Word2vec Meets Cheminformatics
Mol2vec is a Method paper that introduces an unsupervised approach for learning dense vector representations of molecular substructures. The core idea is a direct analogy to Word2vec from natural language processing: molecular substructures (derived from the Morgan algorithm) are treated as “words,” and entire molecules are treated as “sentences.” By training on a large unlabeled corpus of 19.9 million compounds, Mol2vec produces embeddings where chemically related substructures occupy nearby regions of vector space. Compound-level vectors are then obtained by summing constituent substructure vectors, and these can serve as features for downstream supervised learning tasks.
Sparse Fingerprints and Their Limitations
Molecular fingerprints, particularly Morgan fingerprints (extended-connectivity fingerprints, ECFP), are among the most widely used molecular representations in cheminformatics. They perform well for similarity searching, virtual screening, and activity prediction. However, they suffer from several practical drawbacks:
- High dimensionality and sparsity: Morgan fingerprints are typically hashed to fixed-length binary vectors (e.g., 2048 or 4096 bits), resulting in very sparse representations.
- Bit collisions: The hashing step can map distinct substructures to the same bit position, losing structural information.
- No learned relationships: Each bit is independent, so the representation does not encode any notion of chemical similarity between substructures.
At the time of this work (2017), NLP techniques had started to appear in cheminformatics. The tf-idf method had been applied to Morgan fingerprints for compound-protein interaction prediction, and Latent Dirichlet Allocation had been used for chemical topic modeling. The Word2vec concept had been adapted for protein sequences (ProtVec) but had not yet been applied to small molecules. Mol2vec fills this gap.
From Substructure Identifiers to Dense Embeddings
The central insight of Mol2vec is that the Morgan algorithm already produces a natural “vocabulary” of molecular substructures, and the order in which these substructures appear in a molecule provides local context, analogous to word order in a sentence.
Corpus Construction
The training corpus was assembled from ZINC v15 and ChEMBL v23, merged and deduplicated, then filtered by molecular weight (12-600), heavy atom count (3-50), clogP (-5 to 7), and allowed elements (H, B, C, N, O, F, P, S, Cl, Br). This yielded 19.9 million compounds.
Sentence Generation
For each molecule, the Morgan algorithm generates atom identifiers at radius 0 and radius 1. Each atom contributes two identifiers (one per radius), ordered according to the atom order in the canonical SMILES. This sequence of identifiers forms a “sentence” for Word2vec training.
Word2vec Training
The model was trained using the gensim implementation of Word2vec. After evaluating both CBOW and Skip-gram architectures with window sizes of 5, 10, and 20, and embedding dimensions of 100 and 300, the best configuration was:
- Architecture: Skip-gram
- Window size: 10
- Embedding dimension: 300
Rare identifiers appearing fewer than 3 times in the corpus were replaced with a special “UNSEEN” token, which learns a near-zero vector. This allows the model to handle novel substructures at inference time.
Compound Vector Generation
The final vector for a molecule is the sum of all its substructure vectors:
$$\mathbf{v}_{\text{mol}} = \sum_{i=1}^{N} \mathbf{v}_{s_i}$$
where $\mathbf{v}_{s_i}$ is the 300-dimensional embedding for the $i$-th substructure identifier in the molecule. This summation implicitly captures substructure counts and importance through vector amplitude.
Benchmarking Across Regression and Classification Tasks
Datasets
The authors evaluated Mol2vec on four datasets:
| Dataset | Task | Size | Description |
|---|---|---|---|
| ESOL | Regression | 1,144 | Aqueous solubility prediction |
| Ames | Classification | 6,511 | Mutagenicity (balanced: 3,481 positive, 2,990 negative) |
| Tox21 | Classification | 8,192 | 12 human toxicity targets (imbalanced) |
| Kinase | Classification | 284 kinases | Bioactivity from ChEMBL v23 |
Machine Learning Methods
Three ML methods were compared using both Mol2vec and Morgan FP features:
- Random Forest (RF): scikit-learn, 500 estimators
- Gradient Boosting Machine (GBM): XGBoost, 2000 estimators, max depth 3, learning rate 0.1
- Deep Neural Network (DNN): Keras/TensorFlow, 4 hidden layers with 2000 neurons each for Mol2vec; 1 hidden layer with 512 neurons for Morgan FP
All models were validated using 20x 5-fold cross-validation with the Wilcoxon signed-rank test for statistical comparison.
