An Empirical Critique of Molecular Generation Evaluation
This is an Empirical paper that critically examines evaluation practices for molecular generative models. Rather than proposing a new generative method, the paper exposes systematic weaknesses in both distribution-learning metrics and goal-directed optimization scoring functions. The primary contributions are: (1) demonstrating that a trivially simple “AddCarbon” model can achieve near-perfect scores on widely used distribution-learning benchmarks, and (2) introducing an experimental framework with optimization scores and control scores that reveals model-specific and data-specific biases when ML models serve as scoring functions for goal-directed generation.
Evaluation Gaps in De Novo Molecular Design
The rapid growth of deep learning methods for molecular generation (RNN-based SMILES generators, VAEs, GANs, graph neural networks) created a need for standardized evaluation. Benchmarking suites like GuacaMol and MOSES introduced metrics for validity, uniqueness, novelty, KL divergence over molecular properties, and Frechet ChemNet Distance (FCD). For goal-directed generation, penalized logP became a common optimization target.
However, these metrics leave significant blind spots. Distribution-learning metrics do not detect whether a model merely copies training molecules with minimal modifications. Goal-directed benchmarks often use scoring functions that fail to capture the full requirements of drug discovery (synthetic feasibility, drug-likeness, absence of reactive substructures). When ML models serve as scoring functions, the problem worsens because generated molecules can exploit artifacts of the learned model rather than exhibiting genuinely desirable properties.
At the time of writing, wet-lab validations of generative models remained scarce, with only a handful of studies (Merk et al., Zhavoronkov et al.) demonstrating in vitro activity for generated compounds. The lack of rigorous evaluation left the field unable to distinguish meaningfully innovative methods from those that simply exploit metric weaknesses.
The Copy Problem and Control Score Framework
The paper introduces two key conceptual contributions.
The AddCarbon Model for Distribution-Learning
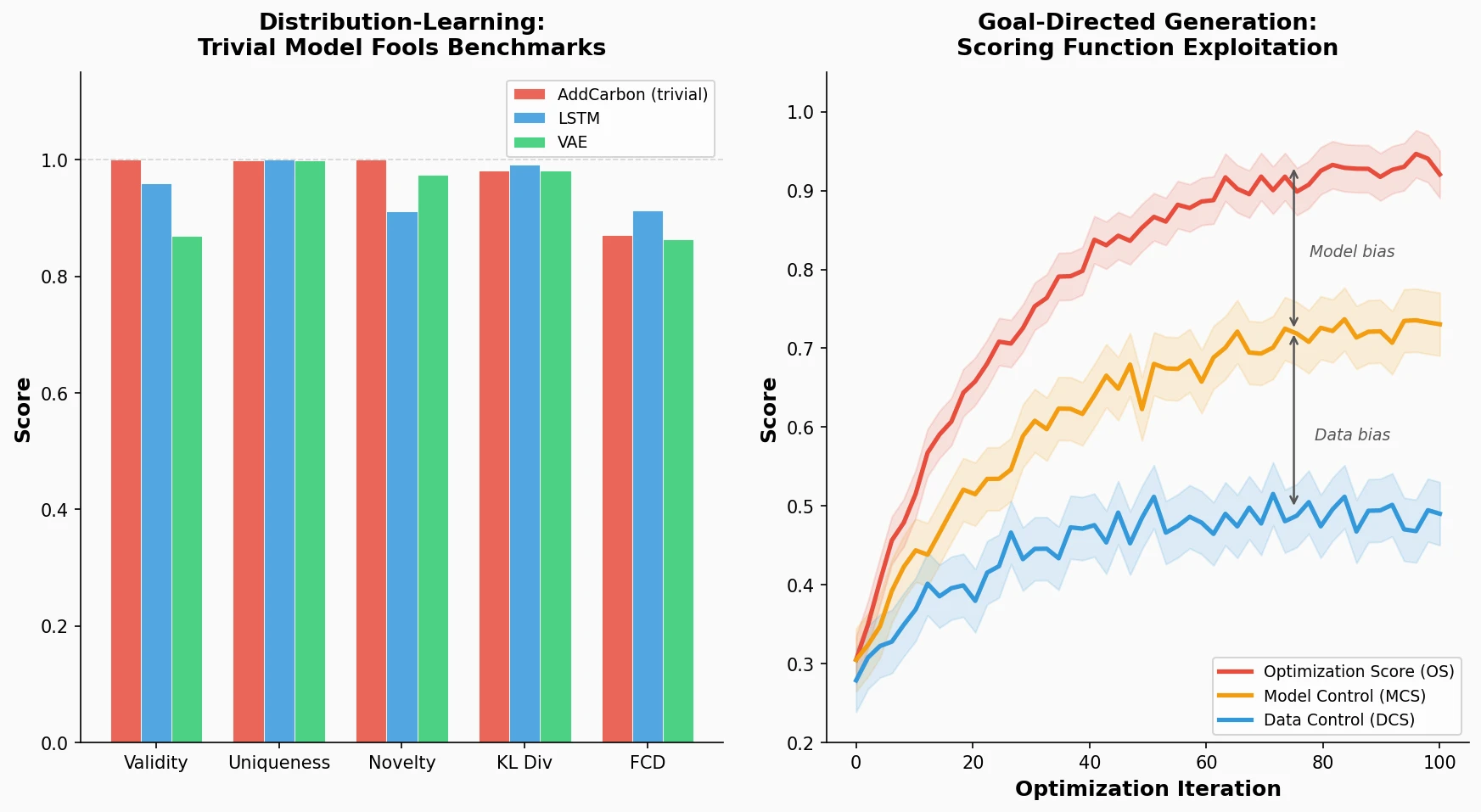
The AddCarbon model is deliberately trivial: it samples a molecule from the training set, inserts a single carbon atom at a random position in its SMILES string, and returns the result if it produces a valid, novel molecule. This model achieves near-perfect scores across most GuacaMol distribution-learning benchmarks:
| Benchmark | RS | LSTM | GraphMCTS | AAE | ORGAN | VAE | AddCarbon |
|---|---|---|---|---|---|---|---|
