BERT-Based Molecular Representations with Auxiliary Pre-Training Tasks
This is a Method paper that introduces MolBERT, a bidirectional Transformer (BERT) architecture applied to SMILES-based molecular representations for drug discovery. The primary contribution is a systematic study of how different domain-relevant self-supervised pre-training tasks affect the quality of learned molecular embeddings, paired with a model that achieves state-of-the-art performance on virtual screening and quantitative structure-activity relationship (QSAR) benchmarks.
Why Domain-Relevant Pre-Training Matters for Molecular Language Models
Molecular representations are foundational for predictive, generative, and analytical tasks in drug discovery. Language models applied to text-based molecular representations like SMILES have demonstrated strong performance across property prediction, reaction prediction, and molecular generation. However, several open questions remained at the time of this work:
- Task selection for pre-training: Prior work explored masked token prediction, input translation, and property concatenation, but there was no systematic comparison of how different self-supervised tasks affect downstream performance.
- SMILES ambiguity: The same molecule can be encoded as many different SMILES strings depending on how the molecular graph is traversed. Canonicalization algorithms address this but introduce their own artifacts that may distract the model.
- Domain knowledge integration: Standard NLP pre-training objectives (e.g., masked language modeling) do not explicitly encode chemical knowledge. It was unclear whether incorporating chemistry-specific supervision during pre-training could improve representation quality.
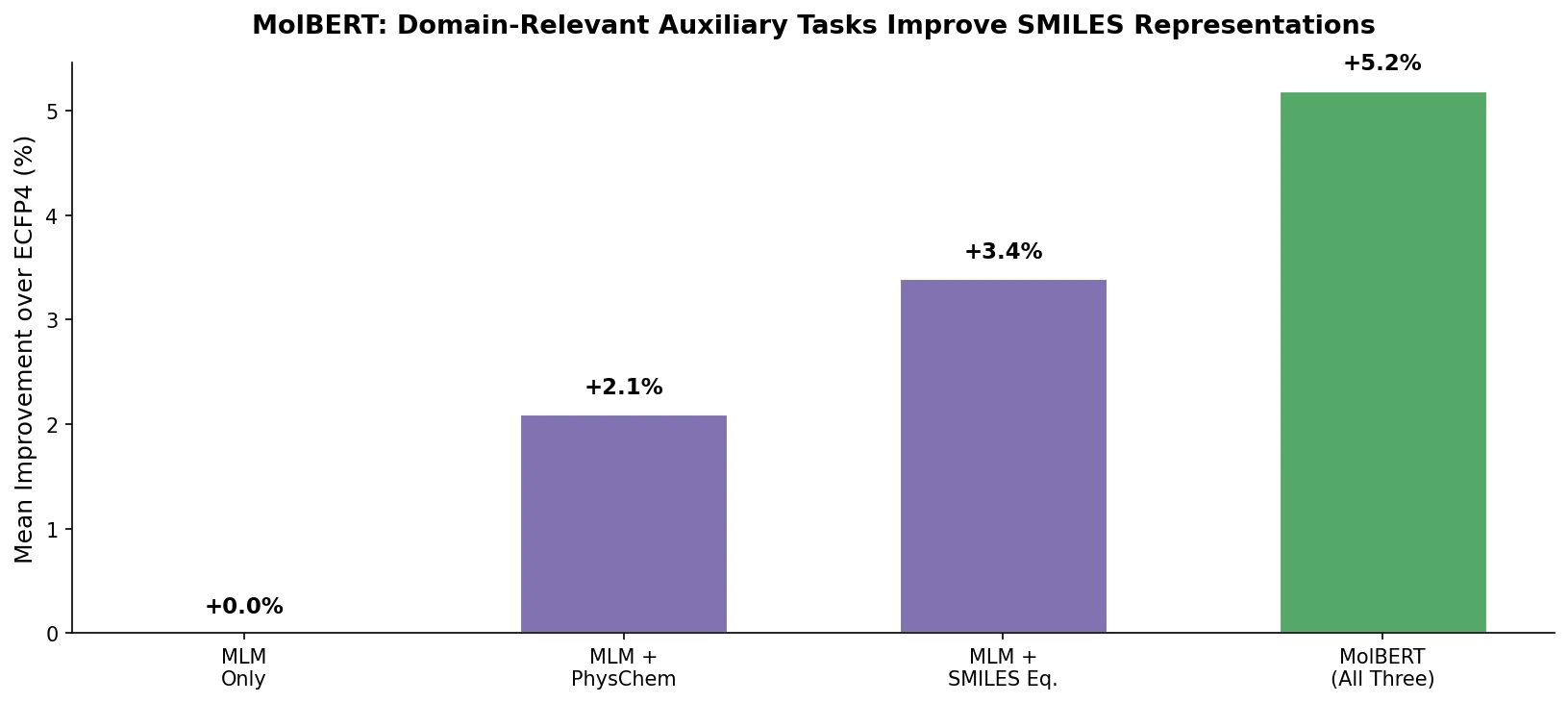
MolBERT addresses these gaps by evaluating three pre-training tasks, including a novel physicochemical property prediction objective, and measuring their individual and combined effects on downstream drug discovery benchmarks.
Three Auxiliary Tasks for Chemistry-Aware Pre-Training
