This group covers models designed primarily to learn fixed-dimensional molecular representations from chemical string notations (SMILES, SELFIES, InChI). These encoders serve as feature extractors for downstream tasks like property prediction, virtual screening, and molecular similarity search. For models that fuse molecular strings or graphs with additional modalities (text, property vectors, knowledge graphs), see Multimodal Molecular Models.
| Paper | Year | Architecture | Key Idea |
|---|---|---|---|
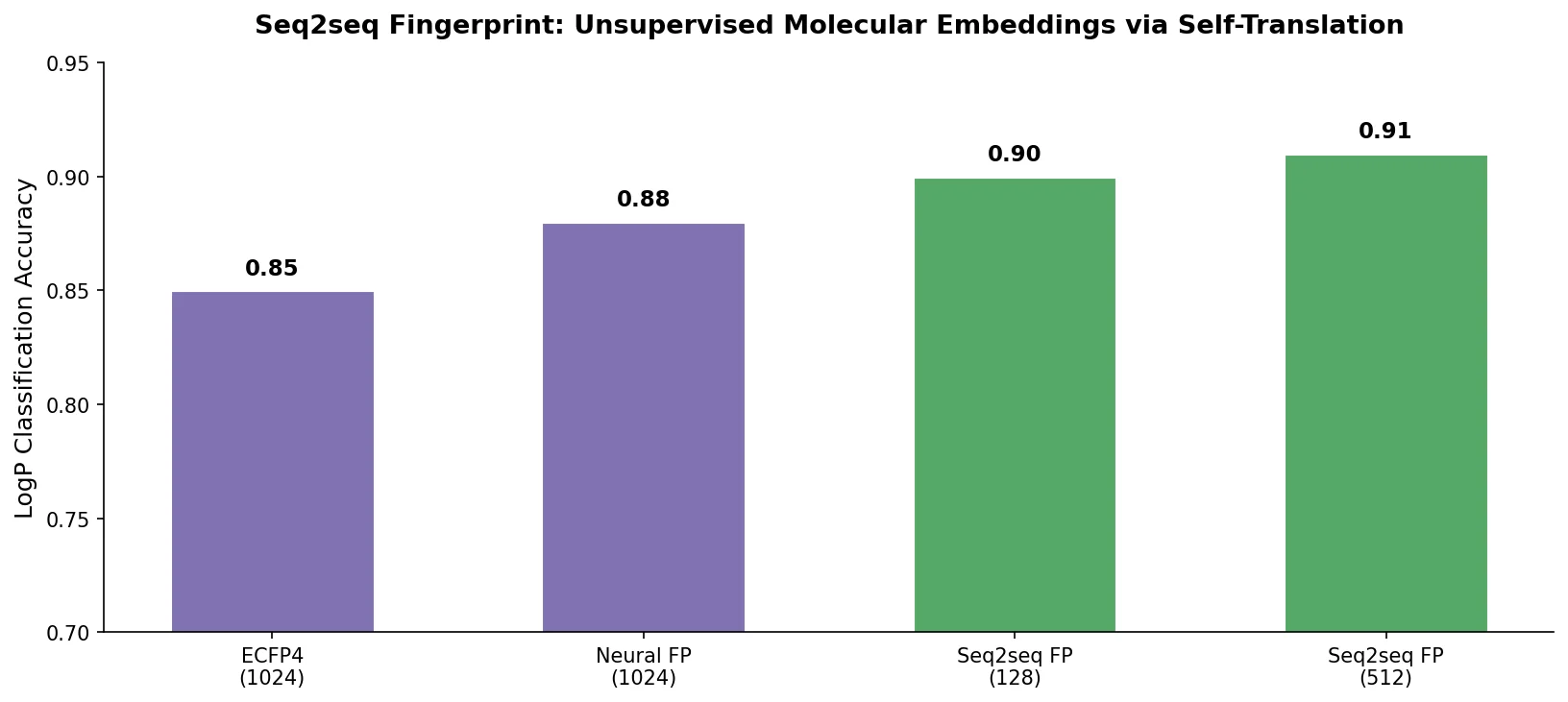
| Seq2seq Fingerprint | 2017 | GRU encoder-decoder | Unsupervised fingerprints learned from SMILES translation |
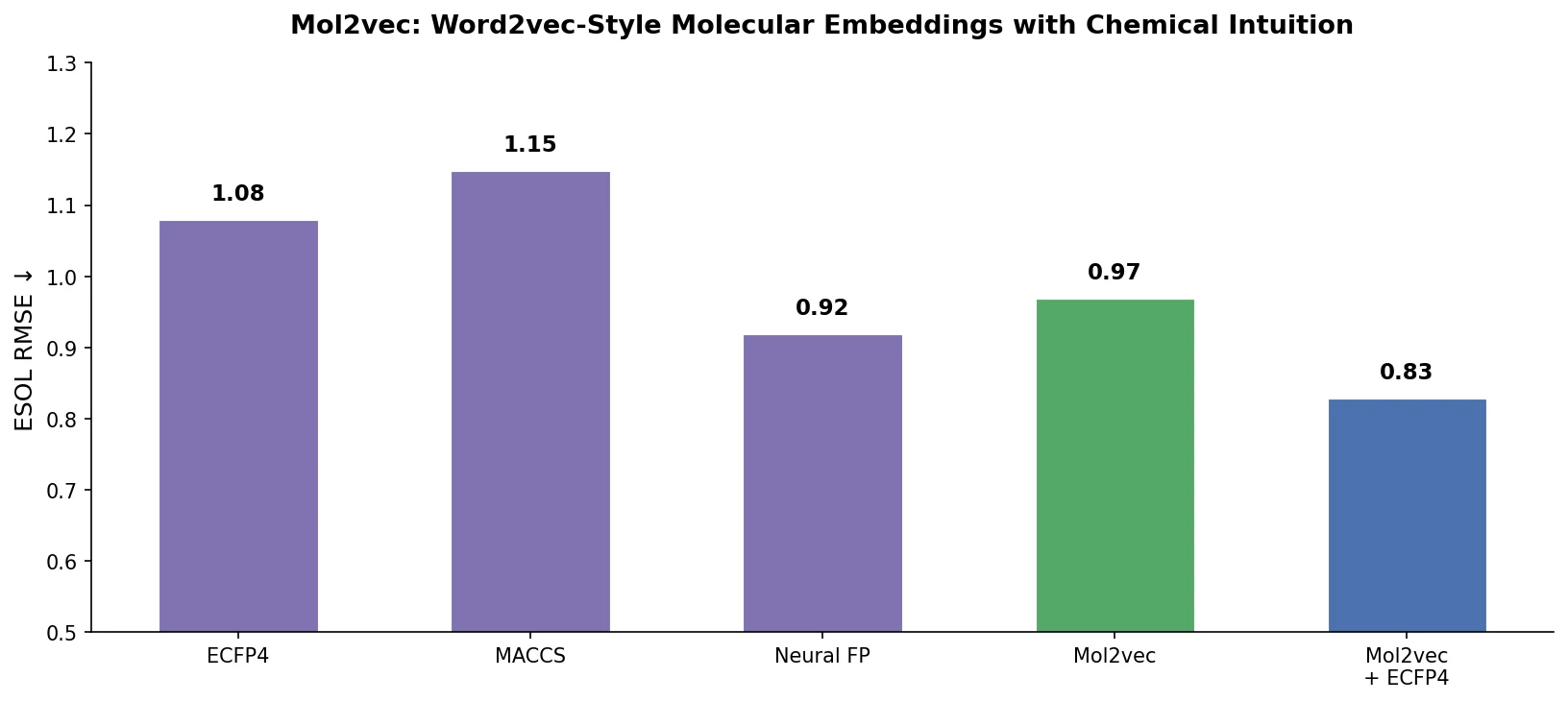
| Mol2vec | 2018 | Word2vec | Substructure embeddings inspired by NLP word vectors |
