A Foundational Method for Continuous Molecular Representation
This is a Method paper that introduces a variational autoencoder (VAE) framework for mapping discrete molecular representations (SMILES strings) into a continuous latent space. The primary contribution is demonstrating that this continuous representation enables three key capabilities: (1) automatic generation of novel molecules by decoding random or perturbed latent vectors, (2) smooth interpolation between molecules in latent space, and (3) gradient-based optimization of molecular properties using a jointly trained property predictor. This work is widely regarded as one of the earliest and most influential applications of deep generative models to molecular design.
The Challenge of Searching Discrete Chemical Space
Molecular design is fundamentally an optimization problem: identify molecules that maximize some set of desirable properties. The search space is enormous (estimated $10^{23}$ to $10^{60}$ drug-like molecules) and discrete, making systematic exploration difficult. Prior approaches fell into two categories, each with significant limitations:
- Virtual screening over fixed libraries: effective but monolithic, costly to enumerate, and requiring hand-crafted rules to avoid impractical chemistries.
- Discrete local search (e.g., genetic algorithms): requires manual specification of mutation and crossover heuristics, and cannot leverage gradient information to guide the search.
The core insight is that mapping molecules into a continuous vector space sidesteps these problems entirely. In a continuous space, new compounds can be generated by vector perturbation (no hand-crafted mutation rules), optimization can follow property gradients (enabling larger and more directed jumps), and large unlabeled chemical databases can be leveraged through unsupervised representation learning.
A VAE Architecture for SMILES Strings with Joint Property Prediction
The architecture consists of three coupled neural networks trained jointly:
Encoder: Converts SMILES character strings into fixed-dimensional continuous vectors (the latent representation). Uses three 1D convolutional layers followed by a fully connected layer. For ZINC molecules, the latent space has 196 dimensions; for QM9, 156 dimensions.
Decoder: Converts latent vectors back into SMILES strings character by character using three layers of gated recurrent units (GRUs). The output is stochastic, as each character is sampled from a probability distribution over the SMILES alphabet.
Property Predictor: A multilayer perceptron that predicts molecular properties directly from the latent representation. Joint training with the autoencoder reconstruction loss organizes the latent space so that molecules with similar properties cluster together.
The VAE Objective
The model uses the variational autoencoder framework of Kingma and Welling. The training objective combines three terms:
$$\mathcal{L} = \mathcal{L}_{recon} + \beta \cdot D_{KL}(q(z|x) | p(z)) + \lambda \cdot \mathcal{L}_{prop}$$
where $\mathcal{L}_{recon}$ is the reconstruction loss (cross-entropy over SMILES characters), $D_{KL}$ is the KL divergence regularizer that encourages the latent distribution $q(z|x)$ to match a standard Gaussian prior $p(z)$, and $\mathcal{L}_{prop}$ is the property prediction regression loss. Both the variational loss and the property prediction loss are annealed in using a sigmoid schedule after 29 epochs over a total of 120 epochs of training.
The KL regularization is critical: it forces the decoder to handle a wider variety of latent points, preventing “dead areas” in latent space that would decode to invalid molecules.
Gradient-Based Optimization
After training, a Gaussian process (GP) surrogate model is fit on top of the latent representations to predict the target property. Optimization proceeds by:
- Encoding a seed molecule into the latent space
- Using the GP model to define a smooth property surface over the latent space
- Optimizing the latent vector $z$ to maximize the predicted property via gradient ascent
- Decoding the optimized $z$ back into a SMILES string
The objective used for demonstration is $5 \times \text{QED} - \text{SAS}$, balancing drug-likeness (QED) against synthetic accessibility (SAS).
Experiments on ZINC and QM9 Datasets
Two autoencoder systems were trained:
- ZINC: 250,000 drug-like molecules from the ZINC database, with a 196-dimensional latent space. Properties predicted: logP, QED, SAS.
- QM9: 108,000 molecules with fewer than 9 heavy atoms, with a 156-dimensional latent space. Properties predicted: HOMO energy, LUMO energy, electronic spatial extent ($\langle R^2 \rangle$).