MolBERT uses the BERT-Base architecture (12 attention heads, 12 layers, 768-dimensional hidden states, approximately 85M parameters) and explores three self-supervised pre-training tasks:
Masked Language Modeling (MaskedLM): The standard BERT objective where 15% of input tokens are masked and the model predicts their identity. The loss is cross-entropy between predicted and true tokens.
SMILES Equivalence (SMILES-Eq): A binary classification task where the model receives two SMILES strings and predicts whether they represent the same molecule. The second string is either a random permutation of the first (same molecule, different traversal) or a randomly sampled molecule. This is optimized with cross-entropy loss.
Physicochemical Property Prediction (PhysChemPred): Using RDKit, a set of 200 real-valued molecular descriptors are computed for each molecule. The model predicts these normalized descriptors from the SMILES input using mean squared error:
$$\mathcal{L}_{\text{PhysChemPred}} = \frac{1}{D} \sum_{d=1}^{D} (y_d - \hat{y}_d)^2$$
where $D = 200$ is the number of descriptors, $y_d$ is the true normalized descriptor value, and $\hat{y}_d$ is the model’s prediction.
The final training loss is the arithmetic mean of all active task losses:
$$\mathcal{L}_{\text{total}} = \frac{1}{|\mathcal{T}|} \sum_{t \in \mathcal{T}} \mathcal{L}_t$$
where $\mathcal{T}$ is the set of active pre-training tasks.
Additionally, MolBERT supports SMILES permutation augmentation during training, where each input molecule is represented by a randomly sampled non-canonical SMILES string rather than the canonical form. The model uses a fixed vocabulary of 42 tokens, a sequence length of 128, and relative positional embeddings (from Transformer-XL) to support arbitrary-length SMILES at inference time.
Ablation Study and Benchmark Evaluation
Pre-Training Setup
All models were pre-trained on the GuacaMol benchmark dataset, consisting of approximately 1.6M compounds curated from ChEMBL, using an 80%/5% train/validation split. Training used the Adam optimizer with a learning rate of $3 \times 10^{-5}$ for 20 epochs (ablation) or 100 epochs (final model).
