A Hybrid RL Strategy for De Novo Molecule Generation
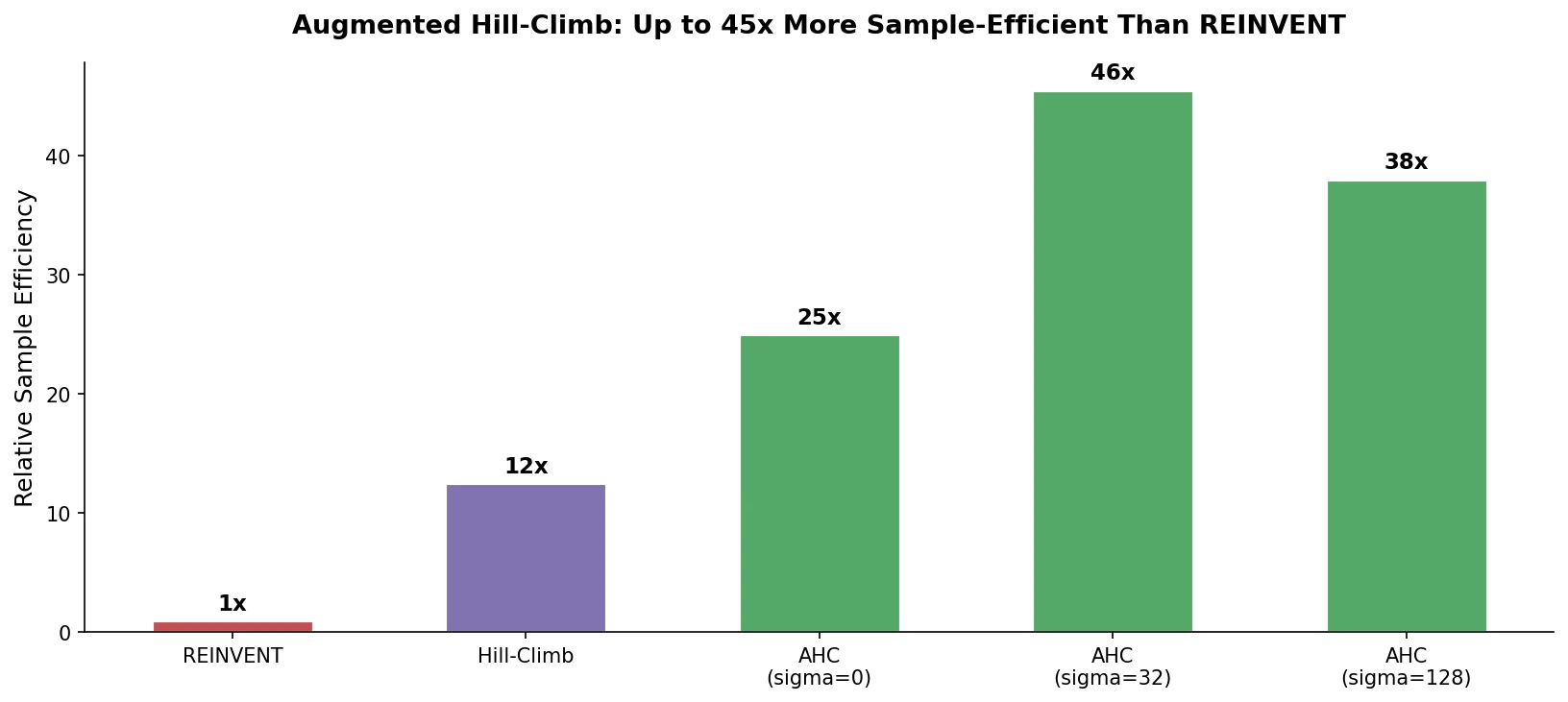
This is a Method paper that proposes Augmented Hill-Climb (AHC), a reinforcement learning strategy for conditioning SMILES-based language models during de novo molecule generation. The primary contribution is a simple hybrid between the REINVENT and Hill-Climb (HC) RL strategies that computes the REINVENT loss function only on the top-k highest-scoring molecules per batch (as in HC), thereby removing the counterproductive regularization effect of low-scoring molecules. The authors demonstrate that AHC improves optimization ability ~1.5-fold and sample efficiency ~45-fold compared to REINVENT across docking tasks against four GPCR targets, and that the approach generalizes to transformer architectures.
