An Empirical Re-evaluation of Generative Model Benchmarks
This is an Empirical paper. The primary contribution is a critical reassessment of the Practical Molecular Optimization (PMO) benchmark for de novo molecule generation. Rather than proposing a new generative model, the authors modify existing benchmark metrics to account for chemical desirability (molecular weight, LogP, topological novelty) and molecular diversity. They then re-evaluate all 25 generative models from the original PMO benchmark plus the recently proposed Augmented Hill-Climb (AHC) method.
Sample Efficiency and Chemical Quality in Drug Design
Deep generative models for de novo molecule generation often require large numbers of oracle evaluations (up to $10^5$ samples) to optimize toward a target objective. This is a practical limitation when using computationally expensive scoring functions like molecular docking. The PMO benchmark by Gao et al. addressed this by reformulating performance as maximizing an objective within a fixed budget of 10,000 oracle calls, finding REINVENT to be the most sample-efficient model across 23 tasks.
However, the authors identify a key limitation: the PMO benchmark measures only sample efficiency without considering the chemical quality of proposed molecules. Investigating the top-performing REINVENT model on the JNK3 task, they find that 4 of 5 replicate runs produce molecules with molecular weight and LogP distributions far outside the training data (ZINC250k). The resulting molecules contain large structures with repeating substructures that are undesirable from a medicinal chemistry perspective. This disconnect between benchmark performance and practical utility motivates the modified evaluation metrics.
Modified Metrics: Property Filters and Diversity Requirements
The core innovation is the introduction of three modified AUC Top-10 metrics that extend the original PMO benchmark evaluation:
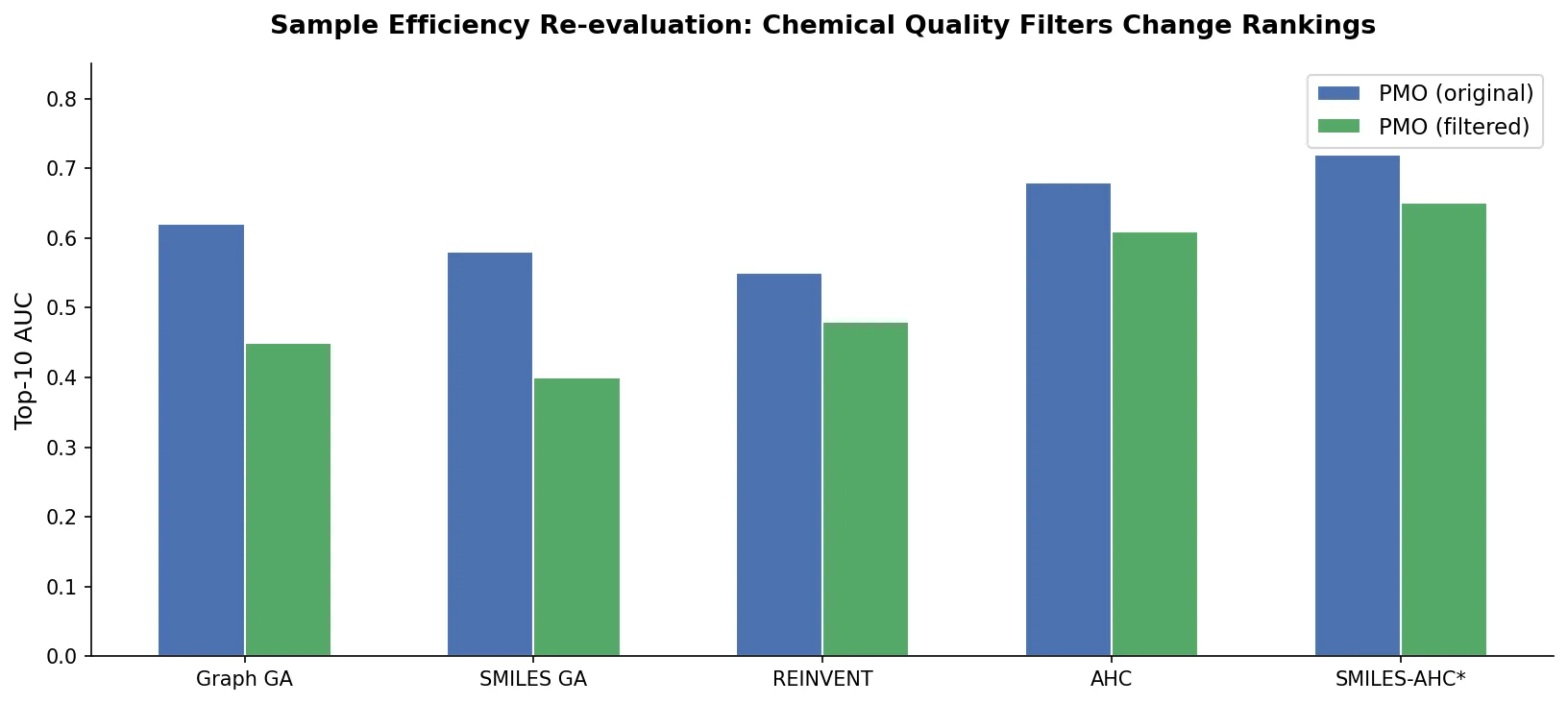
AUC Top-10 (Filtered): Molecules are excluded if their molecular weight or LogP falls beyond 4 standard deviations from the mean of the ZINC250k pre-training dataset ($\mu \pm 4\sigma$, covering approximately 99.99% of a normal distribution). Molecules with more than 10% de novo (unobserved in ZINC250k) ECFP4 fingerprint bits are also filtered out. This ensures the generative model does not drift beyond its applicability domain.
AUC Top-10 (Diverse): The top 10 molecules are selected iteratively, where a molecule is only added if its Tanimoto similarity (by ECFP4 fingerprints) to any previously selected compound does not exceed 0.35. This threshold corresponds to an approximately 80% probability that more-similar molecules belong to the same bioactivity class, enforcing that distinct candidates possess different profiles.
AUC Top-10 (Combined): Applies both property filters and diversity filters simultaneously, providing the most stringent evaluation of practical performance.
Benchmark Setup and Generative Models Evaluated
Implementation Details
The authors re-implement the PMO benchmark using the original code and data (MIT license) with no changes beyond adding AHC and the new metrics. For Augmented Hill-Climb, the architecture follows REINVENT: an embedding layer of size 128 and 3 layers of Gated Recurrent Units (GRU) with size 512. The prior is trained on ZINC250k using SMILES notation with batch size 128 for 5 epochs.
Two AHC variants are benchmarked:
- SMILES-AHC: Hyperparameters optimized via the standard PMO procedure, yielding batch size 256, $\sigma = 120$, $K = 0.25$
- SMILES-AHC*: Uses $\sigma = 60$, chosen based on prior knowledge that lower $\sigma$ values maintain better regularization and chemical quality
Both omit diversity filters and non-unique penalization for standardized comparison, despite these being shown to improve performance in prior work.
Models Compared
The benchmark includes 25 generative models from the original PMO paper spanning diverse architectures: REINVENT (RNN + RL), Graph GA (graph-based genetic algorithm), GP BO (Gaussian process Bayesian optimization), SMILES GA (SMILES-based genetic algorithm), SELFIES-based VAEs, and others. The 23 objective tasks derive primarily from the GuacaMol benchmark.
Re-ranked Results and Augmented Hill-Climb Performance
The modified metrics substantially re-order the ranking of generative models:
SMILES-AHC achieves top performance on AUC Top-10 (Combined)*, where both property filters and diversity are enforced. The use of domain-informed hyperparameter selection ($\sigma = 60$) proves critical.
SMILES-AHC (data-driven hyperparameters) ranks first when accounting for property filters alone, diversity alone, or both combined, demonstrating that the AHC algorithm itself provides strong performance even without manual tuning.
REINVENT retains its first-place rank under property filters alone, suggesting that the minority of compounds staying within acceptable property space still perform well. However, it drops when diversity is also required.
Evolutionary algorithms (Graph GA, GP BO, SMILES GA) drop significantly under the new metrics. This is expected because rule-based methods are not constrained by the ZINC250k distribution and tend to propose molecules that diverge from drug-like chemical space.
Both AHC variants excel on empirically difficult tasks, including isomer-based tasks, Zaleplon MPO, and Sitagliptin MPO, where other methods struggle.
Limitations
The authors acknowledge several limitations:
- Results are preliminary because generative models have not undergone hyperparameter optimization against the new metrics
- Property filter thresholds are subjective, and the 10% de novo ECFP4 bit threshold was chosen by visual inspection
- Comparing rule-based models against distribution-based models using ZINC250k similarity introduces a bias toward distribution-based approaches
- Six objective task reference molecules sit in the lowest 0.01% of ZINC250k property space, raising questions about whether distribution-based models can reasonably optimize for these objectives
- Property filters and diversity could alternatively be incorporated directly into the objective function as additional oracles, though this would not necessarily produce the same results
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Pre-training | ZINC250k | ~250K molecules | Subset of ZINC15, provided by PMO benchmark |
| Evaluation | PMO benchmark tasks | 23 objectives | Derived primarily from GuacaMol |
Algorithms
- Augmented Hill-Climb: RL strategy from Thomas et al. (2022), patience of 5
- Hyperparameters (SMILES-AHC): batch size 256, $\sigma = 120$, $K = 0.25$
- Hyperparameters (SMILES-AHC)*: $\sigma = 60$ (domain-informed selection)
- Prior training: 5 epochs, batch size 128, SMILES notation
- Oracle budget: 10,000 evaluations per task
- Replicates: 5 per model per task
Models
- Architecture: Embedding (128) + 3x GRU (512), following REINVENT
- All 25 PMO benchmark models re-evaluated using original implementations
Evaluation
| Metric | Description | Notes |
|---|---|---|
| AUC Top-10 (Original) | Area under curve of average top 10 molecules | Standard PMO metric |
| AUC Top-10 (Filtered) | Original with MW/LogP and ECFP4 novelty filters | $\mu \pm 4\sigma$ from ZINC250k |
| AUC Top-10 (Diverse) | Top 10 selected with Tanimoto < 0.35 diversity | ECFP4 fingerprints |
| AUC Top-10 (Combined) | Both filters and diversity applied | Most stringent metric |
Hardware
Hardware requirements are not specified in the paper. The benchmark uses 10,000 oracle evaluations per task with 5 replicates, which is computationally modest compared to standard generative model training.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| MolScore | Code | MIT | Scoring and benchmarking framework by the first author |
| PMO Benchmark | Code | MIT | Original benchmark code and data |
Paper Information
Citation: Thomas, M., O’Boyle, N. M., Bender, A., & de Graaf, C. (2022). Re-evaluating sample efficiency in de novo molecule generation. arXiv preprint arXiv:2212.01385.
@misc{thomas2022reevaluating,
title={Re-evaluating sample efficiency in de novo molecule generation},
author={Thomas, Morgan and O'Boyle, Noel M. and Bender, Andreas and de Graaf, Chris},
year={2022},
eprint={2212.01385},
archivePrefix={arXiv},
primaryClass={cs.LG},
doi={10.48550/arxiv.2212.01385}
}