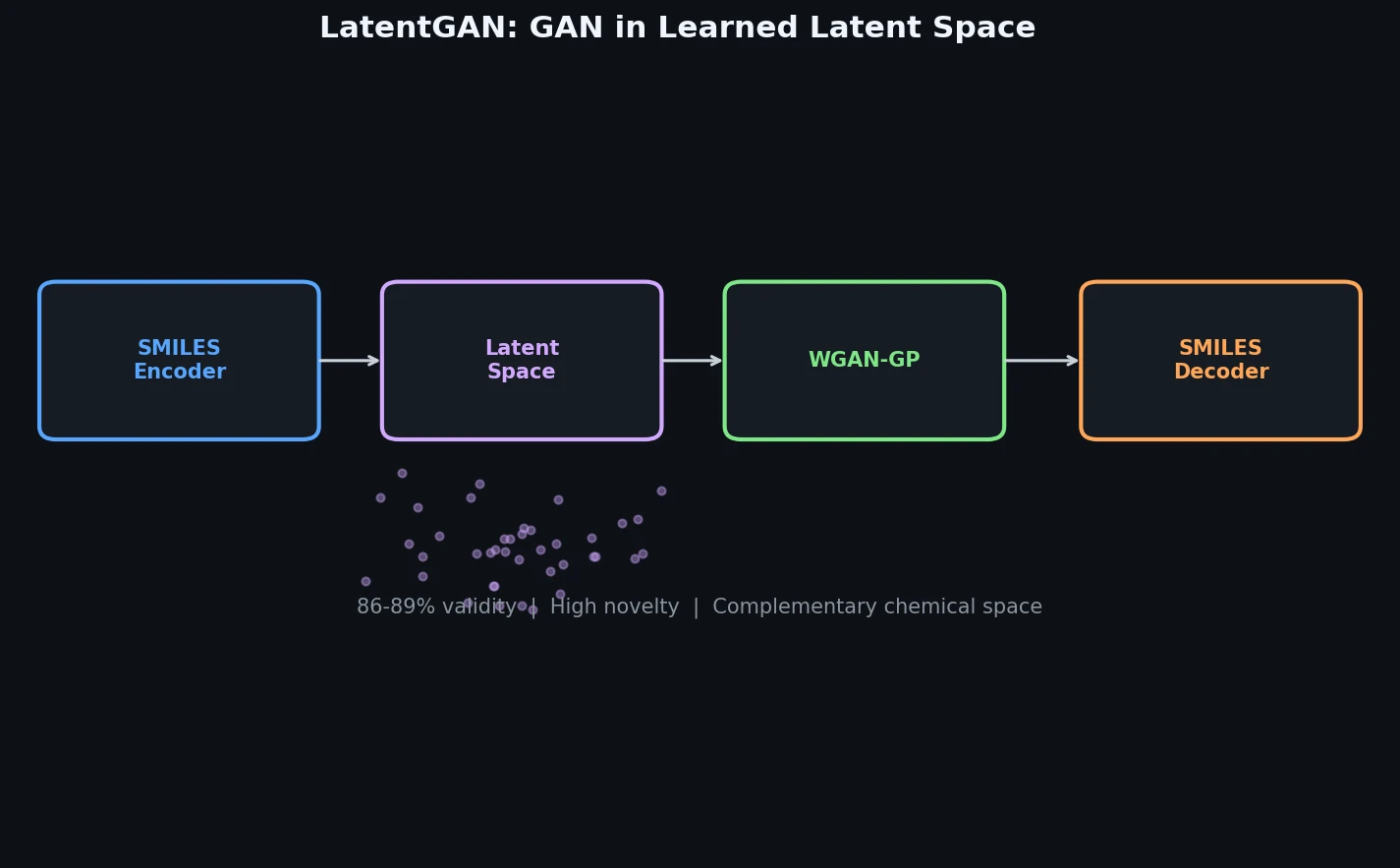
A GAN Operating in Learned Latent Space for Molecular Design
LatentGAN is a Method paper that introduces a two-stage architecture for de novo molecular generation. The first stage trains a heteroencoder to map SMILES strings into a continuous latent vector space. The second stage trains a Wasserstein GAN with gradient penalty (WGAN-GP) to generate new latent vectors that, when decoded, produce valid and novel molecular structures. The key contribution is decoupling the GAN from direct SMILES string generation, allowing the adversarial training to focus on learning the distribution of molecular latent representations rather than character-level sequence generation.