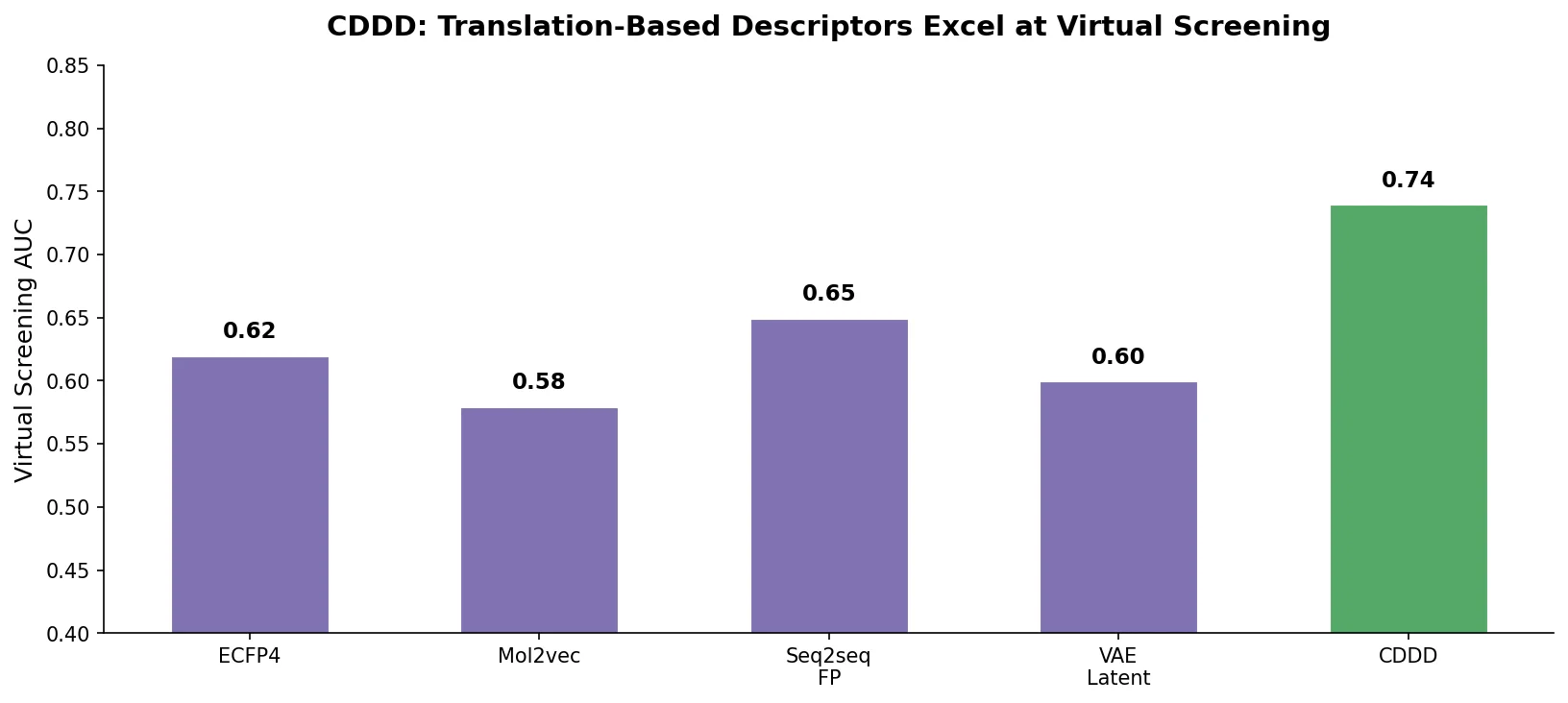
| CDDD | 2019 | Encoder-decoder | Descriptors learned by translating between SMILES and InChI |
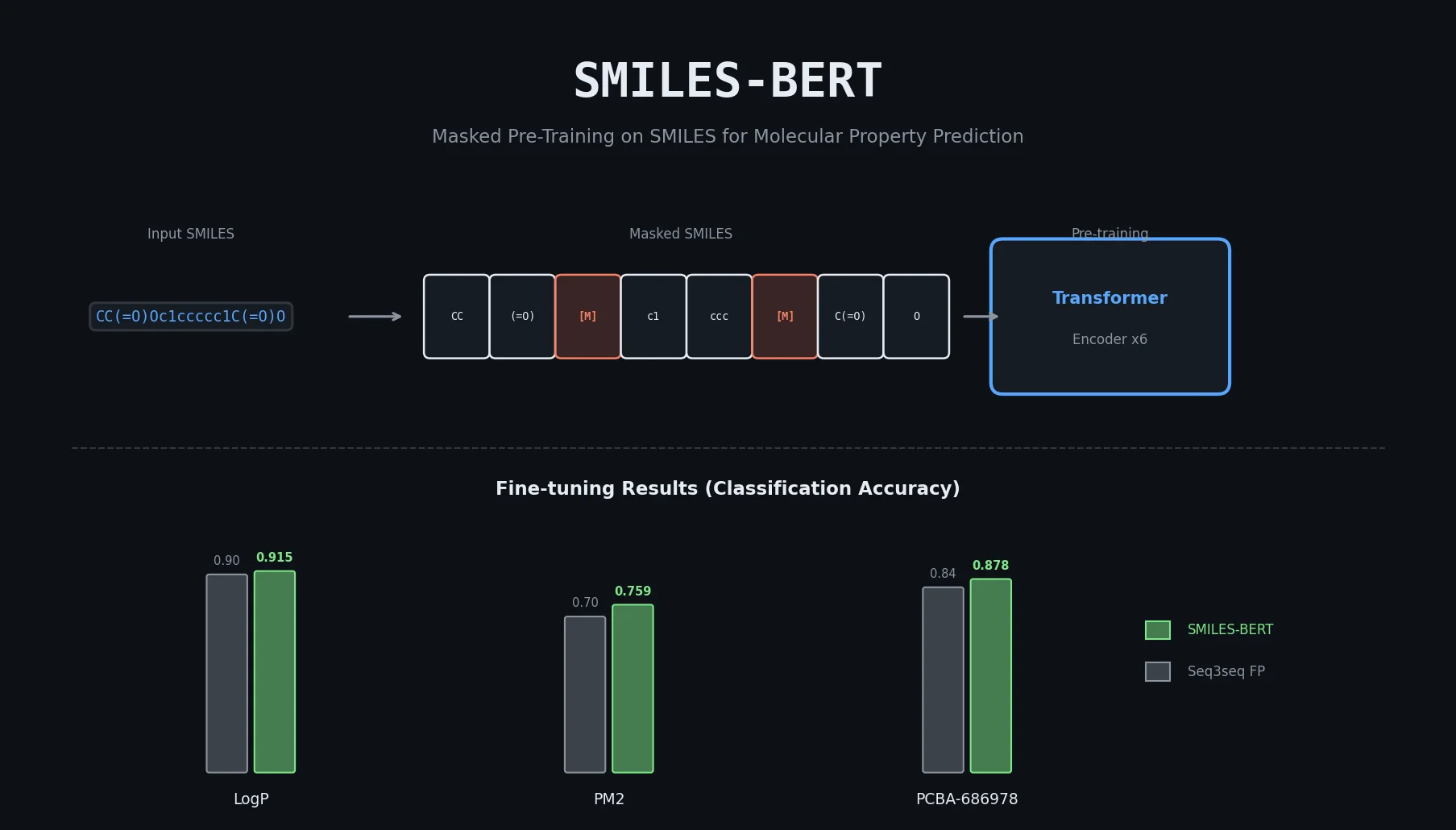
| SMILES-BERT | 2019 | BERT | Masked pretraining on 18M+ SMILES from ZINC |