Latent Space Quality
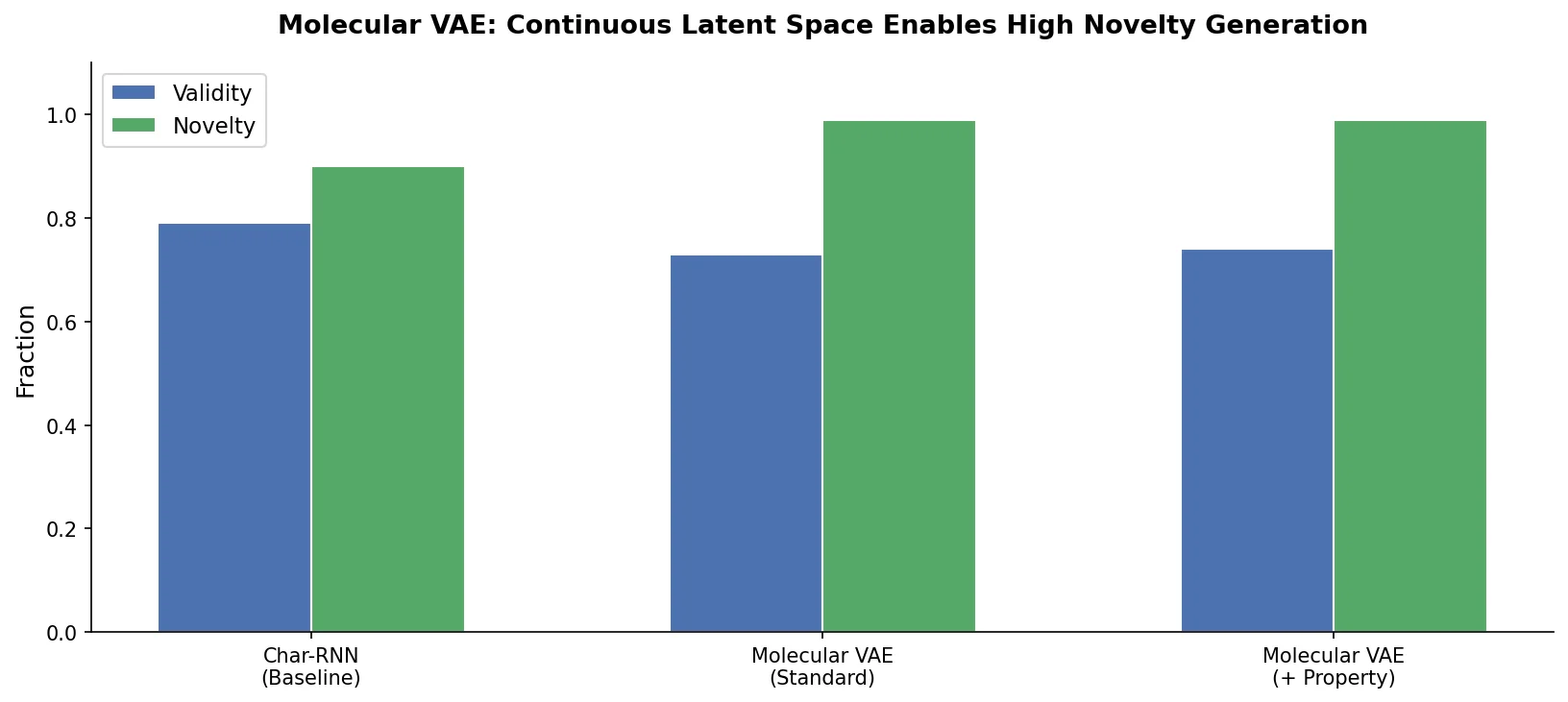
The encoded latent dimensions follow approximately normal distributions as enforced by the variational regularizer. Decoding is stochastic: sampling the same latent point multiple times yields different SMILES strings, with the most frequent decoding tending to be closest to the original point in latent space. Decoding validity rates are 73-79% for points near known molecules but only 4% for randomly selected latent points.
