A Transformer Approach to Learned Molecular Fingerprints
This is a Method paper that introduces SMILES Transformer (ST), a Transformer-based sequence-to-sequence model pre-trained on unlabeled SMILES strings to produce continuous, data-driven molecular fingerprints. The primary contribution is demonstrating that unsupervised pre-training on chemical text representations yields fingerprints that generalize well under low-data conditions, outperforming both rule-based fingerprints (ECFP) and graph convolution models on several MoleculeNet benchmarks. A secondary contribution is the Data Efficiency Metric (DEM), a scalar metric for evaluating model performance across varying training set sizes.
The Low-Data Problem in Molecular Property Prediction
Machine learning for drug discovery depends on molecular representations, but labeled datasets of experimentally validated properties are typically small. Conventional approaches fall into two camps: rule-based fingerprints like ECFP that hash substructures into sparse binary vectors, and graph-based methods like GraphConv that learn representations end-to-end. Rule-based fingerprints perform poorly with shallow models or limited data, while graph-based methods are designed for large fully-labeled settings.
Pre-training on unlabeled data had shown strong results in NLP (ELMo, BERT, XLNet), and prior work in cheminformatics had explored RNN-based and VAE-based pre-training on SMILES (Seq2Seq fingerprints, Grammar VAE, heteroencoders). However, none of these studies systematically evaluated performance in small-data settings. Honda et al. fill this gap by applying Transformer-based pre-training to SMILES and measuring data efficiency explicitly.
Transformer Pre-training on SMILES with Pooled Fingerprint Extraction
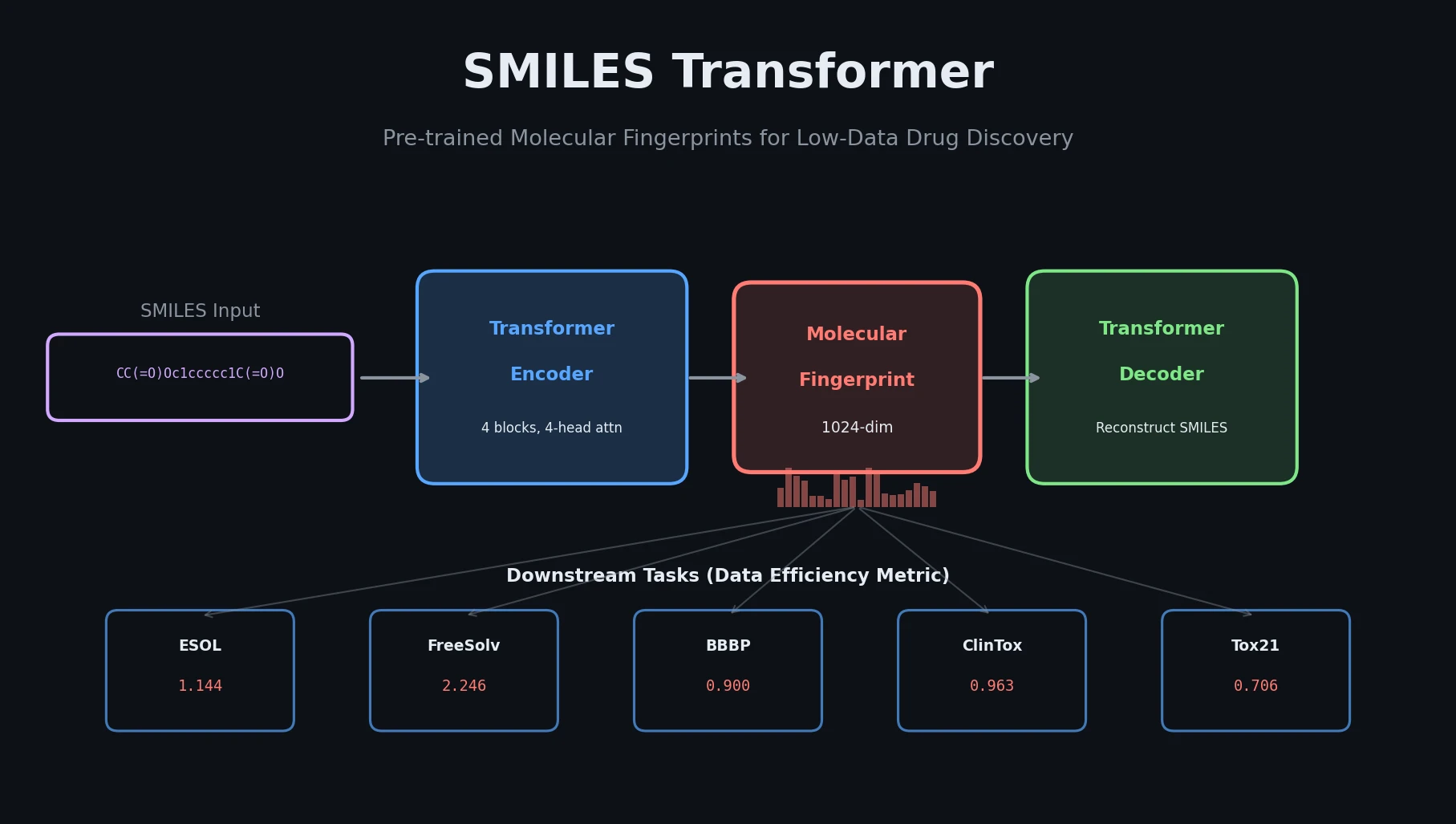
The core innovation is a Transformer encoder-decoder architecture pre-trained as an autoencoder on SMILES strings, with a specific fingerprint extraction strategy that pools the encoder outputs into a fixed-length vector.
