Semi-Supervised Data Augmentation for Molecular Tasks
This is a Method paper that introduces back translation, a semi-supervised technique from neural machine translation, to the domain of molecular generation. The primary contribution is a general-purpose data augmentation strategy that leverages large pools of unlabeled molecules (from databases like ZINC) to improve the performance of both sequence-based and graph-based models on molecule optimization and retrosynthesis prediction tasks.
Bridging the Labeled Data Gap in Molecular Generation
Molecular generation tasks, such as property optimization and retrosynthesis, require paired training data: an input molecule (or property specification) mapped to a desired output molecule. Obtaining these labeled pairs is expensive and labor-intensive. Meanwhile, enormous databases of unlabeled molecules exist. ZINC alone contains over 750 million compounds, and PubChem has 109 million.
Prior approaches to using unlabeled molecular data include variational autoencoders (VAEs) for learning latent representations, conditional recurrent neural networks for inverse design, and pretraining techniques borrowed from NLP. However, these methods either focus on representation learning rather than direct generation, or require task-specific architectural modifications. The authors identify back translation, a well-established technique in machine translation, as a natural fit for molecular generation tasks that can be treated as sequence-to-sequence mappings.
Back Translation as Molecular Data Augmentation
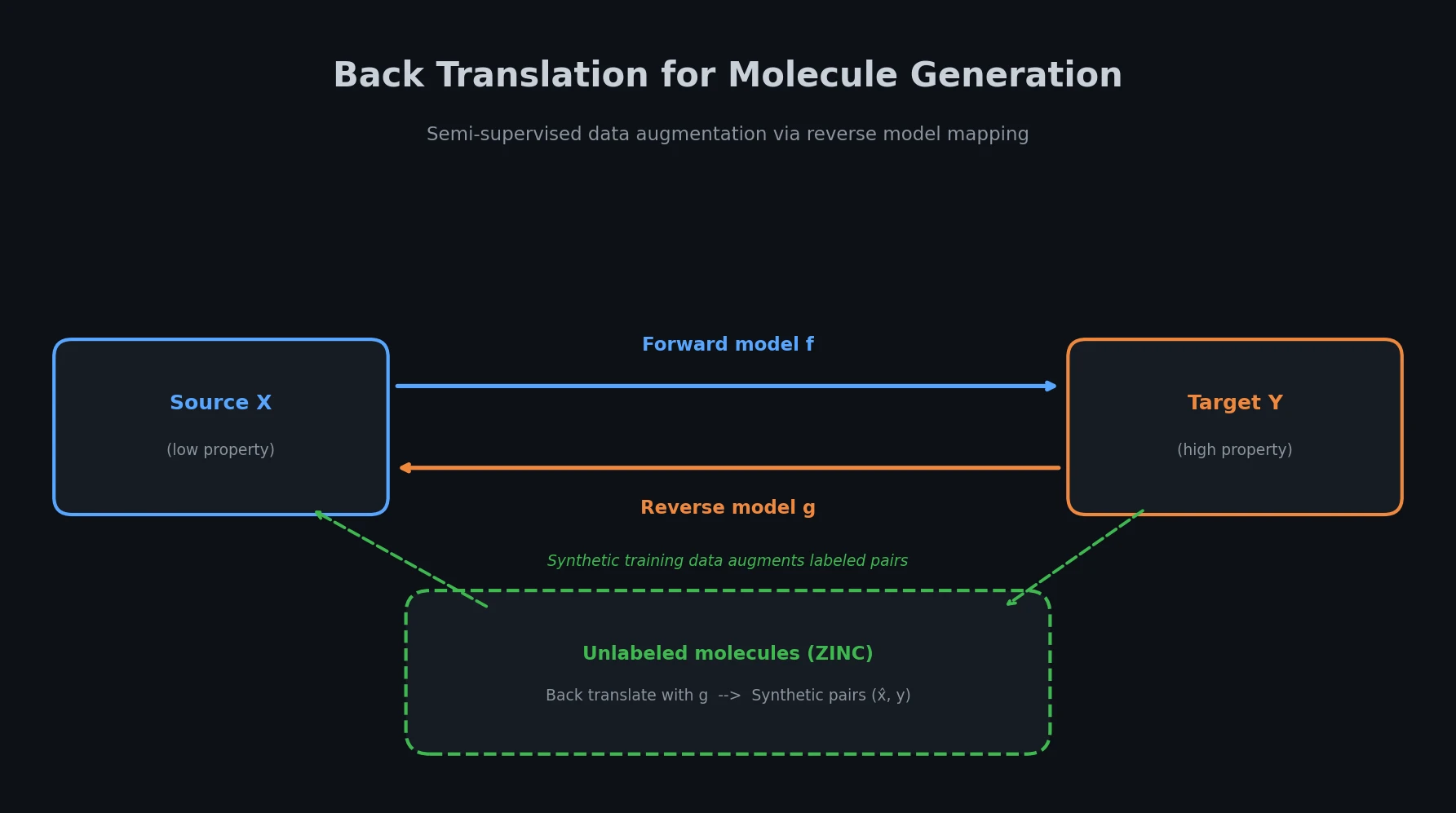
The core idea is straightforward. Given a main task that maps from source domain $\mathcal{X}$ to target domain $\mathcal{Y}$ (e.g., mapping low-QED molecules to high-QED molecules), the method trains a reverse model $g$ that maps from $\mathcal{Y}$ back to $\mathcal{X}$. This reverse model then “back translates” unlabeled molecules from $\mathcal{Y}$ to generate synthetic source molecules, creating pseudo-labeled training pairs.