Limitations of Direct SMILES Generation with GANs
Prior GAN-based molecular generation methods such as ORGAN and ORGANIC operated directly on SMILES strings. This created a fundamental challenge: the generator had to simultaneously learn valid SMILES syntax and the distribution of chemically meaningful molecules. ORGAN struggled with optimizing discrete molecular properties like Lipinski’s Rule of Five, while ORGANIC showed limited success beyond the QED drug-likeness score. Other approaches (RANC, ATNC) substituted more advanced recurrent architectures but still operated in the discrete SMILES space.
Meanwhile, variational autoencoders (VAEs) demonstrated that working in continuous latent space could enable molecular generation, but they relied on forcing the latent distribution to match a Gaussian prior through KL divergence. This assumption is not necessarily appropriate for chemical space, which is inherently discontinuous.
RNN-based methods with transfer learning offered an alternative for target-biased generation, but the authors hypothesized that combining GANs with learned latent representations could produce complementary chemical space coverage.
Heteroencoder Plus Wasserstein GAN Architecture
The core innovation of LatentGAN is separating molecular representation learning from adversarial generation through a two-component pipeline.
Heteroencoder
The heteroencoder is an autoencoder trained on pairs of different non-canonical (randomized) SMILES representations of the same molecule. This is distinct from a standard autoencoder because the input and target SMILES are different representations of the same structure.
The encoder uses a two-layer bidirectional LSTM with 512 units per layer (256 forward, 256 backward). The concatenated output feeds into a 512-dimensional feed-forward layer. During training, zero-centered Gaussian noise with $\sigma = 0.1$ is added to the latent vector as regularization. The decoder is a four-layer unidirectional LSTM with a softmax output layer. Batch normalization with momentum 0.9 is applied to all hidden layers except the noise layer.
Training uses teacher forcing with categorical cross-entropy loss for 100 epochs. The learning rate starts at $10^{-3}$ for the first 50 epochs and decays exponentially to $10^{-6}$ by the final epoch. After training, the noise layer is deactivated for deterministic encoding and decoding.
An important design choice is that the heteroencoder makes no assumption about the latent space distribution (unlike VAEs with their KL divergence term). The latent space is shaped purely by reconstruction loss, and the GAN later learns to sample from this unconstrained distribution.
Wasserstein GAN with Gradient Penalty
The GAN uses the WGAN-GP formulation. The critic (discriminator) consists of three feed-forward layers of 256 dimensions each with leaky ReLU activations (no activation on the final layer). The generator has five feed-forward layers of 256 dimensions each with batch normalization and leaky ReLU between layers.
The training ratio is 5:1, with five critic updates for every generator update. The generator takes random vectors sampled from a uniform distribution and learns to produce latent vectors indistinguishable from the real encoded molecular latent vectors.
The WGAN-GP loss for the critic is:
$$L_{\text{critic}} = \mathbb{E}_{\tilde{x} \sim \mathbb{P}_g}[D(\tilde{x})] - \mathbb{E}_{x \sim \mathbb{P}_r}[D(x)] + \lambda \mathbb{E}_{\hat{x} \sim \mathbb{P}_{\hat{x}}}[(|\nabla_{\hat{x}} D(\hat{x})|_2 - 1)^2]$$
where $\lambda$ is the gradient penalty coefficient, $\mathbb{P}_r$ is the real data distribution (encoded latent vectors), $\mathbb{P}_g$ is the generator distribution, and $\mathbb{P}_{\hat{x}}$ samples uniformly along straight lines between pairs of real and generated points.
