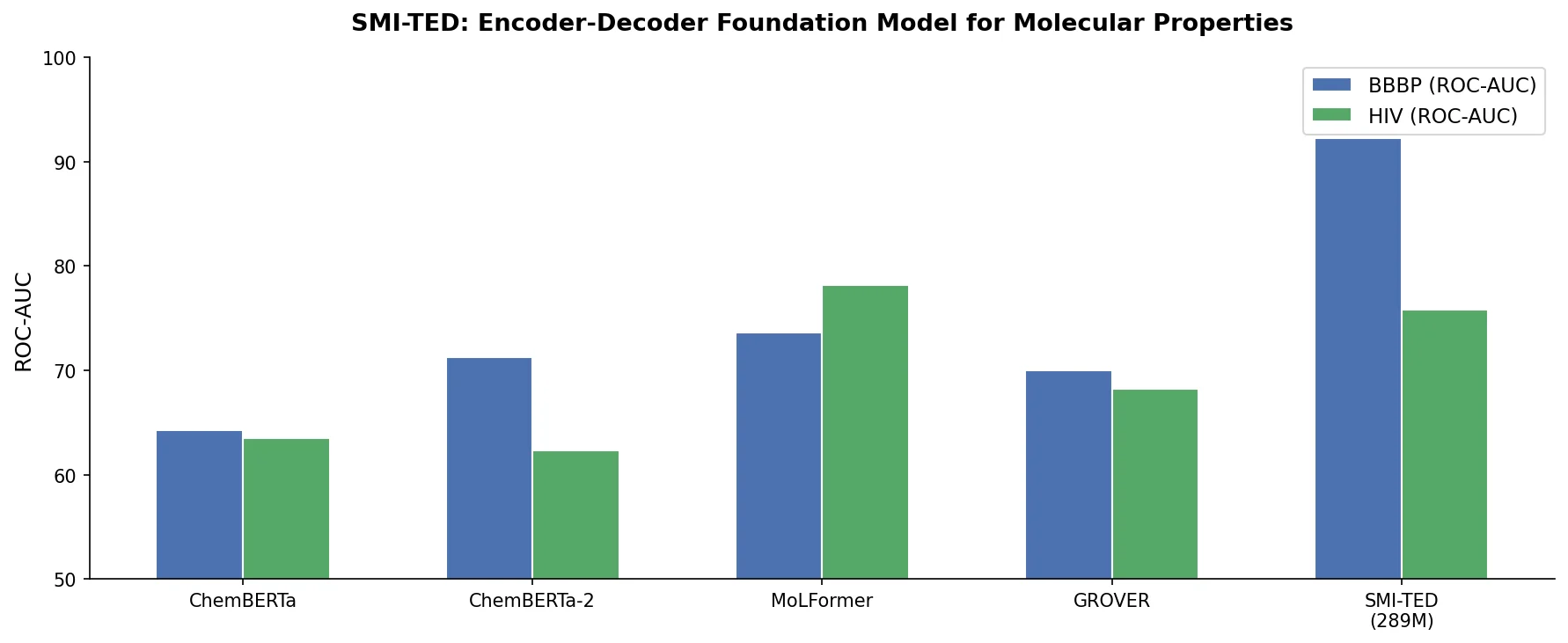
An Encoder-Decoder Chemical Foundation Model Family
SMI-TED is a Method paper that introduces a family of encoder-decoder transformer-based foundation models for chemistry. The primary contribution is the SMI-TED289M architecture, a 289-million parameter model pre-trained on 91 million curated SMILES from PubChem, along with a Mixture-of-Experts variant (MoE-OSMI) that scales to 8x289M parameters. The models support molecular property prediction, molecule reconstruction, reaction yield prediction, and few-shot reasoning over molecular embeddings. All model weights and code are open-sourced under an Apache 2.0 license.
Bridging Encoding and Decoding for Molecular Representations
Chemical language models based on SMILES have gained traction for molecular property prediction and generation. Most existing models, such as MoLFormer and ChemBERTa, are encoder-only architectures that produce molecular embeddings through mean pooling. While effective for downstream classification and regression, this encoder-only approach has a limitation: mean pooling has no natural inverse, meaning the model cannot reconstruct the input molecule from its latent representation. This restricts the model’s utility for generative tasks and limits the interpretability of the learned latent space.