The theoretical motivation comes from maximizing the reconstruction probability. Given an unlabeled molecule $y_u \in \mathcal{U}_y$, the logarithmic reconstruction probability through the reverse model $g$ and forward model $f$ is:
$$ \log P(y_u = \hat{y}_u \mid y_u; g, f) = \log \sum_{\hat{x}_u \in \mathcal{X}} P(\hat{x}_u \mid y_u; g) P(y_u = \hat{y}_u \mid \hat{x}_u; f) $$
Since summing over the exponentially large space $\mathcal{X}$ is intractable, the authors apply Jensen’s inequality to obtain a lower bound:
$$ \log P(y_u = \hat{y}_u \mid y_u; g, f) \geq \mathbb{E}_{\hat{x}_u \sim P(\cdot \mid y_u; g)} \log P(y_u = \hat{y}_u \mid \hat{x}_u; f) $$
This lower bound is optimized via Monte Carlo sampling in three steps:
Step 1: Train both forward model $f$ and reverse model $g$ on the labeled data $\mathcal{L}$:
$$ \begin{aligned} \min_{\theta_f} \sum_{(x,y) \in \mathcal{L}} -\log P(y \mid x; \theta_f) \\ \min_{\theta_g} \sum_{(x,y) \in \mathcal{L}} -\log P(x \mid y; \theta_g) \end{aligned} $$
Step 2: Use the trained reverse model $g$ to back translate each unlabeled molecule $y_u \in \mathcal{U}_y$, producing synthetic pairs:
$$ \hat{\mathcal{L}} = {(\hat{x}_u, y_u) \mid y_u \in \mathcal{U}_y, \hat{x}_u \text{ sampled from } P(\cdot \mid y_u; \theta_g)} $$
Step 3: Retrain the forward model $f$ on the combined labeled and synthetic data $\mathcal{L} \cup \hat{\mathcal{L}}$, warm-starting from the parameters obtained in Step 1:
$$ \min_{\theta_f^} \sum_{(x,y) \in \mathcal{L} \cup \hat{\mathcal{L}}} -\log P(y \mid x; \theta_f^) $$
A key practical finding is that data filtration matters. When using large amounts of unlabeled data (1M molecules), keeping only the synthetic pairs that satisfy the same constraints as the labeled data (e.g., similarity thresholds and property ranges) significantly improves performance over using all back-translated data unfiltered.