Architecture
The model uses 4 Transformer blocks for both the encoder and decoder, each with 4-head attention and 256 embedding dimensions plus 2 linear layers. Input SMILES are tokenized at the symbol level (e.g., ‘c’, ‘Br’, ‘=’, ‘(’, ‘2’) and one-hot encoded. Following Vaswani et al. (2017), the input uses the sum of token encoding and positional encoding.
Pre-training
The model is pre-trained on 861,000 unlabeled SMILES sampled from ChEMBL24 to minimize cross-entropy between input and output SMILES (i.e., reconstruction). SMILES enumeration (Bjerrum, 2017) randomly generates non-canonical SMILES at each epoch to reduce representation bias. Training runs for 5 epochs with Adam optimization, reaching a perplexity of 1.0 (perfect decoding).
Fingerprint Extraction
Since the Transformer outputs symbol-level (atom-level) representations, a pooling strategy produces molecule-level fingerprints. Four vectors are concatenated:
- Mean-pooled output of the last encoder layer
- Max-pooled output of the last encoder layer
- First output token of the last encoder layer
- First output token of the penultimate encoder layer
This produces a 1024-dimensional fingerprint, matching the dimensionality of ECFP for fair comparison.
Data Efficiency Metric
The paper proposes DEM to measure how well a model performs across different training set sizes:
$$ M_{DE}(f, m) = \frac{1}{|I|} \sum_{i \in I} m(f_i, X_i, Y_i) $$
where $f_i$ is the model trained on the fraction $i$ of training data, $m$ is the task metric, and $I = {0.0125, 0.025, 0.05, 0.1, 0.2, 0.4, 0.8}$ doubles the training percentage at each step. This captures average performance across a range of data availability, giving a single scalar that balances accuracy and data efficiency.
Benchmarking Across MoleculeNet with Data Efficiency Focus
Datasets
The evaluation uses 10 datasets from MoleculeNet spanning three categories:
| Category | Dataset | Tasks | Type | Molecules | Metric |
|---|---|---|---|---|---|
| Physical chemistry | ESOL | 1 | Regression | 1,128 | RMSE |
| Physical chemistry | FreeSolv | 1 | Regression | 643 | RMSE |
| Physical chemistry | Lipophilicity | 1 | Regression | 4,200 | RMSE |
| Biophysics | MUV | 17 | Classification | 93,127 | PRC-AUC |
| Biophysics | HIV | 1 | Classification | 41,913 | ROC-AUC |
| Biophysics | BACE | 1 | Classification | 1,522 | ROC-AUC |
| Physiology | BBBP | 1 | Classification | 2,053 | ROC-AUC |
| Physiology | Tox21 | 12 | Classification | 8,014 | ROC-AUC |
| Physiology | SIDER | 27 | Classification | 1,427 | ROC-AUC |
| Physiology | ClinTox | 2 | Classification | 1,491 | ROC-AUC |
Baselines
- ECFP4: Rule-based extended-connectivity fingerprint with 1024 dimensions
- RNNS2S: RNN-based Seq2Seq pre-trained fingerprint (3-layer bidirectional GRU, same pre-training data as ST)
- GraphConv: Graph convolution network trained end-to-end on labeled data
Experimental Setup
All fingerprint methods use a simple MLP classifier/regressor from scikit-learn with default hyperparameters to isolate the fingerprint quality from model capacity. Datasets are randomly split (stratified for classification), and results are averaged over 20 trials. Note that random splits are used rather than scaffold splits for the DEM experiments.
Data Efficiency Results (DEM)
| Dataset | ST+MLP | ECFP+MLP | RNNS2S+MLP | GraphConv |
|---|---|---|---|---|
| ESOL (RMSE, lower is better) | 1.144 | 1.741 | 1.317 | 1.673 |
| FreeSolv (RMSE, lower is better) | 2.246 | 3.043 | 2.987 | 3.476 |
| Lipophilicity (RMSE, lower is better) | 1.169 | 1.090 | 1.219 | 1.062 |
| MUV (PRC-AUC, higher is better) | 0.009 | 0.036 | 0.010 | 0.004 |
| HIV (ROC-AUC, higher is better) | 0.683 | 0.697 | 0.682 | 0.723 |
| BACE (ROC-AUC, higher is better) | 0.719 | 0.769 | 0.717 | 0.744 |
| BBBP (ROC-AUC, higher is better) | 0.900 | 0.760 | 0.884 | 0.795 |
| Tox21 (ROC-AUC, higher is better) | 0.706 | 0.616 | 0.702 | 0.687 |
| SIDER (ROC-AUC, higher is better) | 0.559 | 0.588 | 0.558 | 0.557 |
| ClinTox (ROC-AUC, higher is better) | 0.963 | 0.515 | 0.904 | 0.936 |
ST achieves the best DEM in 5 of 10 datasets (ESOL, FreeSolv, BBBP, Tox21, ClinTox), with particularly strong margins on ClinTox (+0.027 over GraphConv) and BBBP (+0.016 over RNNS2S).
Linear Model Experiments
To further isolate fingerprint quality, the authors replace MLP with ridge/logistic regression with L2 penalty. On 8 datasets (excluding MUV and SIDER due to class imbalance issues), ST achieves best DEM in 5 of 8, confirming the fingerprint quality holds regardless of downstream model.
Stratified Analysis by Molecule Size
On BBBP stratified by SMILES length, ST’s ROC-AUC increases with longer SMILES, similar to RNNS2S but unlike GraphConv which shows stable performance across lengths. This suggests text-based models extract richer information from longer sequences.
Comparison with Record Scores (Large Data)
Under the large-data setting (80/10/10 train/val/test split with hyperparameter tuning via Optuna), ST achieves first place only in ClinTox (0.954) but performs comparably to ECFP and graph-based models on the other datasets. This confirms that ST’s main advantage is in the low-data regime.
Strong Low-Data Performance with Caveats on Scalability
Key Findings
- Transformer-based unsupervised pre-training on SMILES produces fingerprints that excel in low-data molecular property prediction, achieving best data efficiency on 5 of 10 MoleculeNet tasks.
- The advantage is most pronounced on small datasets (ESOL with 1,128 molecules, FreeSolv with 643, BBBP with 2,053, ClinTox with 1,491) where pre-training enables good generalization.
- With sufficient labeled data and hyperparameter tuning, ST fingerprints perform comparably to (but do not surpass) graph-based methods.
- Longer SMILES provide richer information for text-based models, as shown by the stratified analysis on BBBP.
Limitations
- Random splits are used for most DEM experiments rather than scaffold splits, which may inflate performance estimates for drug discovery applications where training and test molecules are structurally distinct.
- The pre-training corpus (861K SMILES from ChEMBL24) is relatively small by modern standards.
- MUV performance is poor across all methods (PRC-AUC near zero), suggesting the DEM framework may not be informative for extremely imbalanced or noisy datasets.
- No comparison with BERT-style masked language model pre-training, which later work (ChemBERTa) would show as a viable alternative.
Future Directions
The authors propose three directions: (1) replacing the Transformer with Transformer-XL to handle longer SMILES, (2) multi-task pre-training that jointly predicts molecular descriptors (e.g., molecular weight, LogP) alongside SMILES reconstruction, and (3) better exploitation of enumerated SMILES to constrain the latent space.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Pre-training | ChEMBL24 | 861,000 SMILES | Unlabeled, randomly sampled |
| Evaluation | MoleculeNet (10 datasets) | 643 to 93,127 molecules | See Table 1 for per-dataset details |
Algorithms
- Transformer encoder-decoder: 4 blocks each, 4-head attention, 256 embedding dimensions
- Pre-training: 5 epochs, Adam optimizer, cross-entropy loss, SMILES enumeration for augmentation
- Fingerprint: 1024 dimensions from concatenated mean pool, max pool, and first-token outputs
- Downstream: scikit-learn MLP (default hyperparameters) for DEM experiments; ridge/logistic regression for linear model experiments; Optuna for hyperparameter search in large-data comparison
Models
| Artifact | Type | License | Notes |
|---|---|---|---|
| smiles-transformer | Code | MIT | Official implementation (Jupyter notebooks) |
Evaluation
- DEM averaged over 7 training fractions (1.25% to 80%), 20 trials each
- Random splits for DEM; scaffold splits for HIV, BACE, BBBP in large-data comparison
- Metrics: RMSE (regression), ROC-AUC or PRC-AUC (classification) per MoleculeNet conventions
Hardware
The paper does not specify GPU type or training time for the pre-training phase.
Paper Information
Citation: Honda, S., Shi, S., & Ueda, H. R. (2019). SMILES Transformer: Pre-trained Molecular Fingerprint for Low Data Drug Discovery. arXiv preprint arXiv:1911.04738.
@article{honda2019smiles,
title={SMILES Transformer: Pre-trained Molecular Fingerprint for Low Data Drug Discovery},
author={Honda, Shion and Shi, Shoi and Ueda, Hiroki R.},
journal={arXiv preprint arXiv:1911.04738},
year={2019}
}