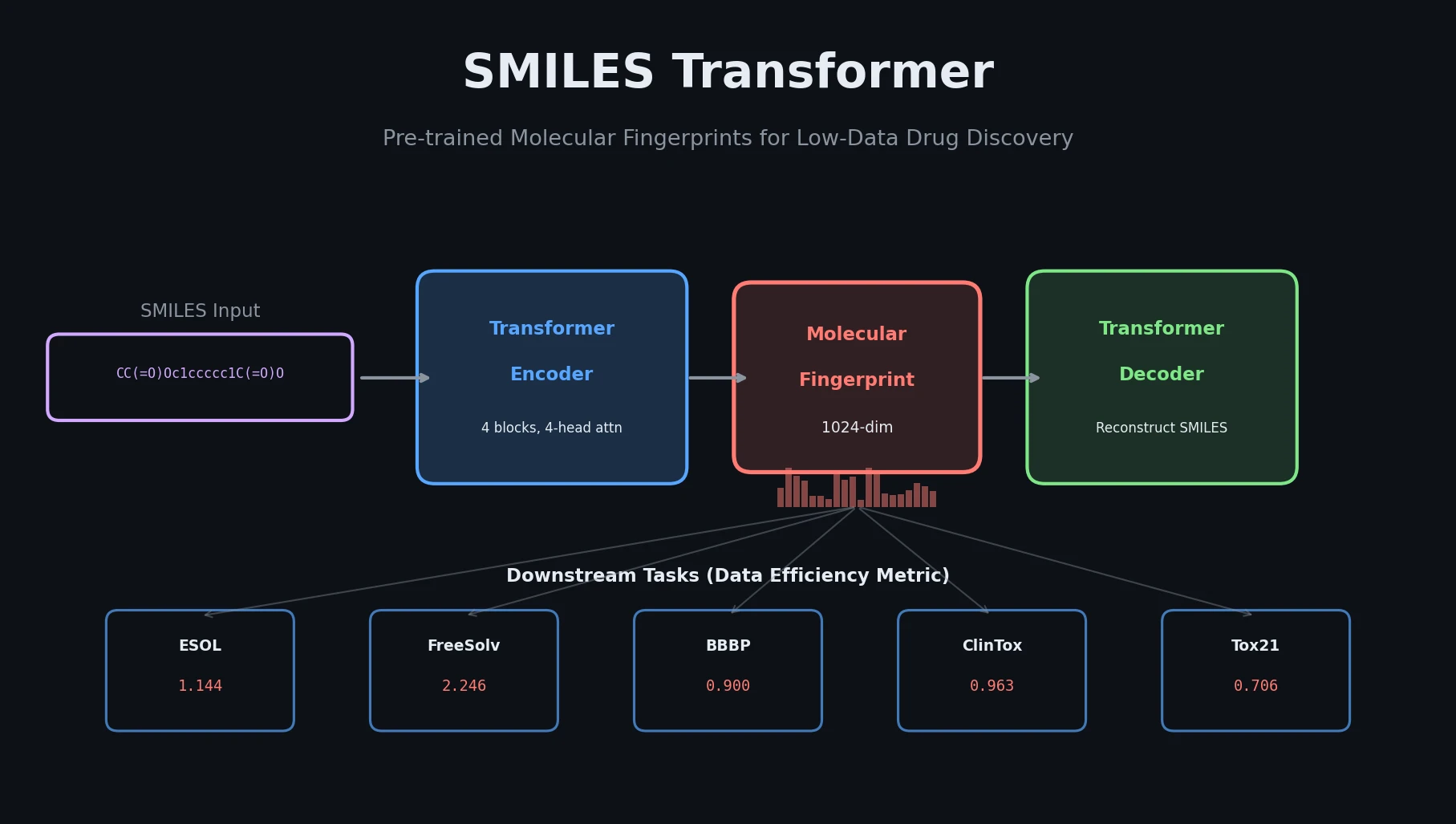
| SMILES Transformer | 2019 | Transformer | Unsupervised transformer fingerprints for low-data tasks |
| ChemBERTa | 2020 | RoBERTa | RoBERTa evaluation on 77M PubChem SMILES |