The authors argue that adding a decoder with a reconstruction objective forces the model to encode a more complete set of structural features. Prior work has shown that the quality of pre-training data matters more than the choice of SMILES vs. SELFIES, and that large-scale pre-training can yield useful chemical representations. SMI-TED builds on these observations by combining an encoder-decoder architecture with a carefully curated 91-million molecule dataset from PubChem.
Invertible Pooling and Two-Phase Pre-Training
The core architectural innovation in SMI-TED is a learned pooling mechanism that replaces standard mean or max pooling with an invertible projection. Given token embeddings $\mathbf{x} \in \mathbb{R}^{D \times L}$ (where $D = 202$ is the maximum token count and $L = 768$ is the embedding dimension), the submersion into the latent space $\mathbf{z} \in \mathbb{R}^{L}$ is computed as:
$$ \mathbf{z} = \left(\text{LayerNorm}\left(\text{GELU}\left(\mathbf{W}_1^T \mathbf{x} + \mathbf{b}_1\right)\right)\right) \mathbf{W}_2 $$
where $\mathbf{W}_1 \in \mathbb{R}^{D \times L}$, $\mathbf{b}_1 \in \mathbb{R}^{L}$, and $\mathbf{W}_2 \in \mathbb{R}^{L \times L}$. The immersion (inverse mapping) back to the token space is:
$$ \tilde{\mathbf{x}}^T = \left(\text{LayerNorm}\left(\text{GELU}\left(\mathbf{z} \mathbf{W}_3 + \mathbf{b}_3\right)\right)\right) \mathbf{W}_4 $$
where $\mathbf{W}_3 \in \mathbb{R}^{L \times L}$, $\mathbf{b}_3 \in \mathbb{R}^{L}$, and $\mathbf{W}_4 \in \mathbb{R}^{L \times D}$. A decoder language model then predicts the next token from $\tilde{\mathbf{x}}$.
The encoder uses a modified RoFormer attention mechanism with rotary position embeddings:
$$ \text{Attention}_m(Q, K, V) = \frac{\sum_{n=1}^{N} \langle \varphi(R_m q_m), \varphi(R_n k_n) \rangle v_n}{\sum_{n=1}^{N} \langle \varphi(R_m q_m), \varphi(R_n k_n) \rangle} $$
where $R_m$ are position-dependent rotation matrices and $\varphi$ is a random feature map.
Two-phase pre-training strategy:
- Phase 1: The token encoder is pre-trained on 95% of the data using masked language modeling (15% token selection, of which 80% masked, 10% random, 10% unchanged). The remaining 5% trains the encoder-decoder layer, preventing convergence issues from unstable early embeddings.
- Phase 2: After the token embeddings converge, both the encoder and decoder train on 100% of the data jointly.
Mixture-of-Experts (MoE-OSMI): The MoE variant composes 8 fine-tuned SMI-TED289M expert models with a gating network. Given an input embedding $x$, the output is:
$$ y = \sum_{i=1}^{n} G(x)_i E_i(\hat{x}) $$
where $G(x) = \text{Softmax}(\text{TopK}(x \cdot W_g))$ selects the top $k = 2$ experts per input, setting all other gate values to zero.