Spherical interpolation (slerp) between molecules in latent space produces smooth structural transitions, accounting for the geometry of high-dimensional Gaussian distributions where linear interpolation would pass through low-probability regions.
Molecular Generation Comparison
| Source | Dataset | Samples | logP | SAS | QED | % in ZINC | % in eMolecules |
|---|---|---|---|---|---|---|---|
| Data | ZINC | 249k | 2.46 (1.43) | 3.05 (0.83) | 0.73 (0.14) | 100 | 12.9 |
| GA | ZINC | 5303 | 2.84 (1.86) | 3.80 (1.01) | 0.57 (0.20) | 6.5 | 4.8 |
| VAE | ZINC | 8728 | 2.67 (1.46) | 3.18 (0.86) | 0.70 (0.14) | 5.8 | 7.0 |
| Data | QM9 | 134k | 0.30 (1.00) | 4.25 (0.94) | 0.48 (0.07) | 0.0 | 8.6 |
| GA | QM9 | 5470 | 0.96 (1.53) | 4.47 (1.01) | 0.53 (0.13) | 0.018 | 3.8 |
| VAE | QM9 | 2839 | 0.30 (0.97) | 4.34 (0.98) | 0.47 (0.08) | 0.0 | 8.9 |
The VAE generates molecules whose property distributions closely match the training data, outperforming a genetic algorithm baseline that biases toward higher chemical complexity and decreased drug-likeness. Only 5.8% of VAE-generated ZINC molecules were found in the original ZINC database, indicating genuine novelty.
Property Prediction
| Dataset/Property | Mean Baseline | ECFP | Graph Conv. | 1-hot SMILES | Encoder Only | VAE |
|---|---|---|---|---|---|---|
| ZINC/logP | 1.14 | 0.38 | 0.05 | 0.16 | 0.13 | 0.15 |
| ZINC/QED | 0.112 | 0.045 | 0.017 | 0.041 | 0.037 | 0.054 |
| QM9/HOMO (eV) | 0.44 | 0.20 | 0.12 | 0.12 | 0.13 | 0.16 |
| QM9/LUMO (eV) | 1.05 | 0.20 | 0.15 | 0.11 | 0.14 | 0.16 |
| QM9/Gap (eV) | 1.07 | 0.30 | 0.18 | 0.16 | 0.18 | 0.21 |
The VAE latent representation achieves property prediction accuracy comparable to graph convolutions for some properties, though graph convolutions generally perform best. The primary purpose of joint training is not to maximize prediction accuracy but to organize the latent space for optimization.
Optimization Results
Bayesian optimization with a GP model on the jointly trained latent space consistently produces molecules with higher percentile scores on the $5 \times \text{QED} - \text{SAS}$ objective compared to both random Gaussian search and genetic algorithm baselines. Starting from molecules in the bottom 10th percentile of the ZINC dataset, the optimizer reliably discovers molecules in regions of high objective value. Training the GP with 1000 molecules (vs. 2000) produces a wider diversity of solutions by optimizing to multiple local optima rather than a single global optimum.
Key Findings, Limitations, and Legacy
Key Findings
- A continuous latent representation of molecules enables gradient-based search through chemical space, a qualitatively different approach from discrete enumeration or genetic algorithms.
- Joint training with property prediction organizes the latent space by property values, creating smooth gradients that optimization can follow.
- The VAE generates novel molecules with realistic property distributions, and the latent space encodes an estimated 7.5 million molecules despite training on only 250,000.
Acknowledged Limitations
- The SMILES-based decoder sometimes produces formally valid but chemically undesirable molecules (acid chlorides, anhydrides, cyclopentadienes, aziridines, etc.) because the grammar of valid SMILES does not capture all synthetic or stability constraints.
- Character-level SMILES generation is fragile: the decoder must implicitly learn which strings are valid SMILES, making the learning problem harder than necessary.
