A Grammar-Constrained VAE for Discrete Data Generation
This is a Method paper that introduces the Grammar Variational Autoencoder (GVAE), a variant of the variational autoencoder that operates directly on parse trees from context-free grammars (CFGs) rather than on raw character sequences. The primary contribution is a decoding mechanism that uses a stack and grammar-derived masks to restrict the output at every timestep to only syntactically valid production rules. This guarantees that every decoded output is a valid string under the grammar, addressing a fundamental limitation of character-level VAEs when applied to structured discrete data such as SMILES molecular strings and arithmetic expressions.
Why Character-Level VAEs Fail on Structured Discrete Data
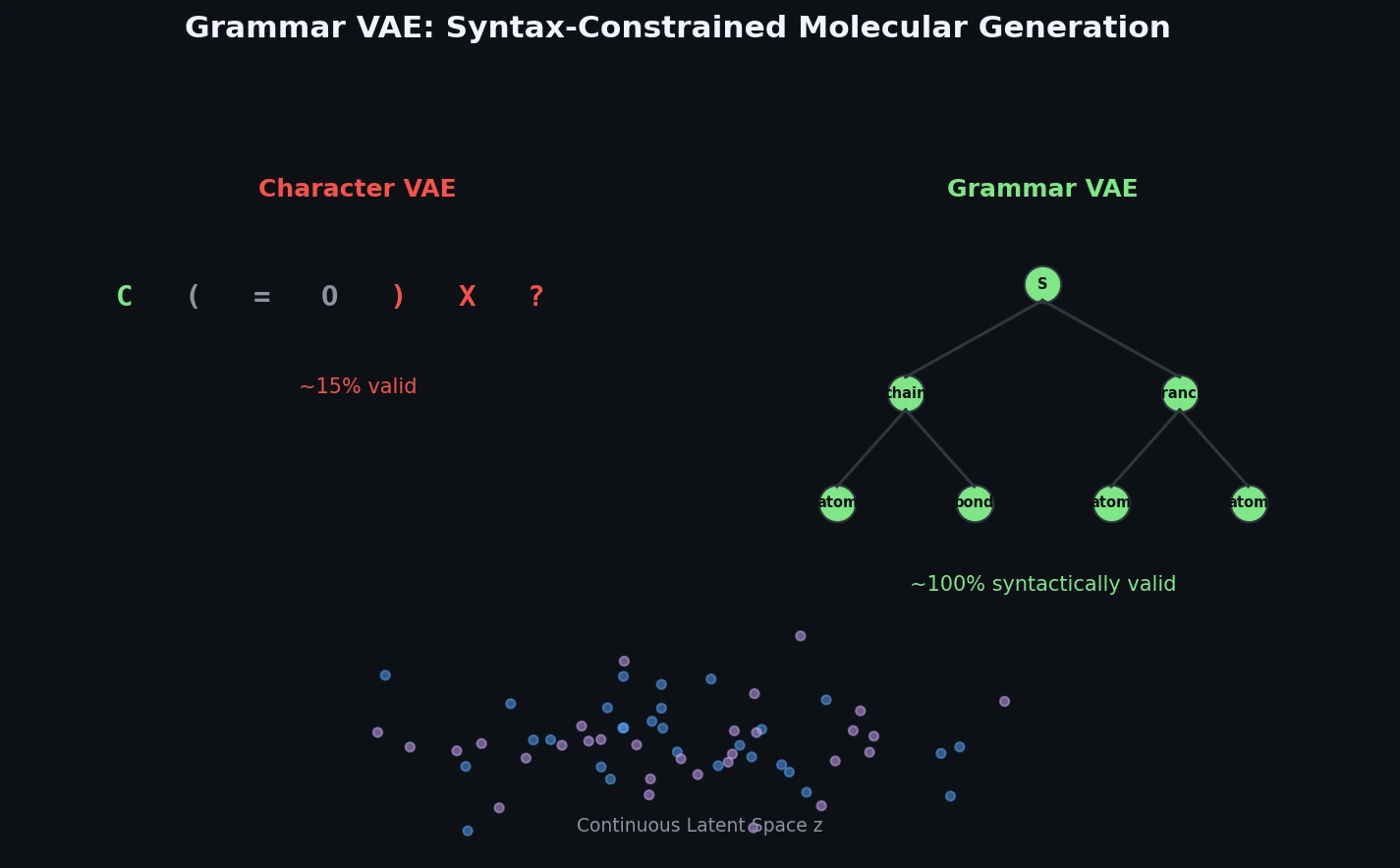
Generative models for continuous data (images, audio) had achieved impressive results by 2017, but generating structured discrete data remained difficult. The key challenge is that string representations of molecules and mathematical expressions are brittle: small perturbations to a character sequence often produce invalid outputs. Gomez-Bombarelli et al. (2016) demonstrated a character-level VAE (CVAE) for SMILES strings that could encode molecules into a continuous latent space and decode them back, enabling latent-space optimization for molecular design. However, the CVAE frequently decoded latent points into strings that were not valid SMILES, particularly when exploring regions of latent space far from training data.
