proprietary Schrodinger license for Glide/LigPrep docking
- No direct experimental (wet-lab) validation was performed in this study
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Prior training | ChEMBL v27 (reaction-sliced) | Not specified | Filtered for drug-like compounds, then reaction-based slicing with SMIRKS |
| Validation | Held-out Bemis-Murcko scaffolds | 287 scaffolds | Held out from training set |
| SMILES augmentation | Randomized SMILES per epoch | Same tuples, different representations | Improves generalizability |
Algorithms
- Architecture: Encoder-decoder RNN with 3 hidden layers of 512 LSTM cells, embedding size 256
- RL loss: DAP (Difference of Augmented and Posterior likelihoods)
- Batch size: 128 molecules per epoch
- Diversity filter: Bemis-Murcko scaffold buckets of size 25
- Score threshold: 0 (to store all molecules for analysis)
- Scoring function: Weighted geometric mean of component scores
Models
- Single pre-trained prior used across all experiments
- Agent initialized as copy of prior, updated via RL
- Pre-trained prior available at GitHub repository
Evaluation
- Molecular docking via DockStream with Glide/LigPrep backend
- Triplicate runs for all experiments
- Metrics: docking scores, unique Bemis-Murcko scaffold counts, binding pose overlap
Hardware
Hardware specifications are not reported in the paper.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| REINVENT (Link-INVENT code) | Code | Apache-2.0 | Main codebase for Link-INVENT |
| ReinventCommunity (data + tutorial) | Code + Data | MIT | Training/validation data, reaction SMIRKS, pre-trained prior, Jupyter tutorial |
Reproducibility status: Partially Reproducible. Code, training data, and pre-trained prior are publicly available. However, reproducing the docking-based experiments (1a, 1b, and 2) requires a proprietary Schrodinger license for Glide and LigPrep. The PROTAC experiments (Experiment 3) that use only physicochemical scoring are fully reproducible with the open-source code.
Paper Information
Citation: Guo, J., Knuth, F., Margreitter, C., Janet, J. P., Papadopoulos, K., Engkvist, O., & Patronov, A. (2023). Link-INVENT: generative linker design with reinforcement learning. Digital Discovery, 2, 392-408. https://doi.org/10.1039/D2DD00115B
@article{guo2023link,
title={Link-INVENT: generative linker design with reinforcement learning},
author={Guo, Jeff and Knuth, Franziska and Margreitter, Christian and Janet, Jon Paul and Papadopoulos, Kostas and Engkvist, Ola and Patronov, Atanas},
journal={Digital Discovery},
volume={2},
number={2},
pages={392--408},
year={2023},
publisher={Royal Society of Chemistry},
doi={10.1039/D2DD00115B}
}