Ablation: Impact of Task Combinations on Virtual Screening
The ablation study evaluated all seven possible task combinations on the RDKit virtual screening benchmark (69 datasets, 5 query molecules per target). Results measured by AUROC and BEDROC20 (an early enrichment metric with $\alpha = 20$):
| MaskedLM | PhysChemPred | SMILES-Eq | AUROC (w/ perm) | BEDROC20 (w/ perm) | AUROC (w/o perm) | BEDROC20 (w/o perm) |
|---|---|---|---|---|---|---|
| Yes | Yes | Yes | 0.685 +/- 0.069 | 0.246 +/- 0.041 | 0.707 +/- 0.059 | 0.280 +/- 0.042 |
| Yes | Yes | No | 0.738 +/- 0.060 | 0.323 +/- 0.071 | 0.740 +/- 0.066 | 0.322 +/- 0.065 |
| Yes | No | Yes | 0.483 +/- 0.092 | 0.092 +/- 0.069 | 0.493 +/- 0.068 | 0.108 +/- 0.070 |
| No | Yes | Yes | 0.476 +/- 0.077 | 0.064 +/- 0.034 | 0.514 +/- 0.165 | 0.084 +/- 0.014 |
| Yes | No | No | 0.696 +/- 0.058 | 0.283 +/- 0.077 | 0.676 +/- 0.060 | 0.250 +/- 0.073 |
| No | Yes | No | 0.719 +/- 0.057 | 0.293 +/- 0.071 | 0.716 +/- 0.061 | 0.290 +/- 0.076 |
| No | No | Yes | 0.129 +/- 0.067 | 0.005 +/- 0.037 | 0.508 +/- 0.068 | 0.048 +/- 0.035 |
Key findings from the ablation:
- PhysChemPred had the highest individual impact (average BEDROC20 of 0.292 alone vs. 0.266 for MaskedLM alone).
- Combining MaskedLM + PhysChemPred achieved the best performance (BEDROC20 of 0.323), though the additive gain from MaskedLM was modest (+0.031).
- The SMILES-Eq task consistently decreased performance when added to other task combinations.
A further sub-ablation on PhysChemPred descriptor groups showed that surface descriptors alone (49 of 200 descriptors) achieved nearly the same performance as the full set, suggesting molecular surface properties provide particularly informative supervision.
Virtual Screening Results
Using the best task combination (MaskedLM + PhysChemPred) trained for 100 epochs:
| Method | AUROC | BEDROC20 |
|---|---|---|
| MolBERT (100 epochs) | 0.743 +/- 0.062 | 0.344 +/- 0.062 |
| CDDD | 0.725 +/- 0.057 | 0.310 +/- 0.080 |
| RDKit descriptors | 0.633 +/- 0.027 | 0.217 +/- 0.000 |
| ECFC4 | 0.603 +/- 0.056 | 0.170 +/- 0.079 |
MolBERT outperformed all baselines including CDDD (the prior state of the art), RDKit calculated descriptors, and extended-connectivity fingerprints (ECFC4).
QSAR Results
On MoleculeNet regression tasks (RMSE, lower is better):
| Dataset | RDKit (norm) | ECFC4 | CDDD | MolBERT | MolBERT (finetune) |
|---|---|---|---|---|---|
| ESOL | 0.687 +/- 0.08 | 0.902 +/- 0.06 | 0.567 +/- 0.06 | 0.552 +/- 0.07 | 0.531 +/- 0.04 |
| FreeSolv | 1.671 +/- 0.45 | 2.876 +/- 0.38 | 1.456 +/- 0.43 | 1.523 +/- 0.66 | 0.948 +/- 0.33 |
| Lipophilicity | 0.738 +/- 0.04 | 0.770 +/- 0.03 | 0.669 +/- 0.02 | 0.602 +/- 0.01 | 0.561 +/- 0.03 |
On MoleculeNet classification tasks (AUROC, higher is better):
| Dataset | RDKit (norm) | ECFC4 | CDDD | MolBERT | MolBERT (finetune) |
|---|---|---|---|---|---|
| BACE | 0.831 | 0.845 | 0.833 | 0.849 | 0.866 |
| BBBP | 0.696 | 0.678 | 0.761 | 0.750 | 0.762 |
| HIV | 0.708 | 0.714 | 0.753 | 0.747 | 0.783 |
Fine-tuned MolBERT achieved the best performance on all six QSAR datasets. When used as a fixed feature extractor with an SVM, MolBERT embeddings outperformed other representations on three of six tasks.