ESOL Regression Results
| Features | Method | $R^2_{\text{ext}}$ | MSE | MAE |
|---|---|---|---|---|
| Descriptors | MLR | 0.81 +/- 0.01 | 0.82 | 0.69 |
| Molecular Graph | CNN | 0.93 | 0.31 +/- 0.03 | 0.40 +/- 0.00 |
| Morgan FP | GBM | 0.66 +/- 0.00 | 1.43 +/- 0.00 | 0.88 +/- 0.00 |
| Mol2vec | GBM | 0.86 +/- 0.00 | 0.62 +/- 0.00 | 0.60 +/- 0.00 |
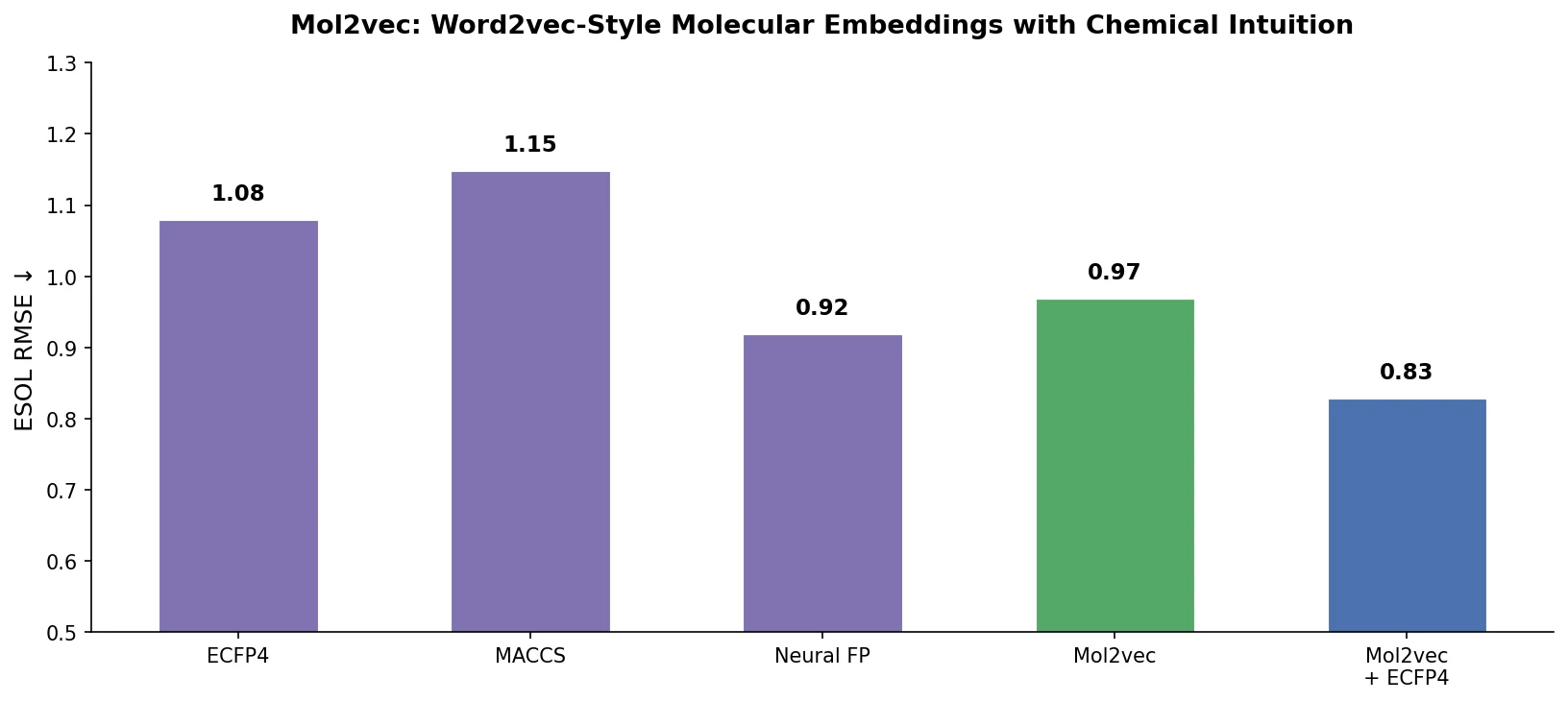
Mol2vec substantially outperformed Morgan FP ($R^2_{\text{ext}}$ 0.86 vs. 0.66) but did not match the best graph convolution methods ($R^2_{\text{ext}}$ ~0.93).
Classification Results (Ames and Tox21)
On the Ames dataset, Mol2vec and Morgan FP performed comparably (AUC 0.87 vs. 0.88), both matching or exceeding prior SVM and Naive Bayes results. On Tox21, both achieved an average AUC of 0.83, outperforming literature results from graph convolution (0.71) and DNN/SVM approaches (0.71-0.72).