Generation Pipeline
At inference time, the full pipeline operates as: (1) sample a random vector, (2) pass through the trained generator to produce a latent vector, (3) decode the latent vector into a SMILES string using the pretrained heteroencoder decoder.
Experiments on Drug-Like and Target-Biased Generation
Datasets
The heteroencoder was trained on 1,347,173 SMILES from ChEMBL 25, standardized with MolVS and restricted to molecules with atoms from {H, C, N, O, S, Cl, Br} and at most 50 heavy atoms.
For general drug-like generation, a random subset of 100,000 ChEMBL compounds was used to train the GAN model for 30,000 epochs.
For target-biased generation, three datasets were extracted from ExCAPE-DB for EGFR, HTR1A, and S1PR1 targets. These were clustered into training and test sets to ensure chemical series were not split across sets.
| Target | Training Set | Test Set | SVM ROC-AUC | SVM Kappa |
|---|---|---|---|---|
| EGFR | 2,949 | 2,326 | 0.850 | 0.56 |
| HTR1A | 48,283 | 23,048 | 0.993 | 0.90 |
| S1PR1 | 49,381 | 23,745 | 0.995 | 0.91 |
SVM target prediction models using 2048-bit FCFP6 fingerprints were built with scikit-learn to evaluate generated compounds.
Baselines
RNN-based generative models with transfer learning served as the primary baseline. A prior RNN model was trained on the same ChEMBL set, then fine-tuned on each target dataset. The LatentGAN was also benchmarked on the MOSES platform against VAE, JTN-VAE, and AAE architectures.
Heteroencoder Performance
The heteroencoder achieved 99% valid SMILES on the training set and 98% on the test set. Reconstruction error (decoding to a different molecule) was 18% on training and 20% on test. Notably, decoding to a different valid SMILES of the same molecule is not counted as an error.
Target-Biased Generation Results
From 50,000 sampled SMILES per target model:
| Target | Arch. | Valid (%) | Unique (%) | Novel (%) | Active (%) | Recovered Actives (%) | Recovered Neighbors |
|---|---|---|---|---|---|---|---|
| EGFR | GAN | 86 | 56 | 97 | 71 | 5.26 | 196 |
| EGFR | RNN | 96 | 46 | 95 | 65 | 7.74 | 238 |
| HTR1A | GAN | 86 | 66 | 95 | 71 | 5.05 | 284 |
| HTR1A | RNN | 96 | 50 | 90 | 81 | 7.28 | 384 |
| S1PR1 | GAN | 89 | 31 | 98 | 44 | 0.93 | 24 |
| S1PR1 | RNN | 97 | 35 | 97 | 65 | 3.72 | 43 |
MOSES Benchmark
On the MOSES benchmark (trained on a ZINC subset of 1,584,663 compounds, sampled 30,000 SMILES), LatentGAN showed comparable or better results than JTN-VAE and AAE on Frechet ChemNet Distance (FCD), Fragment similarity, and Scaffold similarity, while producing slightly worse nearest-neighbor cosine similarity (SNN). The standard VAE showed signs of mode collapse with high test metric overlap and low novelty.
Complementary Generation and Drug-Likeness Preservation
Key Findings
Validity and novelty: LatentGAN achieved 86-89% validity on target-biased tasks (lower than RNN’s 96-97%) but produced higher uniqueness on two of three targets and comparable or higher novelty (95-98%).
Complementary chemical space: The overlap between LatentGAN-generated and RNN-generated active compounds was very small at both compound and scaffold levels. A probabilistic analysis showed that the RNN model would be very unlikely to eventually cover the LatentGAN output space. This suggests the two architectures can work complementarily in de novo design campaigns.
Drug-likeness: QED score distributions of LatentGAN-generated compounds closely matched training set distributions across all three targets, with training compounds showing only slightly higher drug-likeness. SA score distributions were similarly well-preserved.
Chemical space coverage: PCA analysis using MQN fingerprints confirmed that generated compounds occupy most of the chemical space of the training sets. Some regions of the PCA plots contained compounds predicted as inactive, which corresponded to non-drug-like outliers in the training data.