Sample Efficiency Bottleneck in RL-Guided Molecular Generation
Recurrent neural networks trained on SMILES have become a standard approach for de novo molecule generation, with RL strategies like REINVENT and Hill-Climb achieving top performance on benchmarks such as GuacaMol and MOSES. However, RL-guided generation can be highly sample-inefficient, often requiring $10^5$ or more molecules to optimize complex objectives. This is acceptable for cheap scoring functions (e.g., QSAR models, property calculators) but becomes a practical bottleneck when using computationally expensive scoring functions like molecular docking or computer-aided synthesis planning.
The REINVENT strategy regularizes the agent by computing a loss based on the difference between the agent’s policy and an “augmented likelihood” that combines the prior policy with a scaled reward. When low-scoring molecules are sampled ($R_T \approx 0$), the augmented likelihood reduces to the prior likelihood, causing the agent to trend back toward the prior policy. This negates useful learnings, especially early in training or when the objective is difficult. Meanwhile, Hill-Climb simply fine-tunes the RNN on the top-k molecules per batch, which is sample-efficient but lacks explicit regularization, leading to mode collapse and generation of invalid SMILES.
Previous work by Neil et al. compared RL strategies but did not clearly quantify sample-efficiency differences, and modifications to the REINVENT loss function by Fialkova et al. showed no significant improvement. The best agent reminder (BAR) mechanism offered modest gains but was originally tested on graph-based models.
Core Innovation: Filtering Low-Scoring Molecules from the REINVENT Loss
Augmented Hill-Climb combines the loss formulation of REINVENT with the top-k selection mechanism of Hill-Climb. The agent samples a batch of molecules, ranks them by reward, and computes the REINVENT loss only on the top-k molecules. This removes the counterproductive regularization caused by low-scoring molecules while retaining the prior-based regularization for high-scoring molecules.
The REINVENT loss defines an augmented likelihood:
$$ \log P_{\mathbb{U}}(A) = \log P_{prior}(A) + \sigma R_T $$
where $\sigma$ is a scaling coefficient controlling the reward contribution. The agent loss is the squared difference between the augmented likelihood and the agent’s log-likelihood:
$$ L(\theta) = \left[\log P_{\mathbb{U}}(A) - \log P_{agent}(A)\right]^2 $$
In standard REINVENT, this loss is computed over all molecules in the batch. When $R_T \approx 0$, the augmented likelihood collapses to the prior likelihood, pushing the agent back toward the prior. AHC avoids this by computing the loss only on the top-k molecules ranked by reward, exactly as Hill-Climb selects molecules for fine-tuning.
The key insight is that high-scoring molecules are still regularized by the prior component of the augmented likelihood ($\log P_{prior}(A)$), preventing catastrophic forgetting. Low-scoring molecules, which would otherwise pull the agent back toward the prior, are simply excluded from the loss computation.
Diversity Filters to Prevent Mode Collapse
AHC is more susceptible to mode collapse than REINVENT because it focuses learning on high-scoring molecules. The authors address this with diversity filters (DFs) that penalize the reward of molecules similar to previously generated ones. Through a hyperparameter search over 825 configurations on three GuacaMol tasks, they identify an optimal DF configuration (DF2) with:
- Minimum score threshold of 0.5 (lower than DF1’s 0.8)
- Linear penalization output mode (softer than binary)
- Bin size of 50 (larger than DF1’s 25)
- Scaffold similarity based on ECFP4 fingerprints
The authors find that stricter DFs (lower thresholds, smaller bins) better prevent mode collapse but reduce optimization performance, while more lenient DFs enable better learning of chemotype-reward associations. DF2 represents a compromise.
Experimental Setup: Docking Tasks and Benchmark Comparisons
The evaluation spans five experiments:
Experiment 1: AHC vs. REINVENT on DRD2 docking over 100 RL updates (6,400 samples), varying $\sigma$ from 30 to 240. RNN trained on the MOSESn dataset (MOSES with neutralized charges, 2.45M molecules).
Experiment 2: AHC + DF2 vs. REINVENT on four GPCR targets (DRD2, OPRM1, AGTR1, OX1R) over 500 RL updates. Docking performed with Glide-SP after ligand preparation with LigPrep.