Benchmarks Across Property Prediction, Generation, and Reaction Yield
MoleculeNet classification (6 datasets, ROC-AUC)
| Method | BBBP | ClinTox | HIV | BACE | SIDER | Tox21 |
|---|---|---|---|---|---|---|
| MoLFormer | 73.6 +/- 0.8 | 91.2 +/- 1.4 | 80.5 +/- 1.65 | 86.3 +/- 0.6 | 65.5 +/- 0.2 | 80.46 +/- 0.2 |
| Uni-Mol | 72.9 +/- 0.6 | 91.9 +/- 1.8 | 80.8 +/- 0.3 | 85.7 +/- 0.2 | 65.9 +/- 1.3 | 79.6 +/- 0.5 |
| GEM | 72.4 +/- 0.4 | 90.1 +/- 1.3 | 80.6 +/- 0.9 | 85.6 +/- 1.1 | 67.2 +/- 0.4 | 78.1 +/- 0.1 |
| SMI-TED289M (pre-trained) | 91.46 +/- 0.47 | 93.49 +/- 0.85 | 80.51 +/- 1.34 | 85.58 +/- 0.92 | 66.01 +/- 0.88 | 81.53 +/- 0.45 |
| SMI-TED289M (fine-tuned) | 92.26 +/- 0.57 | 94.27 +/- 1.83 | 76.85 +/- 0.89 | 88.24 +/- 0.50 | 65.68 +/- 0.45 | 81.85 +/- 1.42 |
SMI-TED achieves the best results in 4 of 6 classification tasks. Notably, the pre-trained version (without fine-tuning) already matches or exceeds many baselines on BBBP, ClinTox, and Tox21.
MoleculeNet regression (5 datasets, MAE for QM9/QM8, RMSE for ESOL/FreeSolv/Lipophilicity)
| Method | QM9 | QM8 | ESOL | FreeSolv | Lipophilicity |
|---|---|---|---|---|---|
| MoLFormer | 1.5894 | 0.0102 | 0.880 | 2.342 | 0.700 |
| D-MPNN | 3.241 | 0.0143 | 0.98 | 2.18 | 0.65 |
| SMI-TED289M (fine-tuned) | 1.3246 | 0.0095 | 0.6112 | 1.2233 | 0.5522 |
SMI-TED289M achieves the best results across all 5 regression tasks when fine-tuned. The improvements are substantial on ESOL (0.61 vs. 0.82 for next best) and FreeSolv (1.22 vs. 1.91 for next best).
Reaction yield prediction (Buchwald-Hartwig C-N cross-coupling)
The model was tested on Pd-catalyzed Buchwald-Hartwig reactions with 3,955 reactions across varying train/test splits. Selected $R^2$ results:
| Split | Yield-BERT (Aug) | DRFP | SMI-TED289M |
|---|---|---|---|
| 70/30 | 0.97 | 0.95 | 0.984 |
| 10/90 | 0.81 | 0.81 | 0.961 |
| 2.5/97.5 | 0.61 | 0.62 | 0.875 |
| Test 1-4 avg | 0.58 | 0.71 | 0.983 |
SMI-TED shows particularly strong performance in low-data regimes. With only 2.5% training data, it achieves $R^2 = 0.875$, compared to 0.61-0.62 for competing methods.
MOSES molecular generation benchmarks
SMI-TED is competitive with baselines including CharRNN, SMILES VAE, JT-VAE, LIMO, MolGen-7b, and GP-MoLFormer on standard metrics (validity, uniqueness, novelty, FCD, internal diversity). It achieves superior scaffold cosine similarity (Scaf) and nearest-neighbor similarity (SNN) scores.
Latent space compositionality
Using six families of carbon chains ($\mathcal{F} = {CC, CO, CN, CS, CF, CP}$), the authors test whether the embedding space respects hierarchical distance structures. A linear regression on SMI-TED embeddings yields $R^2 = 0.99$ and $MSE = 0.002$, compared to $R^2 = 0.55$ and $MSE = 0.237$ for MoLFormer. This indicates that the SMI-TED latent space captures compositional chemical relationships far more faithfully.
For structure-property analysis on QM9, nitrogen-containing molecules represent 9.10% of the dataset but account for 32.81% of the top 10% by HOMO energy. In the SMI-TED latent space, these molecules cluster distinctly (Davies-Bouldin index of 2.82 vs. 4.28 for MoLFormer), suggesting the decoder objective encourages encoding of functional group information.
Strong Performance with a Compositional Latent Space
SMI-TED289M demonstrates competitive or superior performance across molecular property prediction, reaction yield prediction, and molecular generation benchmarks. The key findings include:
- Broad applicability: The single pre-trained model achieves strong results across classification (4/6 best), regression (5/5 best), reaction yield, and generation tasks.
- Low-data robustness: The pre-training on 91M molecules provides chemical knowledge that transfers well to small training sets, as shown by the reaction yield experiments where SMI-TED maintains high accuracy even at 2.5% training data.