| Validity | 1.000 | 0.959 | 1.000 | 0.822 | 0.379 | 0.870 | 1.000 |
| Uniqueness | 0.997 | 1.000 | 1.000 | 1.000 | 0.841 | 0.999 | 0.999 |
| Novelty | 0.000 | 0.912 | 0.994 | 0.998 | 0.687 | 0.974 | 1.000 |
| KL divergence | 0.998 | 0.991 | 0.522 | 0.886 | 0.267 | 0.982 | 0.982 |
| FCD | 0.929 | 0.913 | 0.015 | 0.529 | 0.000 | 0.863 | 0.871 |
The AddCarbon model beats all baselines except the LSTM on the FCD metric, despite being practically useless. This exposes what the authors call the “copy problem”: current metrics check only for exact matches to training molecules, so minimal edits evade novelty detection. The authors argue that likelihood-based evaluation on hold-out test sets, analogous to standard practice in NLP, would provide a more comprehensive metric.
Control Scores for Goal-Directed Generation
For goal-directed generation, the authors introduce a three-score experimental design:
- Optimization Score (OS): Output of a classifier trained on data split 1, used to guide the molecular optimizer.
- Model Control Score (MCS): Output of a second classifier trained on split 1 with a different random seed. Divergence between OS and MCS quantifies model-specific biases.
- Data Control Score (DCS): Output of a classifier trained on data split 2. Divergence between OS and DCS quantifies data-specific biases.
This mirrors the training/test split paradigm in supervised learning. If a generator truly produces molecules with the desired bioactivity, the control scores should track the optimization score. Divergence between them indicates the optimizer is exploiting artifacts of the specific model or training data rather than learning generalizable chemical properties.
Experimental Setup: Three Targets, Three Generators
Targets and Data
The authors selected three biological targets from ChEMBL: Janus kinase 2 (JAK2), epidermal growth factor receptor (EGFR), and dopamine receptor D2 (DRD2). For each target, the data was split into two halves (split 1 and split 2) with balanced active/inactive ratios. Random forest classifiers using binary folded ECFP4 fingerprints (radius 2, size 1024) were trained to produce three scoring functions per target: the OS and MCS on split 1 (different random seeds), and the DCS on split 2.
Generators
Three molecular generators were evaluated:
- Graph-based Genetic Algorithm (GA): Iteratively applies random mutations and crossovers to a population of molecules, retaining the best in each generation. One of the top performers in GuacaMol.
- SMILES-LSTM: An autoregressive model that generates SMILES character by character, optimized via hill climbing (iteratively sampling, keeping top molecules, fine-tuning). Also a top GuacaMol performer.