The fundamental issue is that character-level decoders must implicitly learn the syntactic rules of the target language from data alone. For SMILES, this includes matching parentheses, valid atom types, proper bonding, and ring closure notation. The GVAE addresses this by giving the decoder explicit knowledge of the grammar, so it can focus entirely on learning the semantic structure of the data.
Core Innovation: Stack-Based Grammar Masking in the Decoder
The GVAE encodes and decodes sequences of production rules from a context-free grammar rather than sequences of characters.
Encoding. Given an input string (e.g., a SMILES molecule), the encoder first parses it into a parse tree using the CFG, then performs a left-to-right pre-order traversal of the tree to extract an ordered sequence of production rules. Each rule is represented as a one-hot vector of dimension $K$ (total number of production rules in the grammar). The resulting $T(\mathbf{X}) \times K$ matrix is processed by a convolutional neural network to produce the mean and variance of a Gaussian posterior $q_{\phi}(\mathbf{z} \mid \mathbf{X})$.
Decoding with grammar masks. The decoder maps a latent vector $\mathbf{z}$ through an RNN to produce a matrix of logits $\mathbf{F} \in \mathbb{R}^{T_{max} \times K}$. The key innovation is a last-in first-out (LIFO) stack that tracks the current parsing state. At each timestep $t$, the decoder:
- Pops the top non-terminal $\alpha$ from the stack
- Applies a fixed binary mask $\mathbf{m}_{\alpha} \in {0, 1}^K$ that zeros out all production rules whose left-hand side is not $\alpha$
- Samples a production rule from the masked softmax distribution:
$$ p(\mathbf{x}_{t} = k \mid \alpha, \mathbf{z}) = \frac{m_{\alpha,k} \exp(f_{tk})}{\sum_{j=1}^{K} m_{\alpha,j} \exp(f_{tj})} $$
- Pushes the right-hand-side non-terminals of the selected rule onto the stack (right-to-left, so the leftmost is on top)
This process continues until the stack is empty or $T_{max}$ timesteps are reached. Because the mask restricts selection to only those rules applicable to the current non-terminal, every generated sequence of production rules is guaranteed to be a valid derivation under the grammar.
Training. The model is trained by maximizing the ELBO:
$$ \mathcal{L}(\phi, \theta; \mathbf{X}) = \mathbb{E}_{q(\mathbf{z} \mid \mathbf{X})} \left[ \log p_{\theta}(\mathbf{X}, \mathbf{z}) - \log q_{\phi}(\mathbf{z} \mid \mathbf{X}) \right] $$
where the likelihood factorizes as:
$$ p(\mathbf{X} \mid \mathbf{z}) = \prod_{t=1}^{T(\mathbf{X})} p(\mathbf{x}_{t} \mid \mathbf{z}) $$
During training, the masks at each timestep are determined by the ground-truth production rule sequence, so no stack simulation is needed. The stack-based decoding is only required at generation time.
Syntactic vs. semantic validity. The grammar guarantees syntactic validity but not semantic validity. The GVAE can still produce chemically implausible molecules (e.g., an oxygen atom with three bonds) because such constraints are not context-free. SMILES ring-bond digit matching is also not context-free, so the grammar cannot enforce it. Additionally, sequences that have not emptied the stack by $T_{max}$ are marked invalid.
Experiments on Symbolic Regression and Molecular Optimization
The authors evaluate the GVAE on two domains: arithmetic expressions and molecules. Both use Bayesian optimization (BO) over the learned latent space.
Setup. After training each VAE, the authors encode training data into latent vectors and train a sparse Gaussian process (SGP) with 500 inducing points to predict properties from latent representations. They then run batch BO with expected improvement, selecting 50 candidates per iteration.
Arithmetic Expressions
- Data: 100,000 randomly generated univariate expressions from a simple grammar (3 binary operators, 2 unary operators, 3 constants), each with at most 15 production rules
- Target: Find an expression minimizing $\log(1 + \text{MSE})$ against the true function $1/3 + x + \sin(x \cdot x)$
- BO iterations: 5, averaged over 10 repetitions
| Method | Fraction Valid | Average Score |
|---|---|---|
| GVAE | 0.99 +/- 0.01 | 3.47 +/- 0.24 |
| CVAE | 0.86 +/- 0.06 | 4.75 +/- 0.25 |
The GVAE’s best expression ($x/1 + \sin(3) + \sin(x \cdot x)$, score 0.04) nearly exactly recovers the true function, while the CVAE’s best ($x \cdot 1 + \sin(3) + \sin(3/1)$, score 0.39) misses the sinusoidal component.