Experiment 3: Diversity filter hyperparameter search (825 configurations) on three GuacaMol tasks (Aripiprazole similarity, C11H24 isomers, Osimertinib MPO) using the GuacaMol training set (1.27M molecules from ChEMBL24).
Experiment 4: Benchmark of AHC against REINFORCE, REINVENT (v1 and v2), BAR, and Hill-Climb (with and without KL regularization) on six tasks of varying difficulty:
| Task | Difficulty | Objective |
|---|---|---|
| Heavy atoms | Easy | Maximize number of heavy atoms |
| Risperidone similarity | Easy | Maximize Tanimoto similarity to Risperidone |
| DRD2 activity | Medium | Maximize QSAR-predicted DRD2 activity |
| DRD2 docking | Medium | Minimize Glide-SP docking score |
| DRD2-DRD3 dual | Hard | Maximize predicted activity against both targets |
| DRD2/DRD3 selective | Hard | Maximize selective DRD2 activity over DRD3 |
Experiment 5: AHC vs. REINVENT on transformer (Tr) and gated transformer (GTr) architectures on the same six benchmark tasks. The GTr implements a GRU-style gate in place of residual connections to stabilize RL training.
RNN and Transformer Architectures
Three RNN configurations were used: (1) embedding 128 + 3 GRU layers of 512 (REINVENT v1), (2) embedding 256 + 3 LSTM layers of 512 (REINVENT 2.0), (3) 3 LSTM layers of 512 with dropout 0.2 (GuacaMol). Transformers used 4 encoder layers with hidden dimension 512, 8 attention heads, and feed-forward dimension 1024.
QSAR models for DRD2 and DRD3 activity were random forest classifiers trained on ExCAPE-DB data with GHOST threshold identification for handling class imbalance.
Key Findings: 45-Fold Sample Efficiency Improvement
Experiment 1: AHC Consistently Outperforms REINVENT
AHC improved optimization ability by 1.39-fold over REINVENT averaged across all $\sigma$ values, with maximum optimization of 205% at $\sigma = 240$ (compared to 128% for REINVENT). AHC required ~80 fewer RL steps to match REINVENT’s mean docking score at 100 steps. With DF1 applied, the improvement was 1.45-fold.
AHC showed greater sensitivity to $\sigma$, giving practitioners more control over the regularization-optimization trade-off. At $\sigma = 60$ (heavily regularized), AHC still improved 1.47-fold over REINVENT while maintaining property space defined by the MOSESn training set. At higher $\sigma$ values, AHC extrapolated further outside the training distribution, which can be favorable (novel chemical space) or unfavorable (scoring function exploitation, e.g., larger molecules getting better docking scores due to the additive nature of scoring functions).
Experiment 2: Improvement Across Four GPCR Targets
Across DRD2, OPRM1, AGTR1, and OX1R, AHC + DF2 required on average 7.4-fold fewer training steps and 45.5-fold fewer samples to reach optimization thresholds. The improvement was largest early in training: 19.8-fold fewer steps to reach 120% optimization, and 71.8-fold fewer samples to first produce a molecule exceeding 160% optimization.
AHC + DF2 surpassed the 80% retrospective precision threshold within 100 RL updates for all targets except the challenging OX1R. DF2 successfully stabilized learning, avoiding the convergence-to-threshold failure mode observed with DF1.
Scaffold analysis showed AHC generates similar chemistry to REINVENT. The top 500 scaffolds produced by REINVENT were also generated by AHC, but typically much sooner.
Experiment 4: Benchmark Against All RL Strategies
AHC outperformed all other RL strategies on all six benchmark tasks except maximizing heavy atoms (an extrapolation task of limited practical relevance). AHC was particularly superior during early-stage optimization and for harder objectives (dual activity, selective activity).
Hill-Climb with a smaller batch size (HC*) showed improved early-stage sample efficiency similar to AHC, but rapidly underwent mode collapse. KL regularization did not rescue mode collapse in any case and sometimes worsened performance. BAR performed poorly in most tasks, possibly because the best-agent memory acts as a second regularizer that inhibits learning.
In terms of wall time for the DRD2 docking task, AHC reached 140% optimization in 16 CPU hours vs. 202 CPU hours for REINVENT 2.0. AHC was the only strategy to reach 200% optimization within the allotted time (216 CPU hours). Parallelized over 10 CPUs, this corresponds to ~21.6 hours, making docking-guided generation feasible on local machines.