- Particle Swarm Optimization (PS): Optimizes molecules in the continuous latent space of a SMILES-based sequence-to-sequence model.
Each optimizer was run 10 times per target dataset.
Score Divergence and Exploitable Biases
Optimization vs. Control Score Divergence
Across all three targets and all three generators, the OS consistently outpaced both control scores during optimization. The DCS sometimes stagnated or even decreased while the OS continued to climb. This divergence demonstrates that the generators exploit biases in the scoring function rather than discovering genuinely active compounds.
The MCS also diverged from the OS despite being trained on exactly the same data, confirming model-specific biases: the optimization exploits features unique to the particular random forest instance. The larger gap between OS and DCS (compared to OS and MCS) indicates that data-specific biases contribute more to the divergence than model-specific biases.
Chemical Space Migration
Optimized molecules migrated toward the region of split 1 actives (used to train the OS), as shown by t-SNE embeddings and nearest-neighbor Tanimoto similarity analysis. Optimized molecules had more similar neighbors in split 1 than in split 2, confirming data-specific bias. By the end of optimization, generated molecules occupied different regions of chemical space than known actives when measured by logP and molecular weight, with compounds from the same optimization run forming distinct clusters.
Quality of Generated Molecules
High-scoring generated molecules frequently contained problematic substructures: reactive dienes, nitrogen-fluorine bonds, long heteroatom chains that are synthetically infeasible, and highly uncommon functional groups. The LSTM optimizer showed a bias toward high molecular weight, low diversity, and high logP values. These molecules would be rejected by medicinal chemists despite their high optimization scores.
Key Takeaways
The authors emphasize several practical implications:
- Early stopping: Control scores can indicate when further optimization is exploiting biases rather than finding better molecules. Optimization should stop when control scores plateau.
- Scoring function iteration: In practice, generative models are “highly adept at exploiting” incomplete scoring functions, necessitating several iterations of generation and scoring function refinement.
- Synthetic accessibility: Even high-scoring molecules are useless if they cannot be synthesized. The authors consider this a major challenge for practical adoption.
- Likelihood-based evaluation: For distribution-learning, the authors recommend reporting test-set likelihoods for likelihood-based models, following standard NLP practice.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Bioactivity data | ChEMBL (JAK2, EGFR, DRD2) | See Table S1 | Binary classification tasks, split 50/50 |
| Distribution-learning | GuacaMol training set | Subset of ChEMBL | Used as starting population for GA and PS |
Algorithms
- Scoring function: Random forest classifier (scikit-learn) on binary ECFP4 fingerprints (size 1024, radius 2, RDKit)
- GA: Graph-based genetic algorithm from Jensen (2019)
- LSTM: SMILES-LSTM with hill climbing, pretrained model from GuacaMol
- PS: Particle swarm optimization in latent space of a sequence-to-sequence model (Winter et al. 2019)
- Each optimizer run 10 times per target
Evaluation
| Metric | Description | Notes |
|---|---|---|
| Optimization Score (OS) | RF classifier on split 1 | Guides optimization |
| Model Control Score (MCS) | RF on split 1, different seed | Detects model-specific bias |
| Data Control Score (DCS) | RF on split 2 | Detects data-specific bias |
| GuacaMol metrics | Validity, uniqueness, novelty, KL div, FCD | For distribution-learning |
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| ml-jku/mgenerators-failure-modes | Code | Unknown | Data, code, and results |
Hardware
Not specified in the paper.
Citation
@article{renz2019failure,
title={On failure modes in molecule generation and optimization},
author={Renz, Philipp and Van Rompaey, Dries and Wegner, J{\"o}rg Kurt and Hochreiter, Sepp and Klambauer, G{\"u}nter},
journal={Drug Discovery Today: Technologies},
volume={32-33},
pages={55--63},
year={2019},
publisher={Elsevier},
doi={10.1016/j.ddtec.2020.09.003}
}
Paper Information
Citation: Renz, P., Van Rompaey, D., Wegner, J. K., Hochreiter, S., & Klambauer, G. (2019). On failure modes in molecule generation and optimization. Drug Discovery Today: Technologies, 32-33, 55-63. https://doi.org/10.1016/j.ddtec.2020.09.003
Publication: Drug Discovery Today: Technologies, Volume 32-33, 2019
Additional Resources: