Pre-Training Transformers on SMILES for Molecular Properties
SMILES-BERT is a Method paper that introduces a BERT-inspired pre-training and fine-tuning framework for molecular property prediction. The primary contribution is adapting the masked language model paradigm from NLP to SMILES strings, enabling a Transformer encoder to learn molecular representations from large-scale unlabeled data before fine-tuning on smaller labeled datasets.
Limited Labels in Molecular Property Prediction
Molecular property prediction is central to drug discovery and chemical design, but obtaining labeled data requires expensive biological assays. Deep learning methods for this task fall into three categories: manually designed fingerprints (e.g., ECFP), graph-based methods (GCNs operating on molecular graphs), and sequence-based methods (RNNs or CNNs operating on SMILES strings).
Prior unsupervised approaches like Seq2seq Fingerprint used an encoder-decoder architecture to learn representations from unlabeled SMILES, but the decoder acts as scaffolding that consumes GPU memory during pre-training without contributing to downstream prediction. The semi-supervised Seq3seq Fingerprint improved on this by incorporating labeled data, but retained the encoder-decoder inefficiency. RNN-based methods also suffer from difficulty in parallel training and require careful tuning (gradient clipping, early stopping) to converge.
The authors identify two motivations: (1) building a semi-supervised model that effectively leverages large pools of unlabeled SMILES to improve prediction with limited labels, and (2) designing an architecture where the entire pre-trained model participates in fine-tuning (no wasted decoder parameters) and naturally supports parallel training.
Masked SMILES Recovery with Transformer Encoders
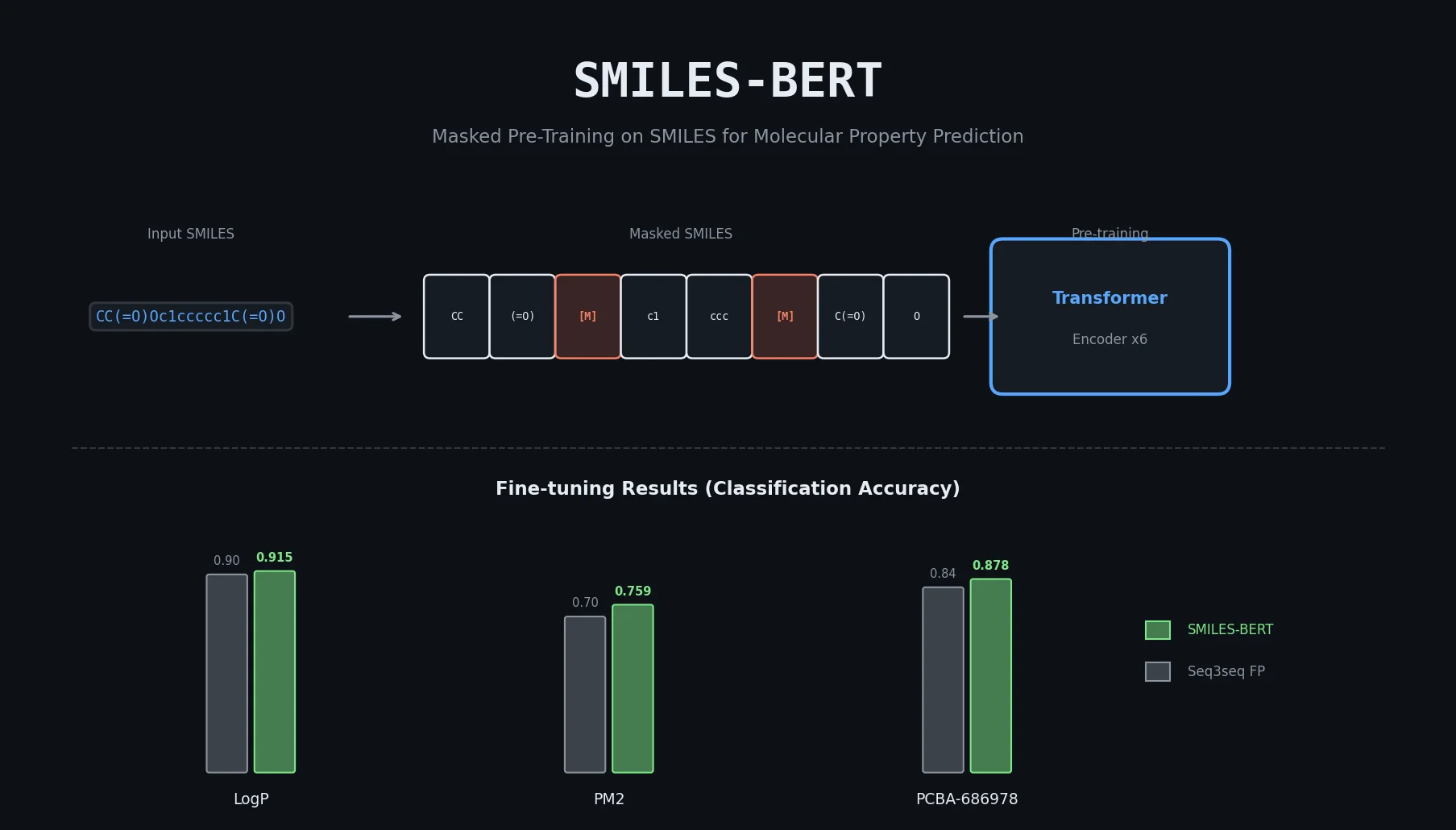
The core innovation is the Masked SMILES Recovery pre-training task, directly analogous to BERT’s masked language modeling. The model architecture is a stack of Transformer encoder layers, making it fully convolutional and parallelizable.
Architecture
SMILES-BERT uses 6 Transformer encoder layers, each with 4-head multi-head self-attention and feed-forward dimension of 1024. Each Transformer layer contains three components: a pre-attention feed-forward network, a self-attention layer, and a post-attention feed-forward network, all followed by layer normalization with residual connections.
The self-attention mechanism uses scaled dot-product attention:
$$ Z = \text{Softmax}\left(\frac{(XW^{Q})(XW^{K})^{T}}{\sqrt{d_{k}}}\right) XW^{V} $$
where $X \in \mathbb{R}^{N \times M}$ is the input feature matrix, $W^{Q}$, $W^{K}$, $W^{V} \in \mathbb{R}^{M \times d_{k}}$ are the query, key, and value weight matrices, and $\sqrt{d_{k}}$ is the scaling factor.
Input SMILES are tokenized at the character level with token embeddings and positional embeddings. A special <GO> token is prepended to each SMILES, and its output representation is used for downstream classification/regression after fine-tuning.
Pre-training: Masked SMILES Recovery
Following BERT’s masking strategy, 15% of tokens in each SMILES are selected for masking (minimum one per SMILES). Of the selected tokens:
- 85% are replaced with a
<MASK>token - 10% are replaced with a random token from the vocabulary
- 5% are kept unchanged
The model is trained to recover the original tokens at masked positions. The loss is computed only on the masked token outputs.
Fine-tuning
After pre-training, a classifier or regressor head is added to the <GO> token output. The entire model (all Transformer layers plus the new head) is fine-tuned on the labeled dataset.
Key differences from the original BERT:
- Only the Masked SMILES Recovery task is used (BERT’s next sentence prediction is dropped since SMILES have no consecutive-sentence structure)
- Segment embeddings are removed
- The architecture is smaller (6 layers, 4 heads, 1024 FFN dim) since SMILES have a much smaller vocabulary and shorter sequences than natural language
The authors compared this configuration against a larger BERT-base setup (12 layers, 12 heads, 3072 FFN dim) and found no meaningful performance difference, confirming that the smaller model is sufficient for SMILES.
Experimental Setup and Baseline Comparisons
Pre-training Data
SMILES-BERT was pre-trained on the ZINC database with 18,671,355 training SMILES, 10,000 for validation, and 10,000 for evaluation. Pre-training ran for 10 epochs using the Adam optimizer with a warm-up strategy (learning rate from $10^{-9}$ to $10^{-4}$ over 4,000 steps, then inverse-square-root decay). Batch size was 256 and dropout was 0.1. The pre-training masked SMILES exact recovery rate reached 82.85% on the validation set.
Fine-tuning Datasets
| Dataset | Source | Size | Task | Metric |
|---|---|---|---|---|
| LogP | NCATS/NIH | 10,850 | Classification (threshold 1.88) | Accuracy |
| PM2 | NCATS/NIH | 323,242 | Classification (threshold 0.024896) | Accuracy |
| PCBA-686978 | PubChem | 302,175 | Classification | Accuracy |
All datasets were split 80/10/10 for train/validation/test. Fine-tuning used Adam with a fixed learning rate for 50 epochs, selecting the best model on validation data.
Baselines
- Circular Fingerprint (CircularFP): Manually designed hash-based fingerprint (ECFP family)
- Neural Fingerprint (NeuralFP): Graph-based neural network replacing hash functions with learned layers
- Seq2seq Fingerprint (Seq2seqFP): Unsupervised encoder-decoder model on SMILES
- Seq3seq Fingerprint (Seq3seqFP): Semi-supervised encoder-decoder model on SMILES
Results
| Method | LogP | PM2 | PCBA-686978 |
|---|---|---|---|
| CircularFP | ~0.90 | 0.6858 | ~0.82 |
| NeuralFP | ~0.90 | 0.6802 | ~0.82 |
| Seq2seqFP | ~0.87 | 0.6112 | ~0.80 |
| Seq3seqFP | ~0.90 | 0.7038 | ~0.84 |
| SMILES-BERT | 0.9154 | 0.7589 | 0.8784 |
SMILES-BERT outperformed all baselines on all three datasets. The improvement over Seq3seqFP was approximately 2% on LogP, 5.5% on PM2, and 3.8% on PCBA-686978. The results on PM2 (the largest labeled dataset) show that pre-training benefits persist even with substantial labeled data.