- Decoding validity drops to only 4% for random latent points far from training data, limiting the ability to explore truly novel regions of chemical space.
Directions Identified
The authors point to several extensions that were already underway at the time of publication:
- Grammar VAE: Using an explicitly defined SMILES grammar instead of forcing the model to learn one (Kusner et al., 2017).
- Graph-based decoders: Directly outputting molecular graphs to avoid the SMILES validity problem.
- Adversarial training: Using GANs for molecular generation (ORGAN, ORGANIC).
- LSTM/RNN generators: Applying recurrent networks directly to SMILES for generation and reaction prediction.
This paper has been cited over 2,900 times and launched a large body of follow-up work in VAE-based, GAN-based, and reinforcement learning-based molecular generation.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Training | ZINC (drug-like subset) | 250,000 molecules | Randomly sampled from ZINC database |
| Training | QM9 | 108,000 molecules | Molecules with fewer than 9 heavy atoms |
| Evaluation | ZINC held-out set | 5,000 molecules | For latent space analysis |
Algorithms
- Encoder: 3 x 1D convolutional layers (ZINC: filters 9,9,10 with kernels 9,9,11; QM9: filters 2,2,1 with kernels 5,5,4), followed by a fully connected layer
- Decoder: 3 x GRU layers (ZINC: hidden dim 488; QM9: hidden dim 500), trained with teacher forcing
- Property Predictor: 2 fully connected layers of 1000 neurons (dropout 0.20) for prediction; smaller 3-layer MLP of 67 neurons (dropout 0.15) for latent space shaping
- Variational loss annealing: Sigmoid schedule after 29 epochs, total 120 epochs
- SMILES validation: Post-hoc filtering with RDKit; invalid outputs discarded
- Optimization: Gaussian process surrogate model trained on 2000 maximally diverse molecules from latent space
Models
Built with Keras and TensorFlow. Latent dimensions: 196 (ZINC), 156 (QM9). SMILES alphabet: 35 characters (ZINC), 22 characters (QM9). Maximum string length: 120 (ZINC), 34 (QM9). Only canonicalized SMILES used for training.
Evaluation
| Metric | Description |
|---|---|
| logP | Water-octanol partition coefficient |
| QED | Quantitative Estimation of Drug-likeness (0-1) |
| SAS | Synthetic Accessibility Score |
| HOMO/LUMO (eV) | Frontier orbital energies (QM9) |
| Decoding validity | Fraction of latent points producing valid SMILES |
| Novelty | Fraction of generated molecules not in training set |
Hardware
Training was performed on the Harvard FAS Odyssey Cluster. Specific GPU types and training times are not reported. The Gaussian process optimization requires only minutes to train on a few thousand molecules.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| chemical_vae | Code | Apache-2.0 | Official implementation with training scripts and pre-trained models |
Paper Information
Citation: Gómez-Bombarelli, R., Wei, J. N., Duvenaud, D., Hernández-Lobato, J. M., Sánchez-Lengeling, B., Sheberla, D., Aguilera-Iparraguirre, J., Hirzel, T. D., Adams, R. P., & Aspuru-Guzik, A. (2018). Automatic Chemical Design Using a Data-Driven Continuous Representation of Molecules. ACS Central Science, 4(2), 268-276. https://doi.org/10.1021/acscentsci.7b00572
@article{gomez2018automatic,
title={Automatic Chemical Design Using a Data-Driven Continuous Representation of Molecules},
author={G{\'o}mez-Bombarelli, Rafael and Wei, Jennifer N. and Duvenaud, David and Hern{\'a}ndez-Lobato, Jos{\'e} Miguel and S{\'a}nchez-Lengeling, Benjam{\'i}n and Sheberla, Dennis and Aguilera-Iparraguirre, Jorge and Hirzel, Timothy D. and Adams, Ryan P. and Aspuru-Guzik, Al{\'a}n},
journal={ACS Central Science},
volume={4},
number={2},
pages={268--276},
year={2018},
publisher={American Chemical Society},
doi={10.1021/acscentsci.7b00572}
}