An Unsupervised Seq2seq Method for Molecular Fingerprints
This is a Method paper that introduces seq2seq fingerprint, an unsupervised molecular embedding approach based on sequence-to-sequence learning. The core idea is to train a GRU encoder-decoder network to translate SMILES strings to themselves, then extract the intermediate fixed-length vector as a molecular fingerprint. These fingerprints are then used with standard supervised classifiers for downstream property prediction tasks such as solubility classification and promiscuity prediction.
The Labeled Data Bottleneck in Drug Discovery
Machine learning approaches to molecular property prediction depend on fixed-length feature vectors as inputs. Traditional molecular fingerprints fall into two categories: hash-based methods like Extended-Connectivity Fingerprints (ECFP) that are fast but lossy and non-invertible, and biologist-guided local-feature fingerprints that require domain expertise and are task-specific. Supervised deep learning fingerprints (e.g., neural fingerprints) can learn representations from data but require large amounts of labeled data, which is expensive to obtain in drug discovery due to the cost of biological experiments.
The authors identify three limitations of existing approaches:
- Hash-based fingerprints discard information during the hashing process and cannot reconstruct the original molecule
- Local-feature fingerprints require expert knowledge and generalize poorly across tasks
- Supervised deep learning fingerprints are data-hungry and fail when labeled data is limited
Self-Translation as Unsupervised Molecular Encoding
The key insight is to adapt the sequence-to-sequence learning framework from machine translation (originally English-to-French) to molecular representation learning by setting both the input and output to the same SMILES string. Since the intermediate vector must contain enough information to reconstruct the original SMILES, it serves as a rich, task-agnostic molecular fingerprint.
The architecture consists of two components:
- Perceiver network: A multi-layer GRU encoder that reads the SMILES string and compresses it into a fixed-length vector
- Interpreter network: A multi-layer GRU decoder that reconstructs the original SMILES from the fingerprint vector
The GRU cell computes a sequence of outputs $(s_1, \ldots, s_T)$ from input sequences $(x_1, \ldots, x_T)$ by iterating:
$$ z_t = \sigma_g(W_z x_t + U_z s_{t-1} + b_z) $$
$$ r_t = \sigma_r(W_r x_t + U_r s_{t-1} + b_r) $$
$$ h_t = \tanh(U_h x_t + W_h(s_{t-1} \circ r_t)) $$
$$ s_t = (1 - z_t) \circ h_{t-1} + z_t \circ s_{t-1} $$
where $z_t$ is the update gate, $r_t$ is the reset gate, $\circ$ denotes element-wise multiplication, and $W$, $U$, $b$ are trainable parameters.
Several adaptations to the original seq2seq framework make this work for molecular data:
- GRU instead of LSTM: GRU provides comparable performance with faster training, which is important given the large training data pool
- Attention mechanism: Establishes a stronger connection between the perceiver and interpreter networks via soft alignment, addressing the challenge of passing information through hidden memory for long sequences (SMILES can be up to 250 characters)
- Dropout layers: Added to input and output gates (but not hidden memory transfer) following the approach of Zaremba et al. to combat overfitting when training on large datasets
- Fingerprint extraction layer: A fixed-unit fully connected layer combined with a GRU cell state concatenation layer is inserted between encoder and decoder to explicitly output the fingerprint vector
- Reverse target sequence: Following Sutskever et al., the target sequence is reversed to improve SGD optimization
- Bucket training: Sequences are distributed into buckets by length and padded to enable GPU parallelization
Classification Experiments on LogP and PM2 Datasets
Training Setup
The unsupervised training used 334,092 valid SMILES representations from combined LogP and PM2-full datasets obtained from the National Center for Advancing Translational Sciences (NCATS) at NIH. Three model variants were trained with fingerprint dimensions of 512, 768, and 1024, differing in the number of GRU layers (2, 3, and 4 respectively) while keeping the latent dimension at 256. Each model was trained for 24 hours on a workstation with an Intel i7-6700K CPU, 16 GB RAM, and an NVIDIA GTX 1080 GPU.
Reconstruction Performance
The models were evaluated on their ability to reconstruct SMILES strings from their fingerprints:
| Model | GRU Layers | Latent Dim | Perplexity | Exact Match Accuracy |
|---|---|---|---|---|
| seq2seq-512 | 2 | 256 | 1.00897 | 94.24% |
| seq2seq-768 | 3 | 256 | 1.00949 | 92.92% |
| seq2seq-1024 | 4 | 256 | 1.01472 | 90.26% |
Deeper models showed lower reconstruction accuracy, possibly because larger fingerprint spaces introduce more null spaces and require longer training to converge.
