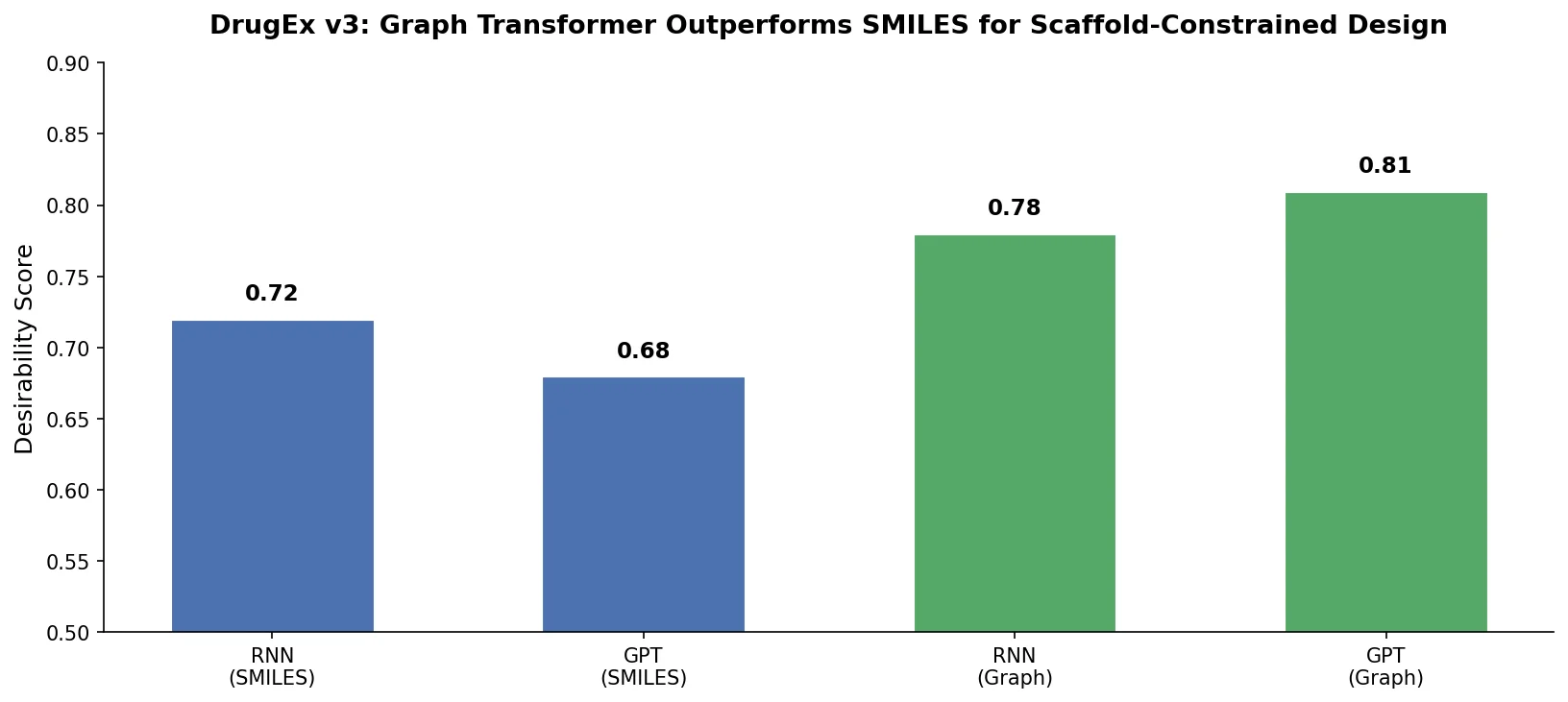
A Graph Transformer Method for Scaffold-Constrained Drug Design
This is a Method paper that introduces DrugEx v3, a Graph Transformer model for scaffold-constrained de novo drug design. The primary contribution is a novel positional encoding scheme for molecular graphs that allows a Transformer architecture to operate on graph-structured molecular data rather than SMILES strings. The model takes user-provided scaffold fragments as input and generates complete molecules through growing and connecting operations, trained with multi-objective reinforcement learning to optimize for both target affinity and drug-likeness.