Proteochemometric (PCM) Extension
Mol2vec was combined with ProtVec (protein sequence embeddings using the same Word2vec approach on 3-grams) by concatenating vectors, forming PCM2vec. This was evaluated using a rigorous 4-level cross-validation scheme:
- CV1: New compound-target pairs
- CV2: New targets
- CV3: New compounds
- CV4: New compounds and targets
On Tox21, PCM2vec improved predictions for new compound-target pairs (CV1: AUC 0.87 vs. 0.79 for Morgan FP) and new compounds (CV3: AUC 0.85 vs. 0.78). On the kinase dataset, PCM2vec approached the performance of classical PCM (Morgan + z-scales) while being alignment-independent, meaning it can be applied to proteins with low sequence similarity.
Chemical Intuition and Practical Value
Embedding Quality
The learned substructure embeddings capture meaningful chemical relationships. Hierarchical clustering of the 25 most common substructures shows expected groupings: aromatic carbons cluster together, aliphatic ring carbons form a separate group, and carbonyl carbons and oxygens are closely related. Similarly, t-SNE projections of amino acid vectors encoded by Mol2vec reproduce known amino acid relationships (e.g., similar distances between Glu/Gln and Asp/Asn pairs, reflecting the carboxylic acid to amide transition).
Key Findings
- Skip-gram with 300-dimensional embeddings provides the best Mol2vec representations, consistent with NLP best practices.
- Mol2vec excels at regression tasks, substantially outperforming Morgan FP on ESOL solubility prediction ($R^2_{\text{ext}}$ 0.86 vs. 0.66).
- Classification performance is competitive with Morgan FP across Ames and Tox21 datasets.
- PCM2vec enables alignment-independent proteochemometrics, extending PCM approaches to diverse protein families with low sequence similarity.
- Tree-based methods (RF, GBM) outperformed DNNs on these tasks, though the authors note further DNN tuning could help.
Limitations
- The compound vector is a simple sum of substructure vectors, which discards information about substructure arrangement and molecular topology.
- Only Morgan identifiers at radii 0 and 1 were used. Larger radii might capture more context but would increase vocabulary size.
- DNN architectures were not extensively optimized, leaving open the question of how well Mol2vec pairs with deep learning.
- The approach was benchmarked against Morgan FP but not against other learned representations such as graph neural networks in a controlled comparison.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Pre-training | ZINC v15 + ChEMBL v23 | 19.9M compounds | Filtered by MW, atom count, clogP, element types |
| Evaluation | ESOL | 1,144 compounds | Aqueous solubility regression |
| Evaluation | Ames | 6,511 compounds | Mutagenicity classification |
| Evaluation | Tox21 | 8,192 compounds | 12 toxicity targets, retrieved via DeepChem |
| Evaluation | Kinase (ChEMBL v23) | 284 kinases | IC50/Kd/Ki binding assays |
| Protein corpus | UniProt | 554,241 sequences | For ProtVec training |
Algorithms
- Word2vec: Skip-gram, window size 10, 300-dimensional embeddings, min count 3
- Morgan algorithm: Radii 0 and 1 (119 and 19,831 unique identifiers respectively)
- UNSEEN token: Replaces identifiers occurring fewer than 3 times
- Compound vector: Sum of all substructure vectors
Models
- RF: scikit-learn, 500 estimators, sqrt features, balanced class weights
- GBM: XGBoost, 2000 estimators, max depth 3, learning rate 0.1
- DNN: Keras/TensorFlow, 4 layers x 2000 neurons (Mol2vec) or 1 layer x 512 neurons (Morgan FP), ReLU activation, dropout 0.1
Evaluation
| Metric | Mol2vec Best | Morgan FP Best | Task |
|---|---|---|---|
| $R^2_{\text{ext}}$ | 0.86 (GBM) | 0.66 (GBM) | ESOL regression |
| AUC | 0.87 (RF) | 0.88 (RF) | Ames classification |
| AUC | 0.83 (RF) | 0.83 (RF) | Tox21 classification |
Hardware
Not specified in the paper.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| mol2vec | Code | BSD-3-Clause | Python package with pre-trained model |
Paper Information
Citation: Jaeger, S., Fulle, S., & Turk, S. (2018). Mol2vec: Unsupervised Machine Learning Approach with Chemical Intuition. Journal of Chemical Information and Modeling, 58(1), 27-35. https://doi.org/10.1021/acs.jcim.7b00616
@article{jaeger2018mol2vec,
title={Mol2vec: Unsupervised Machine Learning Approach with Chemical Intuition},
author={Jaeger, Sabrina and Fulle, Simone and Turk, Samo},
journal={Journal of Chemical Information and Modeling},
volume={58},
number={1},
pages={27--35},
year={2018},
publisher={American Chemical Society},
doi={10.1021/acs.jcim.7b00616}
}