Key Findings and Limitations
Key Findings
- Pre-training task selection matters significantly. The choice of auxiliary tasks during pre-training has a large effect on downstream performance. PhysChemPred provides the strongest individual signal.
- Domain-relevant auxiliary tasks improve representation quality. Predicting physicochemical properties during pre-training encodes chemical knowledge directly into the embeddings, outperforming purely linguistic objectives.
- The SMILES equivalence task hurts performance. Despite being chemically motivated, the SMILES-Eq task consistently degraded results, suggesting it may introduce conflicting learning signals.
- PhysChemPred organizes the embedding space. Analysis of pairwise cosine similarities showed that models trained with PhysChemPred assign high similarity to permutations of the same molecule and low similarity to different molecules, creating a more semantically meaningful representation space.
Limitations
- The paper evaluates only SMILES-based representations, inheriting all limitations of string-based molecular encodings (inability to capture 3D structure, sensitivity to tokenization).
- The virtual screening evaluation uses a fixed number of query molecules ($n = 5$), which may not reflect realistic screening scenarios.
- Cross-validation splits from ChemBench were used for QSAR evaluation rather than scaffold splits, which may overestimate performance on structurally novel compounds.
- The model’s 128-token sequence length limit may truncate larger molecules, though relative positional embeddings partially address this at inference time.
Future Directions
The authors propose extending MolBERT to learn representations for other biological entities such as proteins, and developing more advanced pre-training strategies.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Pre-training | GuacaMol (ChEMBL) | ~1.6M compounds | 80% train / 5% validation split |
| Virtual Screening | RDKit benchmark v1.2 | 69 target datasets | Filtered subset with active/decoy compounds |
| QSAR (Regression) | ESOL, FreeSolv, Lipophilicity | Varies | From MoleculeNet, ChemBench splits |
| QSAR (Classification) | BACE, BBBP, HIV | Varies | From MoleculeNet, ChemBench splits |
Algorithms
- Architecture: BERT-Base (12 heads, 12 layers, 768-dim hidden, ~85M params)
- Optimizer: Adam, learning rate $3 \times 10^{-5}$
- Vocabulary: 42 tokens, sequence length 128
- Masking: 15% of tokenized input
- Positional encoding: relative positional embeddings (Transformer-XL)
- Fine-tuning SVM: $C = 5.0$, RBF kernel (from Winter et al.)
- Fine-tuning head: single linear layer on pooled output
- Embeddings: pooled output (or average sequence output when only MaskedLM is used)
Models
- BERT-Base with ~85M parameters
- Pre-trained weights available at BenevolentAI/MolBERT
Evaluation
| Metric | Task | Notes |
|---|---|---|
| AUROC | Virtual Screening, Classification QSAR | Standard area under ROC curve |
| BEDROC20 | Virtual Screening | Early enrichment metric, $\alpha = 20$ |
| RMSE | Regression QSAR | Root mean squared error |
Hardware
- 2 GPUs, 16 CPUs
- Pre-training time: ~40 hours (20 epochs)
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| BenevolentAI/MolBERT | Code + Model | MIT | Official implementation with pre-trained weights |
Paper Information
Citation: Fabian, B., Edlich, T., Gaspar, H., Segler, M., Meyers, J., Fiscato, M., & Ahmed, M. (2020). Molecular representation learning with language models and domain-relevant auxiliary tasks. arXiv preprint arXiv:2011.13230.
@article{fabian2020molecular,
title={Molecular representation learning with language models and domain-relevant auxiliary tasks},
author={Fabian, Benedek and Edlich, Thomas and Gaspar, H{\'e}l{\'e}na and Segler, Marwin and Meyers, Joshua and Fiscato, Marco and Ahmed, Mohamed},
journal={arXiv preprint arXiv:2011.13230},
year={2020}
}