Classification Results
Two labeled datasets were used for downstream classification:
- LogP: 10,850 samples with water-octanol partition coefficient values, binarized at a threshold of 1.88
- PM2-10k: 10,000 samples with binary promiscuity class labels
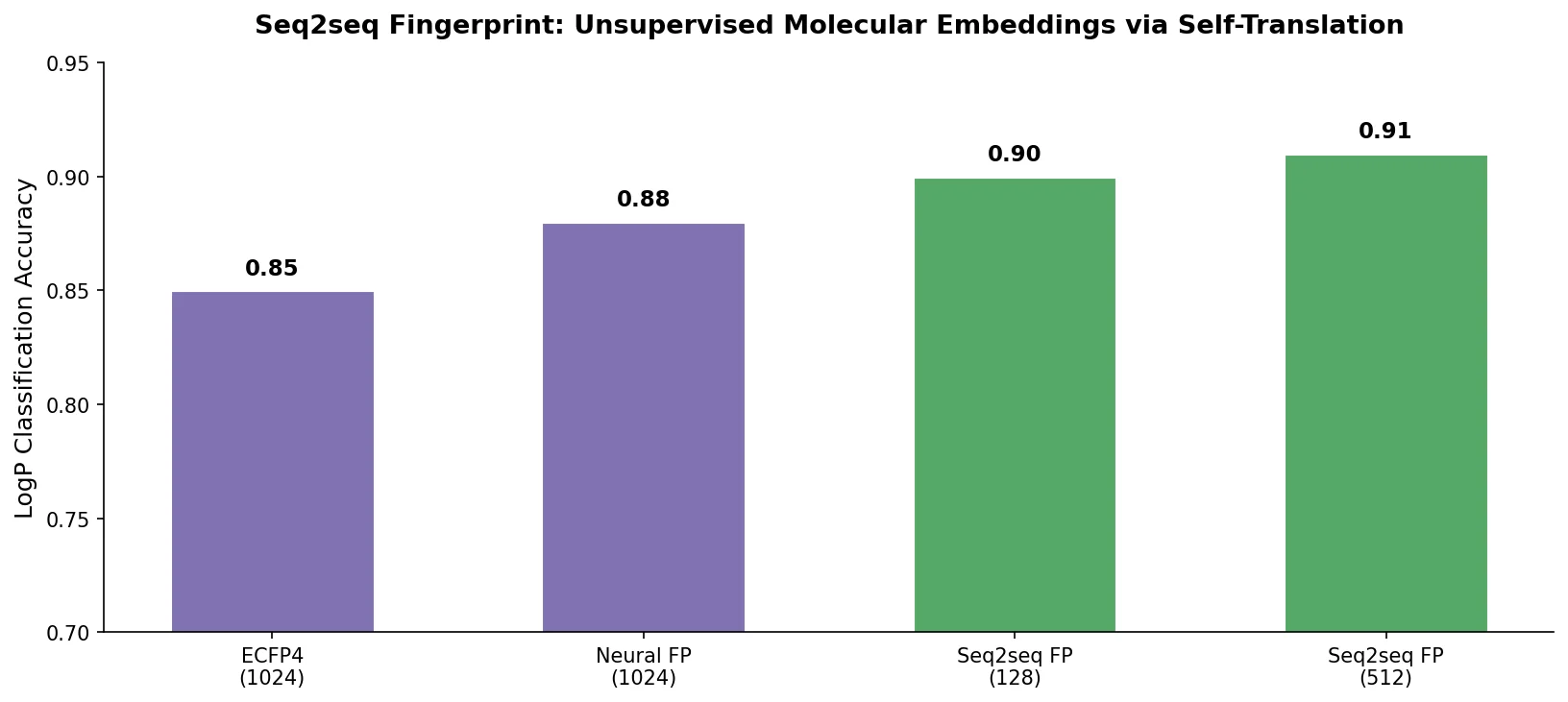
The seq2seq fingerprints were evaluated with three ensemble classifiers (AdaBoost, GradientBoost, RandomForest) against circular fingerprints (ECFP) and neural fingerprints. Results are 100-run averages of 5-fold cross-validation accuracy.