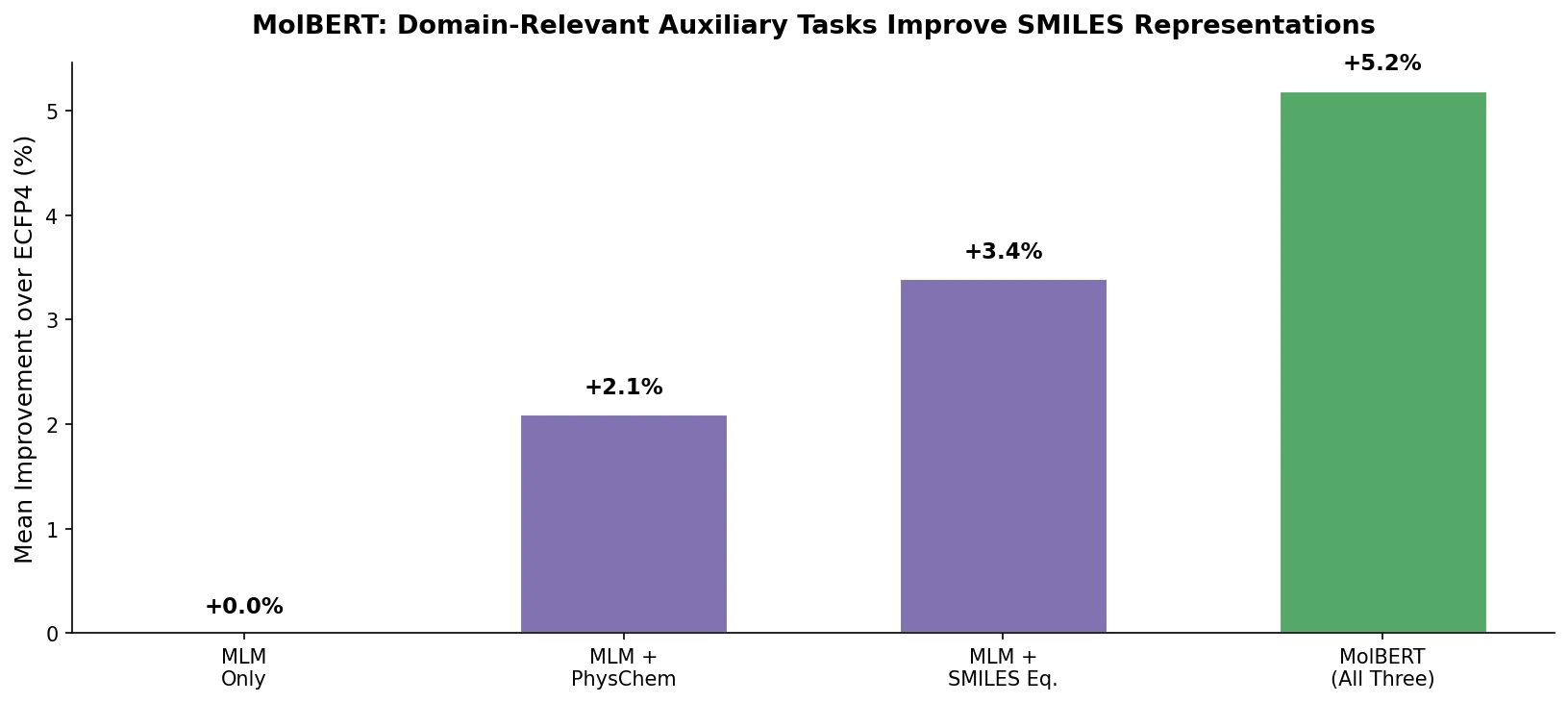
| MolBERT | 2020 | BERT | Domain-relevant auxiliary task pretraining |
| X-MOL | 2020 | Transformer | Large-scale pretraining on 1.1 billion molecules |
| ChemBERTa-2 | 2022 | RoBERTa | Scaling to 77M molecules, comparing MLM vs MTR objectives |