Structure Study
| Configuration | Layers | Attention Heads | FFN Dim | LogP Accuracy |
|---|---|---|---|---|
| SMILES-BERT | 6 | 4 | 1024 | 0.9154 |
| SMILES-BERT (large) | 12 | 12 | 3072 | 0.9147 |
The larger configuration provided no improvement, supporting the choice of the smaller, more efficient architecture.
Findings, Limitations, and Future Directions
SMILES-BERT demonstrated that BERT-style masked pre-training on SMILES strings produces transferable molecular representations that improve property prediction across datasets of varying sizes and property types.
Key findings:
- The Masked SMILES Recovery pre-training task transfers effectively to molecular property prediction
- The full model participates in fine-tuning (no wasted decoder), making SMILES-BERT more parameter-efficient than encoder-decoder alternatives
- A smaller Transformer configuration (6 layers, 4 heads) matches the performance of a BERT-base-sized model for SMILES data
- Pre-training on ~18.7M SMILES from ZINC provides robust initialization across different downstream tasks
Limitations: The evaluation uses only classification accuracy as the metric, without reporting AUC-ROC, F1, or other metrics common in molecular property prediction. The comparison is limited to four baselines, and two of the three evaluation datasets (LogP, PM2) are non-public NIH datasets. The paper does not explore different pre-training dataset sizes or ablate the masking strategy. Only classification tasks are evaluated, though the architecture supports regression.
Future work: The authors propose incorporating Quantitative Estimate of Druglikeness (QED) prediction as an additional pre-training task to warm up the model’s classification capability, analogous to BERT’s next sentence prediction.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Pre-training | ZINC | 18,671,355 SMILES | Publicly available database |
| Fine-tuning | LogP | 10,850 | Non-public, from NCATS/NIH |
| Fine-tuning | PM2 | 323,242 | Non-public, from NCATS/NIH |
| Fine-tuning | PCBA-686978 | 302,175 | Public, from PubChem BioAssay |
Algorithms
- Pre-training: Adam optimizer, warm-up for 4,000 steps ($10^{-9}$ to $10^{-4}$), inverse-square-root LR schedule, batch size 256, dropout 0.1, 10 epochs
- Fine-tuning: Adam optimizer, fixed LR (insensitive to choice among $10^{-5}$, $10^{-6}$, $10^{-7}$), 50 epochs, best model on validation
Models
- 6 Transformer encoder layers, 4-head multi-head attention, FFN dim 1024
- Token embedding + positional embedding,
<GO>special token - Implemented with FairSeq (Facebook AI Research Sequence-to-Sequence Toolkit)
Evaluation
| Metric | SMILES-BERT | Best Baseline (Seq3seqFP) | Notes |
|---|---|---|---|
| LogP Accuracy | 0.9154 | ~0.90 | ~2% improvement |
| PM2 Accuracy | 0.7589 | 0.7038 | ~5.5% improvement |
| PCBA Accuracy | 0.8784 | ~0.84 | ~3.8% improvement |
Hardware
The paper mentions GPU training and NVIDIA GPU donation in acknowledgments but does not specify the exact GPU model or training time beyond noting that pre-training on a single GPU takes over a week for 10 epochs.
| Artifact | Type | License | Notes |
|---|---|---|---|
| No public code or model release identified | - | - | Paper does not provide a GitHub link or model checkpoint |
Reproducibility status: Partially Reproducible. The ZINC pre-training data is public and the architecture is described in detail, but no code or pre-trained weights are released. Two of three evaluation datasets (LogP, PM2) are non-public.
Paper Information
Citation: Wang, S., Guo, Y., Wang, Y., Sun, H., & Huang, J. (2019). SMILES-BERT: Large Scale Unsupervised Pre-Training for Molecular Property Prediction. In Proceedings of the 10th ACM International Conference on Bioinformatics, Computational Biology and Health Informatics (ACM-BCB ‘19), 429-436. https://doi.org/10.1145/3307339.3342186
@inproceedings{wang2019smilesbert,
title={SMILES-BERT: Large Scale Unsupervised Pre-Training for Molecular Property Prediction},
author={Wang, Sheng and Guo, Yuzhi and Wang, Yuhong and Sun, Hongmao and Huang, Junzhou},
booktitle={Proceedings of the 10th ACM International Conference on Bioinformatics, Computational Biology and Health Informatics},
pages={429--436},
year={2019},
publisher={ACM},
doi={10.1145/3307339.3342186}
}