Experiments on Property Optimization and Retrosynthesis
Molecular Property Improvement
The authors evaluate on four tasks from Jin et al. (2019, 2020), each requiring the model to improve a specific molecular property while maintaining structural similarity (measured by Dice similarity on Morgan fingerprints):
- LogP (penalized partition coefficient): two settings with similarity thresholds $\delta \geq 0.4$ and $\delta \geq 0.6$
- QED (quantitative estimation of drug-likeness): translate molecules from QED range [0.7, 0.8] to [0.9, 1.0]
- DRD2 (dopamine type 2 receptor activity): translate inactive ($P < 0.5$) to active ($P \geq 0.5$)
Two backbone architectures are tested: a Transformer (6 layers, 4 heads, 128-dim embeddings, 512-dim FFN) and HierG2G, a hierarchical graph-to-graph translation model. Unlabeled molecules are sampled from ZINC at 250K and 1M scales.
| Method | LogP ($\delta \geq 0.6$) | LogP ($\delta \geq 0.4$) | QED (%) | DRD2 (%) |
|---|---|---|---|---|
| JT-VAE | 0.28 | 1.03 | 8.8 | 3.4 |
| GCPN | 0.79 | 2.49 | 9.4 | 4.4 |
| JTNN | 2.33 | 3.55 | 59.9 | 77.8 |
| Transformer baseline | 2.45 | 3.69 | 71.9 | 60.2 |
| +BT (1M, filtered) | 2.86 | 4.41 | 82.9 | 67.4 |
| HierG2G baseline | 2.49 | 3.98 | 76.9 | 85.9 |
| +BT (250K, filtered) | 2.75 | 4.24 | 79.1 | 87.3 |
Retrosynthesis Prediction
On the USPTO-50K benchmark (50K reactions, 10 reaction types, 80/10/10 train/val/test split), the method is applied to Transformer and GLN (Graph Logic Network) backbones. For other approaches to this benchmark, see Tied Two-Way Transformers and Data Transfer for Retrosynthesis. Unlabeled reactant sets are constructed by sampling molecules from ZINC and concatenating them following the training data’s reactant count distribution ($N_1 : N_2 : N_3 = 29.3% : 70.4% : 0.3%$).
| Method | Top-1 | Top-3 | Top-5 | Top-10 |
|---|---|---|---|---|
| Reaction type given | ||||
| GLN | 64.2 | 79.1 | 85.2 | 90.0 |
| Ours + GLN | 67.9 | 82.5 | 87.3 | 91.5 |
| Transformer | 52.2 | 68.2 | 72.7 | 77.4 |
| Ours + Transformer | 55.9 | 72.8 | 77.8 | 79.7 |
| Reaction type unknown | ||||
| GLN | 52.5 | 69.0 | 75.6 | 83.7 |
| Ours + GLN | 54.7 | 70.2 | 77.0 | 84.4 |
| Transformer | 37.9 | 57.3 | 62.7 | 68.1 |
| Ours + Transformer | 43.5 | 58.8 | 64.6 | 69.7 |
The improvements are largest at lower $k$ values (top-1 and top-3), suggesting that back translation helps the model make more precise high-confidence predictions.
Ablation Studies
Effect of unlabeled data size: On retrosynthesis with Transformer, performance improves as unlabeled data increases from 50K to 250K, then plateaus or declines beyond 250K. The authors attribute this to noise in the back-translated data outweighing the benefits at larger scales.
Effect of labeled data size: With only 5K labeled samples, adding back-translated data hurts performance because the reverse model is too weak to generate useful synthetic data. As labeled data increases (10K, 25K, 50K), the benefit of back translation grows. This confirms that the method requires a reasonably well-trained reverse model to be effective.