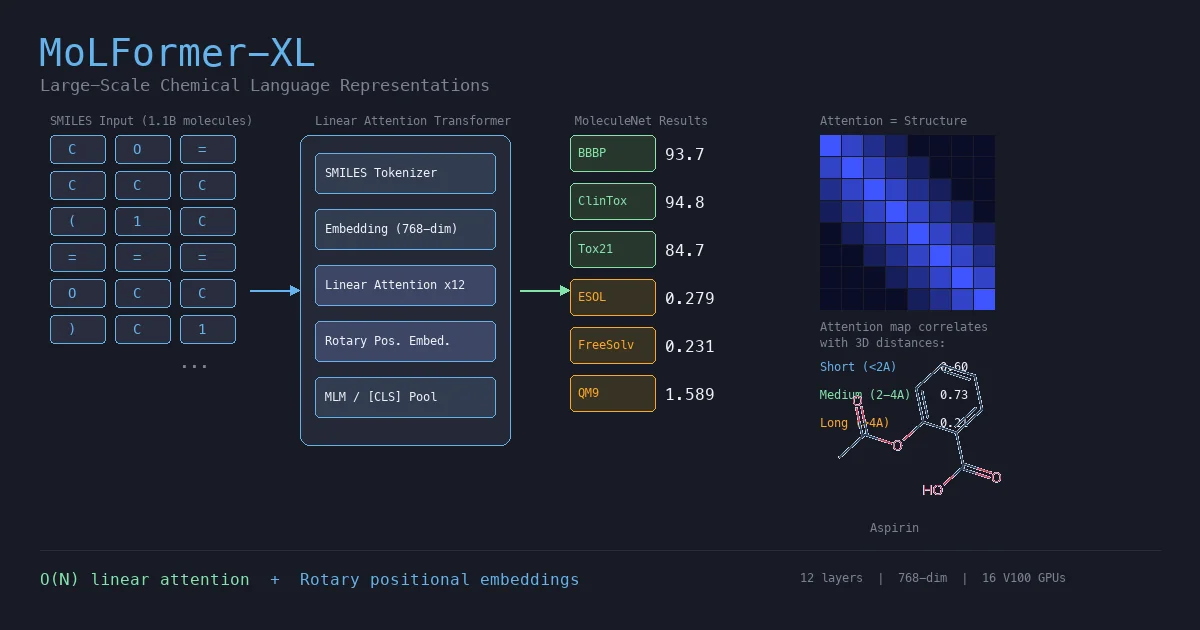
| MoLFormer | 2022 | Linear-attention Transformer | Linear attention at 1.1B molecule scale |
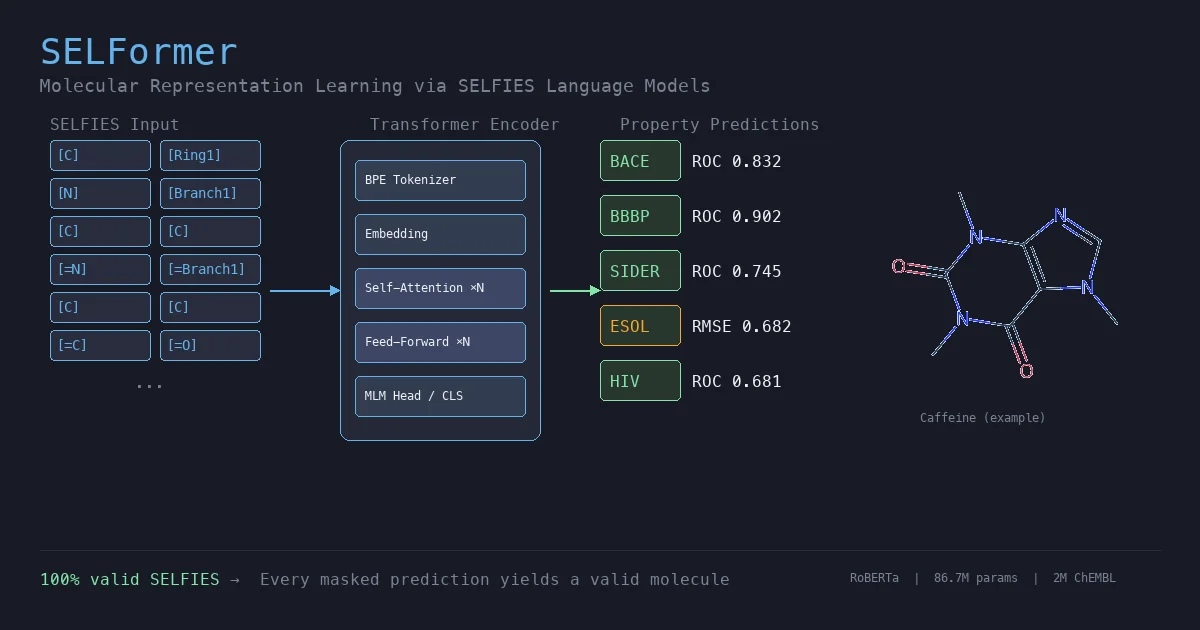
| SELFormer | 2023 | RoBERTa | SELFIES-based pretraining on 2M ChEMBL molecules |