Experiment 5: Generalization to Transformers
AHC outperformed REINVENT on both the standard transformer and the gated transformer architectures. The standard transformer was unstable under RL, readily undergoing mode collapse. The gated transformer (with GRU-style gating replacing residual connections) stabilized RL training. AHC’s efficiency gains generalized to both architectures.
Limitations
The authors acknowledge several limitations:
- Chemistry quality evaluation is complicated by the interaction between RL strategy and scoring function suitability. Greater optimization may lead to unreasonable chemistry due to scoring function exploitation rather than the RL strategy itself.
- The diversity filter hyperparameter search was conducted on GuacaMol toy tasks, which may not fully transfer to docking-based objectives.
- The docking scoring function was system-dependent: DRD2 and OPRM1 were optimized effectively, while AGTR1 and OX1R proved more challenging (especially AGTR1, where the docking algorithm targeted the wrong sub-pocket).
- KL regularization proved ineffective for HC and REINFORCE, suggesting it is not a sufficient regularization method in this context.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| RNN pretraining | MOSESn (MOSES neutralized) | 2,454,087 molecules | ZINC15 clean leads with neutralized charges |
| RNN pretraining | GuacaMol train | 1,273,104 molecules | ChEMBL24 with property filters |
| QSAR training | ExCAPE-DB (DRD2) | 4,609 actives / 343,026 inactives | Random forest with GHOST thresholds |
| QSAR training | ExCAPE-DB (DRD3) | 2,758 actives / 402,524 inactives | Unique subsets for dual/selective tasks |
| DF parameter search | GuacaMol benchmark tasks | 3 tasks | 825 configurations tested |
Algorithms
- AHC: REINVENT loss computed on top-k molecules per batch, ranked by reward
- Baselines: REINFORCE, REINVENT (v1, v2), BAR, Hill-Climb, Hill-Climb + KL regularization
- Hyperparameters: Default values from each original publication (listed in Supplementary Table S3)
- Docking: Glide-SP with Schrodinger Protein Preparation Wizard, LigPrep for ligand preparation
Models
- RNNs: 3 configurations (GRU/LSTM, 512 hidden units, trained 5-10 epochs)
- Transformer: 4 encoder layers, 512 hidden dim, 8 heads, 1024 FFN dim
- Gated Transformer: Same architecture with GRU-style gating replacing residual connections
- QSAR: Random forest classifiers (100 estimators, max depth 15, min leaf 2)
Evaluation
| Metric | AHC + DF2 | REINVENT | Notes |
|---|---|---|---|
| Optimization fold-improvement | 1.45x | baseline | DRD2 docking, averaged across sigma values |
| Sample efficiency | 45.5x fewer samples | baseline | Averaged across 4 GPCR targets |
| Step efficiency | 7.4x fewer steps | baseline | Averaged across 4 GPCR targets |
| CPU hours to 140% (DRD2 docking) | 16h | 202h (REINVENT 2.0) | AMD Threadripper 1920 + RTX 2060 Super |
Hardware
- AMD Threadripper 1920 CPU
- Nvidia GeForce RTX 2060 Super GPU
- DRD2 docking benchmark: 216 CPU hours for AHC to reach 200% optimization (~21.6h parallelized over 10 CPUs)
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| SMILES-RNN | Code | MIT | RNN and transformer generative model code |
| MolScore | Code | MIT | Scoring function platform |
| Figshare datasets | Dataset | CC-BY-4.0 | Supporting data (published under same license as paper) |
Paper Information
Citation: Thomas, M., O’Boyle, N. M., Bender, A., & de Graaf, C. (2022). Augmented Hill-Climb increases reinforcement learning efficiency for language-based de novo molecule generation. Journal of Cheminformatics, 14, 68.
@article{thomas2022augmented,
title={Augmented Hill-Climb increases reinforcement learning efficiency for language-based de novo molecule generation},
author={Thomas, Morgan and O'Boyle, Noel M. and Bender, Andreas and de Graaf, Chris},
journal={Journal of Cheminformatics},
volume={14},
number={1},
pages={68},
year={2022},
doi={10.1186/s13321-022-00646-z}
}