Molecular Optimization
- Data: 250,000 SMILES strings from the ZINC database
- Target: Maximize penalized logP (water-octanol partition coefficient penalized for ring size and synthetic accessibility)
- BO iterations: 10, averaged over 5 trials
| Method | Fraction Valid | Average Score |
|---|---|---|
| GVAE | 0.31 +/- 0.07 | -9.57 +/- 1.77 |
| CVAE | 0.17 +/- 0.05 | -54.66 +/- 2.66 |
The GVAE produces roughly twice as many valid molecules as the CVAE and finds molecules with substantially better penalized logP scores (best: 2.94 vs. 1.98).
Latent Space Quality
Interpolation experiments show that the GVAE produces valid outputs at every intermediate point when linearly interpolating between two encoded expressions, while the CVAE passes through invalid strings. Grid searches around encoded molecules in the GVAE latent space show smooth transitions where neighboring points differ by single atoms.
Predictive Performance
Sparse GP models trained on GVAE latent features achieve better test RMSE and log-likelihood than those trained on CVAE features for both expressions and molecules:
| Metric | GVAE (Expressions) | CVAE (Expressions) | GVAE (Molecules) | CVAE (Molecules) |
|---|---|---|---|---|
| Test LL | -1.320 +/- 0.001 | -1.397 +/- 0.003 | -1.739 +/- 0.004 | -1.812 +/- 0.004 |
| Test RMSE | 0.884 +/- 0.002 | 0.975 +/- 0.004 | 1.404 +/- 0.006 | 1.504 +/- 0.006 |
Reconstruction and Prior Sampling
On held-out molecules, the GVAE achieves 53.7% reconstruction accuracy vs. 44.6% for the CVAE. When sampling from the prior $p(\mathbf{z}) = \mathcal{N}(0, \mathbf{I})$, 7.2% of GVAE samples are valid molecules vs. 0.7% for the CVAE.
Key Findings, Limitations, and Impact
Key findings. Incorporating grammar structure into the VAE decoder consistently improves validity rates, latent space smoothness, downstream predictive performance, and Bayesian optimization outcomes across both domains. The approach is general: any domain with a context-free grammar can benefit.
Limitations acknowledged by the authors.
- The GVAE guarantees syntactic but not semantic validity. For molecules, invalid ring-bond patterns and chemically implausible structures can still be generated.
- The molecular validity rate during BO (31%) is substantially higher than the CVAE (17%) but still means most decoded molecules are invalid, largely due to non-context-free constraints in SMILES.
- The approach requires a context-free grammar for the target domain, which limits applicability to well-defined formal languages.
- Sequences that do not complete parsing within $T_{max}$ timesteps are discarded as invalid.
Impact. The GVAE was an influential early contribution to constrained molecular generation. It directly inspired the Syntax-Directed VAE (SD-VAE) by Dai et al. (2018), which uses attribute grammars for tighter semantic constraints, and contributed to the broader movement toward structured molecular generation methods including graph-based approaches. The paper demonstrated that encoding domain knowledge into the decoder architecture is more effective than relying on the model to learn structural constraints from data alone.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Training (expressions) | Generated arithmetic expressions | 100,000 | Up to 15 production rules each |
| Training (molecules) | ZINC database subset | 250,000 SMILES | Same subset as Gomez-Bombarelli et al. (2016) |
Algorithms
- Encoder: 1D convolutional neural network over one-hot rule sequences
- Decoder: RNN with stack-based grammar masking
- Latent space: 56 dimensions (molecules), isotropic Gaussian prior
- Property predictor: Sparse Gaussian process with 500 inducing points
- Optimization: Batch Bayesian optimization with expected improvement, 50 candidates per iteration, Kriging Believer for batch selection
Models
Architecture details follow Gomez-Bombarelli et al. (2016) with modifications for grammar-based encoding/decoding. Specific layer sizes and hyperparameters are described in the supplementary material.
Evaluation
| Metric | GVAE | CVAE | Notes |
|---|---|---|---|
| Fraction valid (expressions) | 0.99 | 0.86 | During BO |
| Fraction valid (molecules) | 0.31 | 0.17 | During BO |
| Best penalized logP | 2.94 | 1.98 | Best molecule found |
| Reconstruction accuracy | 53.7% | 44.6% | On held-out molecules |
| Prior validity | 7.2% | 0.7% | Sampling from N(0,I) |
Hardware
Not specified in the paper.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| grammarVAE | Code | Not specified | Official implementation |
Paper Information
Citation: Kusner, M. J., Paige, B., & Hernández-Lobato, J. M. (2017). Grammar Variational Autoencoder. Proceedings of the 34th International Conference on Machine Learning (ICML), 1945-1954.
@inproceedings{kusner2017grammar,
title={Grammar Variational Autoencoder},
author={Kusner, Matt J. and Paige, Brooks and Hern{\'a}ndez-Lobato, Jos{\'e} Miguel},
booktitle={Proceedings of the 34th International Conference on Machine Learning},
pages={1945--1954},
year={2017},
publisher={PMLR}
}