LogP classification accuracy:
| Method | Mean Accuracy | Std Dev |
|---|---|---|
| Circular FP (ECFP) | 0.3674 | 0.0074 |
| Neural FP | 0.6080 | 0.0135 |
| Seq2seq-1024 + GradientBoost | 0.7664 | 0.0043 |
| Seq2seq-1024 + AdaBoost | 0.7342 | 0.0042 |
| Seq2seq-512 + GradientBoost | 0.7350 | 0.0060 |
PM2-10k classification accuracy:
| Method | Mean Accuracy | Std Dev |
|---|---|---|
| Circular FP (ECFP) | 0.3938 | 0.0114 |
| Neural FP | 0.5227 | 0.0112 |
| Seq2seq-1024 + GradientBoost | 0.6206 | 0.0198 |
| Seq2seq-1024 + AdaBoost | 0.6036 | 0.0147 |
| Seq2seq-512 + GradientBoost | 0.5741 | 0.0086 |
The seq2seq fingerprint outperformed both baselines across all configurations. Despite the seq2seq-1024 model having lower reconstruction accuracy, it provided the best classification performance, suggesting that the longer fingerprint captures more discriminative information for downstream tasks even if the reconstruction is less exact.
Unsupervised Transfer Learning for Molecular Properties
The results demonstrate that unsupervised pretraining on large unlabeled molecular datasets can produce fingerprints that transfer well to supervised property prediction with limited labels. The key advantages confirmed by the experiments are:
- Label-free training: The unsupervised approach uses essentially unlimited SMILES data, avoiding the expensive label collection process
- Task-agnostic representations: The same fingerprints work across different classification tasks (solubility and promiscuity) without retraining
- Invertibility: The fingerprints contain enough information to reconstruct the original SMILES (up to 94.24% exact match), unlike hash-based methods
Limitations acknowledged by the authors include:
- Long training times (24 hours per model variant), motivating future work on distributed training
- The relationship between fingerprint dimensionality and downstream performance is non-monotonic (768-dim underperforms 512-dim on some tasks), suggesting sensitivity to hyperparameter choices
- Only classification tasks were evaluated; regression performance was not assessed
- The comparison baselines are limited to ECFP and neural fingerprints from 2015
Future directions proposed include distributed training strategies, hyperparameter optimization methods, and semi-supervised extensions that incorporate label information into the fingerprint training.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Unsupervised training | LogP + PM2-full (combined) | 334,092 SMILES | Obtained from NCATS at NIH |
| Classification | LogP | 10,850 samples | Binary labels at LogP threshold 1.88 |
| Classification | PM2-10k | 10,000 samples | Binary promiscuity labels |
Algorithms
- Encoder-decoder: Multi-layer GRU with attention mechanism and dropout
- Fingerprint dimensions: 512, 768, 1024 (with 2, 3, 4 GRU layers respectively)
- Latent dimension: 256 for all variants
- Downstream classifiers: AdaBoost, GradientBoost, RandomForest
- Evaluation: 5-fold cross-validation, 100-run averages
- Baselines: ECFP via RDKit, Neural Fingerprint from HIPS/neural-fingerprint
Models
Three model variants trained for 24 hours each. The paper states code would become publicly available after acceptance, but no public repository has been confirmed.
Evaluation
| Metric | Best Value | Task | Configuration |
|---|---|---|---|
| Classification accuracy | 0.7664 | LogP | seq2seq-1024 + GradientBoost |
| Classification accuracy | 0.6206 | PM2-10k | seq2seq-1024 + GradientBoost |
| Exact match reconstruction | 94.24% | SMILES recovery | seq2seq-512 |
| Perplexity | 1.00897 | SMILES recovery | seq2seq-512 |
Hardware
- Training: Intel i7-6700K @ 4.00 GHz, 16 GB RAM, NVIDIA GTX 1080 GPU
- Hyperparameter search and classifier training: TACC Lonestar 5 cluster
- Training time: 24 hours per model variant
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| Neural Fingerprint (baseline) | Code | MIT | Baseline comparison code |
The authors indicated the seq2seq fingerprint code would be released after acceptance, but no public repository has been found as of this writing. The datasets were sourced from NCATS/NIH.
Paper Information
Citation: Xu, Z., Wang, S., Zhu, F., & Huang, J. (2017). Seq2seq Fingerprint: An Unsupervised Deep Molecular Embedding for Drug Discovery. Proceedings of the 8th ACM International Conference on Bioinformatics, Computational Biology, and Health Informatics (ACM-BCB ‘17), 285-294. https://doi.org/10.1145/3107411.3107424
@inproceedings{xu2017seq2seq,
title={Seq2seq Fingerprint: An Unsupervised Deep Molecular Embedding for Drug Discovery},
author={Xu, Zheng and Wang, Sheng and Zhu, Feiyun and Huang, Junzhou},
booktitle={Proceedings of the 8th ACM International Conference on Bioinformatics, Computational Biology, and Health Informatics},
pages={285--294},
year={2017},
publisher={ACM},
doi={10.1145/3107411.3107424}
}