From Fixed Objectives to User-Guided Scaffold Design
Prior versions of DrugEx (v1 and v2) used RNN-based generators trained with reinforcement learning for de novo drug design, but they operated under fixed objectives and could not accept user-provided structural priors. If a medicinal chemist wanted to explore analogs of a specific scaffold, the model needed retraining from scratch. Meanwhile, SMILES-based molecular generators face inherent limitations for scaffold-constrained design: SMILES is a linear notation, so inserting fragments at multiple positions of a scaffold requires complex grammar handling, and small token changes can produce invalid molecules.
Several approaches had been proposed for scaffold-based generation, including graph generative models (Lim et al., 2019), DeepScaffold (Li et al., 2020), SMILES-based scaffold decorators (Arus-Pous et al., 2020), and SyntaLinker for fragment linking (Yang et al., 2020). DrugEx v3 aims to combine the advantages of graph representations (validity guarantees, local invariance, flexible extension) with the Transformer architecture’s ability to handle complex dependencies, while maintaining the multi-objective reinforcement learning framework from DrugEx v2.
Graph Positional Encoding for Molecular Transformers
The core innovation is adapting the Transformer architecture to work directly with molecular graph representations. Two key modifications make this possible.
Graph word encoding. Since atoms and bonds cannot be processed simultaneously in a graph, the authors combine them into a single index:
$$ W = T_{atom} \times 4 + T_{bond} $$
where $T_{atom}$ is the atom type index and $T_{bond}$ is the bond type index (four bond types: single, double, triple, and none).
Graph positional encoding. Standard sequential position encoding does not capture molecular topology. The authors propose an adjacency-matrix-based positional encoding:
$$ P = I_{Atom} \times L_{max} + I_{Connected} $$
where $I_{Atom}$ is the current atom index, $L_{max}$ is the maximum sequence length, and $I_{Connected}$ is the index of the atom connected by the current bond. This encoding is then processed through the standard sinusoidal positional encoding:
$$ PE_{(p, 2i)} = \sin(pos / 10000^{2i / d_{m}}) $$
$$ PE_{(p, 2i+1)} = \cos(pos / 10000^{2i / d_{m}}) $$
with $d_{m} = 512$.
Molecule generation procedure. Each molecule in the training data is represented as a five-row matrix encoding atom type, bond type, connected atom index, current atom index, and fragment index. The columns are divided into three sections: fragment (the scaffold), growing (new atoms added to fragments), and linking (bonds connecting grown fragments). The decoder uses a GRU-based recurrent layer to sequentially output atom type, bond type, connected atom index, and current atom index at each step, with chemical valence rules enforced at every generation step to guarantee valid molecules.
Multi-objective reinforcement learning. The generator is trained with a policy gradient objective:
$$ J(\theta) = \mathbb{E}\left[R^{*}(y_{1:T}) | \theta\right] = \sum_{t=1}^{T} \log G(y_{t} | y_{1:t-1}) \cdot R^{\ast}(y_{1:T}) $$
where $R^{*}$ is a Pareto-based reward combining target affinity and QED drug-likeness score:
$$ R^{*} = \begin{cases} 0.5 + \frac{k - N_{undesired}}{2N_{desired}}, & \text{if desired} \\ \frac{k}{2N_{undesired}}, & \text{if undesired} \end{cases} $$
with $k$ being the solution’s index in the Pareto rank. An exploration strategy uses two networks: an exploitation network $G_{\theta}$ (updated by policy gradient) and an exploration network $G_{\phi}$ (fixed, pre-trained on ChEMBL), with an exploration rate $\varepsilon$ controlling how many scaffolds are routed to $G_{\phi}$ during training.