Novel scaffolds: About 14% of scaffolds in the sampled sets had similarity below 0.4 to the training set across all three targets, indicating LatentGAN can generate genuinely novel chemical scaffolds. Around 5% of generated compounds were identical to training set compounds, while 21-25% had Tanimoto similarity below 0.4.
Limitations
The paper acknowledges several limitations. The 18-20% heteroencoder reconstruction error means a non-trivial fraction of encoded molecules decode to different structures. Validity rates (86-89%) are lower than RNN baselines (96-97%). The S1PR1 target showed notably lower uniqueness (31%) and predicted activity (44%) compared to the other targets, possibly due to the smaller effective training set of active compounds. The paper does not report specific hardware requirements or training times. No wet-lab experimental validation of generated compounds was performed.
Future Directions
The authors envision LatentGAN as a complementary tool to existing RNN-based generative models, with the two architectures covering different regions of chemical space. The approach of operating in learned latent space rather than directly on SMILES strings offers a general framework that could be extended to other molecular representations or generation objectives.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Heteroencoder training | ChEMBL 25 (subset) | 1,347,173 SMILES | Standardized with MolVS; atoms restricted to H, C, N, O, S, Cl, Br; max 50 heavy atoms |
| General GAN training | ChEMBL 25 (random subset) | 100,000 | Subset of heteroencoder training set |
| Target-biased training | ExCAPE-DB (EGFR) | 2,949 actives | Clustered train/test split |
| Target-biased training | ExCAPE-DB (HTR1A) | 48,283 actives | Clustered train/test split |
| Target-biased training | ExCAPE-DB (S1PR1) | 49,381 actives | Clustered train/test split |
| Benchmarking | ZINC (MOSES subset) | 1,584,663 | Canonical SMILES |
Algorithms
- Heteroencoder: Bidirectional LSTM encoder (2 layers, 512 units) + unidirectional LSTM decoder (4 layers), trained with teacher forcing and categorical cross-entropy for 100 epochs
- GAN: WGAN-GP with 5:1 critic-to-generator training ratio. General model trained 30,000 epochs; target models trained 10,000 epochs
- Evaluation: SVM classifiers with FCFP6 fingerprints (2048 bits) for activity prediction; MQN fingerprints for PCA-based chemical space analysis; Murcko scaffolds for scaffold-level analysis
Models
- Heteroencoder: 512-dim latent space, bidirectional LSTM encoder, unidirectional LSTM decoder
- Generator: 5 feed-forward layers of 256 dims with batch norm and leaky ReLU
- Critic: 3 feed-forward layers of 256 dims with leaky ReLU
Evaluation
| Metric | LatentGAN (EGFR) | RNN Baseline (EGFR) | Notes |
|---|---|---|---|
| Validity | 86% | 96% | Percent valid SMILES |
| Uniqueness | 56% | 46% | Percent unique among valid |
| Novelty | 97% | 95% | Not in training set |
| Predicted active | 71% | 65% | By SVM model |
Hardware
Not specified in the paper.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| LatentGAN source code | Code | Not specified | Includes trained heteroencoder model and training sets |
Paper Information
Citation: Prykhodko, O., Johansson, S.V., Kotsias, P.-C., Arús-Pous, J., Bjerrum, E.J., Engkvist, O., & Chen, H. (2019). A de novo molecular generation method using latent vector based generative adversarial network. Journal of Cheminformatics, 11(1), 74. https://doi.org/10.1186/s13321-019-0397-9
@article{prykhodko2019latentgan,
title={A de novo molecular generation method using latent vector based generative adversarial network},
author={Prykhodko, Oleksii and Johansson, Simon Viet and Kotsias, Panagiotis-Christos and Ar{\'u}s-Pous, Josep and Bjerrum, Esben Jannik and Engkvist, Ola and Chen, Hongming},
journal={Journal of Cheminformatics},
volume={11},
number={1},
pages={74},
year={2019},
publisher={Springer},
doi={10.1186/s13321-019-0397-9}
}