- Compositional embeddings: The encoder-decoder architecture produces a latent space where molecular similarity follows chemical intuition, with near-perfect linear relationships between functional group families ($R^2 = 0.99$).
- Structure-property capture: The reconstruction objective appears to enforce encoding of chemically meaningful features like nitrogen substituent effects on HOMO energy, outperforming encoder-only models in latent space organization.
Limitations: The paper evaluates on MoleculeNet benchmarks, which are well-studied but may not reflect performance on more diverse chemical tasks. The BBBP classification result (92.26) shows a large jump from prior methods (73.6 for MoLFormer), which is worth scrutinizing. The MoE variant is evaluated only in supplementary materials, and scaling behavior beyond 8 experts is not explored.
Future directions: The authors note that compositionality of the learned representations suggests potential for reasoning applications, though they acknowledge that stronger claims require further studies following compositionality analysis methodologies from natural language processing. The model has been integrated into the dZiner agent for inverse molecular design.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Pre-training | PubChem (curated) | 91M molecules, 4B tokens | Deduplicated, canonicalized, validity-checked |
| Classification | MoleculeNet (BBBP, ClinTox, HIV, BACE, SIDER, Tox21) | Varies | Original benchmark splits |
| Regression | MoleculeNet (QM9, QM8, ESOL, FreeSolv, Lipophilicity) | Varies | Original benchmark splits |
| Generation | MOSES | 1.94M molecules | Train/test/scaffold test splits |
| Reaction yield | Buchwald-Hartwig HTE | 3,955 reactions | 3x 1536-well plates |
Algorithms
- Masked language modeling for token encoder (15% selection: 80% masked, 10% random, 10% unchanged)
- Two-phase pre-training (95/5 split then 100% joint training)
- RoFormer attention with rotary position embeddings
- Vocabulary: 2,993 tokens (2,988 molecular + 5 special)
- Maximum sequence length: 202 tokens (covers 99.4% of PubChem)
- Learning rate: 1.6e-4, batch size: 288 molecules
- 40 epochs over the full PubChem corpus
- 10 random seeds per experiment for robustness
Models
| Variant | Parameters | Encoder | Decoder | Description |
|---|---|---|---|---|
| SMI-TED289M base | 289M | 47M | 242M | 12 layers, 12 attention heads, hidden size 768, dropout 0.2 |
| MoE-OSMI | 8x289M | - | - | 8 experts, top-k=2 routing, gating network |
Evaluation
- Classification: ROC-AUC
- Regression: MAE (QM9, QM8), RMSE (ESOL, FreeSolv, Lipophilicity)
- Reaction yield: $R^2$
- Generation: Validity, uniqueness, novelty, FCD, IntDiv, Scaf, SNN (MOSES metrics)
- Latent space: Linear regression $R^2$, MSE, Davies-Bouldin index, t-SNE visualization
Hardware
- 24 NVIDIA V100 GPUs (16GB)
- 4 nodes with DDP (Distributed Data Parallel)
- Pre-training: 40 epochs on 91M molecules
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| IBM/materials (smi_ted) | Code | Apache-2.0 | Training, fine-tuning scripts, Jupyter notebooks |
| ibm/materials.smi-ted | Model | Apache-2.0 | Pre-trained model weights |
| Zenodo archive | Code + Data | Apache-2.0 | Archival copy of scripts |
Paper Information
Citation: Soares, E., Vital Brazil, E., Shirasuna, V., Zubarev, D., Cerqueira, R., & Schmidt, K. (2025). An open-source family of large encoder-decoder foundation models for chemistry. Communications Chemistry, 8(1). https://doi.org/10.1038/s42004-025-01585-0
@article{soares2025smited,
title={An open-source family of large encoder-decoder foundation models for chemistry},
author={Soares, Eduardo and Vital Brazil, Emilio and Shirasuna, Victor and Zubarev, Dmitry and Cerqueira, Renato and Schmidt, Kristin},
journal={Communications Chemistry},
volume={8},
number={1},
year={2025},
publisher={Nature Publishing Group},
doi={10.1038/s42004-025-01585-0}
}