Experimental Setup: Architecture Comparison and RL Optimization
Data
The ChEMBL set (version 27) contained approximately 1.7 million molecules for pre-training, preprocessed via RDKit (charge neutralization, metal/fragment removal). The LIGAND set comprised 10,828 adenosine receptor ligands for fine-tuning. Each molecule was decomposed into fragments using the BRICS algorithm, creating scaffold-molecule pairs (up to 15 pairs per molecule with four fragments). The ChEMBL set yielded 9.3 million training pairs, and the LIGAND set produced 53,888 training pairs.
Architecture comparison
Four architectures were compared:
- Graph Transformer: graph input with novel positional encoding
- Sequential Transformer: SMILES input with standard Transformer
- LSTM-BASE: SMILES encoder-decoder with three recurrent layers
- LSTM+ATTN: LSTM-BASE with an attention mechanism between encoder and decoder
All models were pre-trained on ChEMBL and fine-tuned on the LIGAND set. The bioactivity predictor was a random forest regression model using 2048D ECFP6 fingerprints and 19D physicochemical descriptors, with an activity threshold of pX = 6.5 for the A2A adenosine receptor.
Evaluation metrics
Five metrics were used: validity (parseable molecules), accuracy (scaffold containment), desirability (meeting all objectives), uniqueness, and novelty (not in ChEMBL). Diversity was measured using the Solow-Polasky index with Tanimoto distance on ECFP6 fingerprints:
$$ I(A) = \frac{1}{|A|} \mathbf{e}^{\intercal} F(\mathbf{s})^{-1} \mathbf{e} $$
Hardware
Models were benchmarked on a server with NVIDIA Tesla P100 GPUs.
Key Results: Graph Representation Advantages and RL Trade-offs
Pre-training and fine-tuning performance
The Graph Transformer achieved the best overall performance across all metrics:
| Method | Validity (PT) | Accuracy (PT) | Validity (FT) | Accuracy (FT) | Novelty (FT) | Uniqueness (FT) |
|---|---|---|---|---|---|---|
| Graph Transformer (512) | 100.0% | 99.3% | 100.0% | 99.2% | 68.9% | 82.9% |
| Seq. Transformer (512) | 96.7% | 74.0% | 99.3% | 92.7% | 8.9% | 28.9% |
| LSTM+ATTN (512) | 94.3% | 72.8% | 96.9% | 85.9% | 6.3% | 20.7% |
| LSTM-BASE (512) | 93.9% | 52.4% | 98.7% | 81.6% | 3.9% | 19.2% |
PT = pre-trained, FT = fine-tuned. The Graph Transformer achieved 100% validity due to its explicit valence checking at each generation step. It also produced substantially more novel and unique molecules after fine-tuning compared to SMILES-based methods.
The authors identified four advantages of the graph representation over SMILES: (1) local invariance, where fragment ordering does not affect output; (2) global extendibility, where new atoms can be appended without restructuring existing data; (3) freedom from grammar constraints; and (4) direct accessibility of chemical valence rules for validity enforcement.
Reinforcement learning results
With multi-objective RL (affinity + QED), 74.6% of generated molecules were predicted active at $\varepsilon = 0.0$. The exploration rate $\varepsilon$ trades off desirability against uniqueness:
| $\varepsilon$ | Desirability | Uniqueness | Novelty | Diversity |
|---|---|---|---|---|
| 0.0 | 74.6% | 60.7% | 60.6% | 0.879 |
| 0.1 | 66.8% | 75.0% | 74.6% | 0.842 |
| 0.2 | 61.6% | 80.2% | 79.4% | 0.879 |
| 0.3 | 56.8% | 89.8% | 88.8% | 0.874 |
The authors report that $\varepsilon = 0.3$ produced the best balance between desirability and uniqueness, with 56.8% desired molecules and 89.8% uniqueness. Diversity remained above 0.84 across all settings.