Data filtration: Using 1M unfiltered back-translated molecules sometimes hurts performance (e.g., QED drops from 71.9% to 75.1% vs. 82.9% with filtering), while filtering to enforce the same constraints as the labeled data recovers and exceeds the 250K filtered results.
Consistent Gains Across Architectures and Tasks
The method achieves state-of-the-art results on all four molecular property improvement tasks and the USPTO-50K retrosynthesis benchmark at time of publication. Several observations stand out:
- Architecture agnosticism: Back translation improves both sequence-based (Transformer) and graph-based (HierG2G, GLN) models, confirming that the approach is independent of the underlying architecture.
- Filtration is essential at scale: Unfiltered 1M back-translated data can degrade performance, but filtered data at the same scale consistently outperforms smaller unfiltered sets.
- Training overhead is moderate: On the DRD2 task, back translation with Transformer takes about 2.5x the supervised training time (11.0h vs. 8.5h for initial training), with the back-translation step itself taking under 1 hour.
- Diversity and novelty increase: Back translation improves both diversity (average pairwise distance among generated molecules) and novelty (fraction of generated molecules not seen in training) across QED and DRD2 tasks.
The authors acknowledge limitations: the method does not form a closed loop between forward and reverse models (as in dual learning approaches), and the data filtration strategy is rule-based rather than learned. They suggest joint training of forward and reverse models and learned filtration as future directions.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Training (property improvement) | Jin et al. (2019, 2020) datasets | 34K-99K pairs | LogP, QED, DRD2 tasks |
| Training (retrosynthesis) | USPTO-50K | 40K reactions | 80/10/10 split from Dai et al. (2019) |
| Unlabeled molecules | ZINC | 250K or 1M | Randomly sampled |
| Evaluation | Same as training | 800-1000 test samples | Per-task test sets |
Algorithms
- Back translation with optional data filtration
- Beam search with $k=20$ for inference
- Random sampling for back-translation step (Equation 5)
- Dice similarity on Morgan fingerprints for similarity constraint
Models
- Transformer: 6 layers, 4 attention heads, 128-dim embeddings, 512-dim FFN (for property improvement); 4 layers, 8 heads, 256-dim embeddings, 2048-dim FFN (for retrosynthesis)
- HierG2G: Settings from Jin et al. (2020)
- GLN: Settings from Dai et al. (2019)
Evaluation
| Metric | Task | Best Value | Baseline | Notes |
|---|---|---|---|---|
| LogP improvement | LogP ($\delta \geq 0.6$) | 2.86 | 2.49 (HierG2G) | Transformer + BT(1M, filtered) |
| LogP improvement | LogP ($\delta \geq 0.4$) | 4.41 | 3.98 (HierG2G) | Transformer + BT(1M, filtered) |
| Success rate | QED | 82.9% | 76.9% (HierG2G) | Transformer + BT(1M, filtered) |
| Success rate | DRD2 | 87.3% | 85.9% (HierG2G) | HierG2G + BT(250K, filtered) |
| Top-1 accuracy | USPTO-50K (known type) | 67.9% | 64.2% (GLN) | Ours + GLN |
Hardware
The paper reports training times (8.5h for Transformer, 16.8h for HierG2G on DRD2 with 1M unlabeled data) but does not specify the GPU hardware used.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| BT4MolGen | Code | MIT | Official implementation in Python |
Paper Information
Citation: Fan, Y., Xia, Y., Zhu, J., Wu, L., Xie, S., & Qin, T. (2021). Back translation for molecule generation. Bioinformatics, 38(5), 1244-1251. https://doi.org/10.1093/bioinformatics/btab817
@article{fan2022back,
title={Back translation for molecule generation},
author={Fan, Yang and Xia, Yingce and Zhu, Jinhua and Wu, Lijun and Xie, Shufang and Qin, Tao},
journal={Bioinformatics},
volume={38},
number={5},
pages={1244--1251},
year={2021},
publisher={Oxford University Press},
doi={10.1093/bioinformatics/btab817}
}