Limitations
The Graph Transformer produced molecules with worse synthetic accessibility (SA scores) compared to SMILES-based methods, particularly after fine-tuning on the smaller LIGAND set. The authors attribute this to uncommon ring systems generated when the model handles long-distance dependencies. A kekulization issue also causes a small fraction of molecules to fail scaffold matching: aromatic bond inference during sanitization can alter the scaffold substructure. Without single-objective affinity constraint, the model generates molecules with molecular weight exceeding 500 Da, reducing drug-likeness. All bioactivity predictions rely on a random forest model rather than experimental validation, and the t-SNE analysis suggests some generated molecules fall outside the model’s applicability domain.
Future directions
The authors propose extending the Graph Transformer to accept protein information as input via proteochemometric modeling, enabling design of ligands for targets without known ligands. Lead optimization, where a “hit” serves as input to generate improved analogs, is also identified as a natural extension.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Pre-training | ChEMBL v27 | ~1.7M molecules (9.3M scaffold-molecule pairs) | Preprocessed via RDKit |
| Fine-tuning | LIGAND set (A2A AR ligands from ChEMBL) | 10,828 ligands (53,888 pairs) | Split 8:1:1 train/val/test |
| Bioactivity labels | ChEMBL A2A AR activity data | pX threshold = 6.5 | Average pChEMBL values |
Algorithms
- Fragment decomposition: BRICS algorithm via RDKit (max 4 fragments per molecule)
- Optimizer: Adam with learning rate $10^{-4}$, batch size 256
- Pre-training: 20 epochs; fine-tuning: up to 1,000 epochs with early stopping (patience: 100 epochs)
- Bioactivity predictor: random forest regression (scikit-learn) with 2048D ECFP6 + 19D physicochemical descriptors
- Pareto-based multi-objective ranking with GPU acceleration
Models
- Graph Transformer: 512 hidden units, 8 attention heads, $d_{k} = d_{v} = 64$
- Sequential Transformer: same hidden size, sinusoidal positional encoding
- LSTM-BASE / LSTM+ATTN: 128 embedding units, 512 hidden units, 3 recurrent layers
Evaluation
| Metric | Graph Transformer | Best SMILES Baseline | Notes |
|---|---|---|---|
| Validity (fine-tuned) | 100.0% | 99.6% (LSTM-BASE 1024) | Valence checking guarantees validity |
| Accuracy (fine-tuned) | 99.2% | 94.3% (Seq. Transformer 1024) | Scaffold containment |
| Desirability (RL, $\varepsilon$=0.0) | 74.6% | N/A | Only Graph Transformer used for RL |
| Diversity (RL) | 0.879 | N/A | Solow-Polasky index |
Hardware
NVIDIA Tesla P100 GPUs. Specific training times not reported, but Transformer models trained faster than LSTM models with the same hidden layer size.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| CDDLeiden/DrugEx | Code | MIT | Official implementation (v1, v2, v3) |
| ChEMBL v27 | Dataset | CC-BY-SA 3.0 | Pre-training data source |
Paper Information
Citation: Liu, X., Ye, K., van Vlijmen, H. W. T., IJzerman, A. P., & van Westen, G. J. P. (2023). DrugEx v3: scaffold-constrained drug design with graph transformer-based reinforcement learning. Journal of Cheminformatics, 15, 24. https://doi.org/10.1186/s13321-023-00694-z
@article{liu2023drugex,
title={DrugEx v3: scaffold-constrained drug design with graph transformer-based reinforcement learning},
author={Liu, Xuhan and Ye, Kai and van Vlijmen, Herman W. T. and IJzerman, Adriaan P. and van Westen, Gerard J. P.},
journal={Journal of Cheminformatics},
volume={15},
number={1},
pages={24},
year={2023},
publisher={Springer},
doi={10.1186/s13321-023-00694-z}
}