| BARTSmiles | 2024 | BART | Denoising pretraining on 1.7B SMILES from ZINC20 |
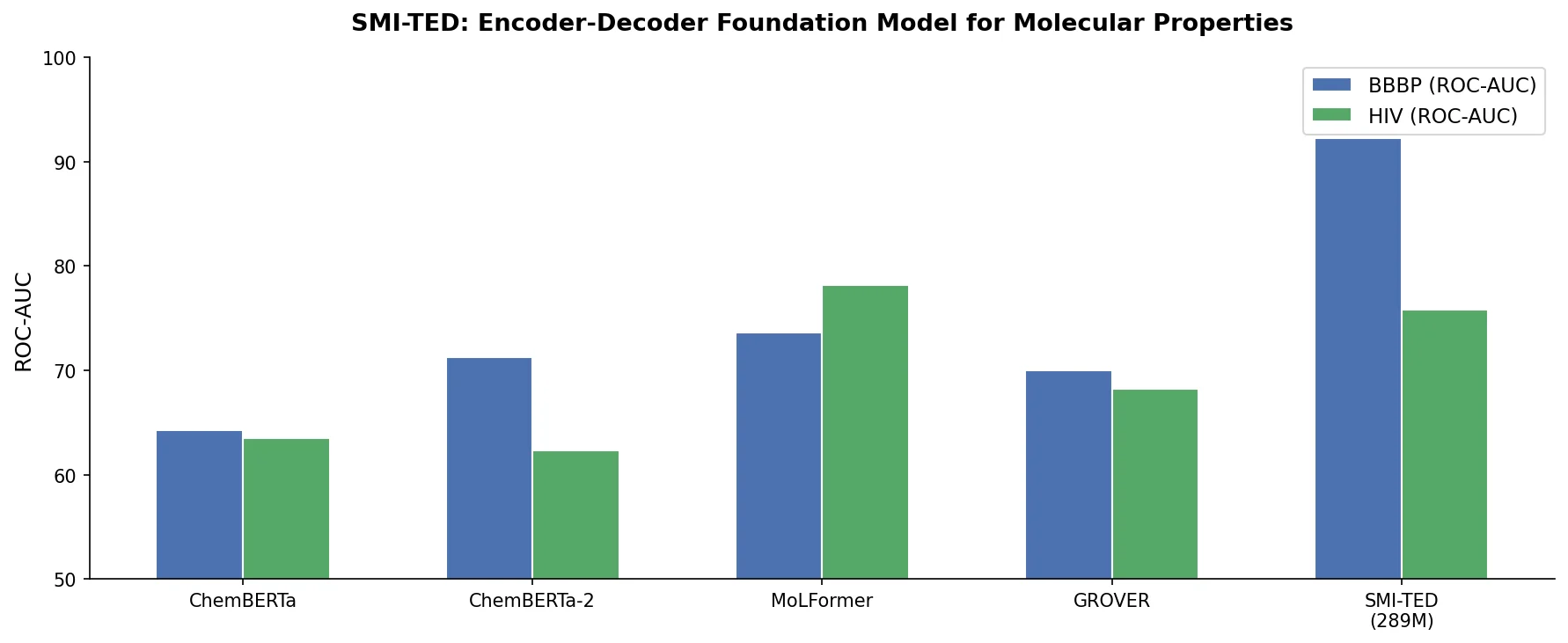
| SMI-TED | 2025 | Encoder-decoder Transformer | Foundation model trained on 91M PubChem molecules |
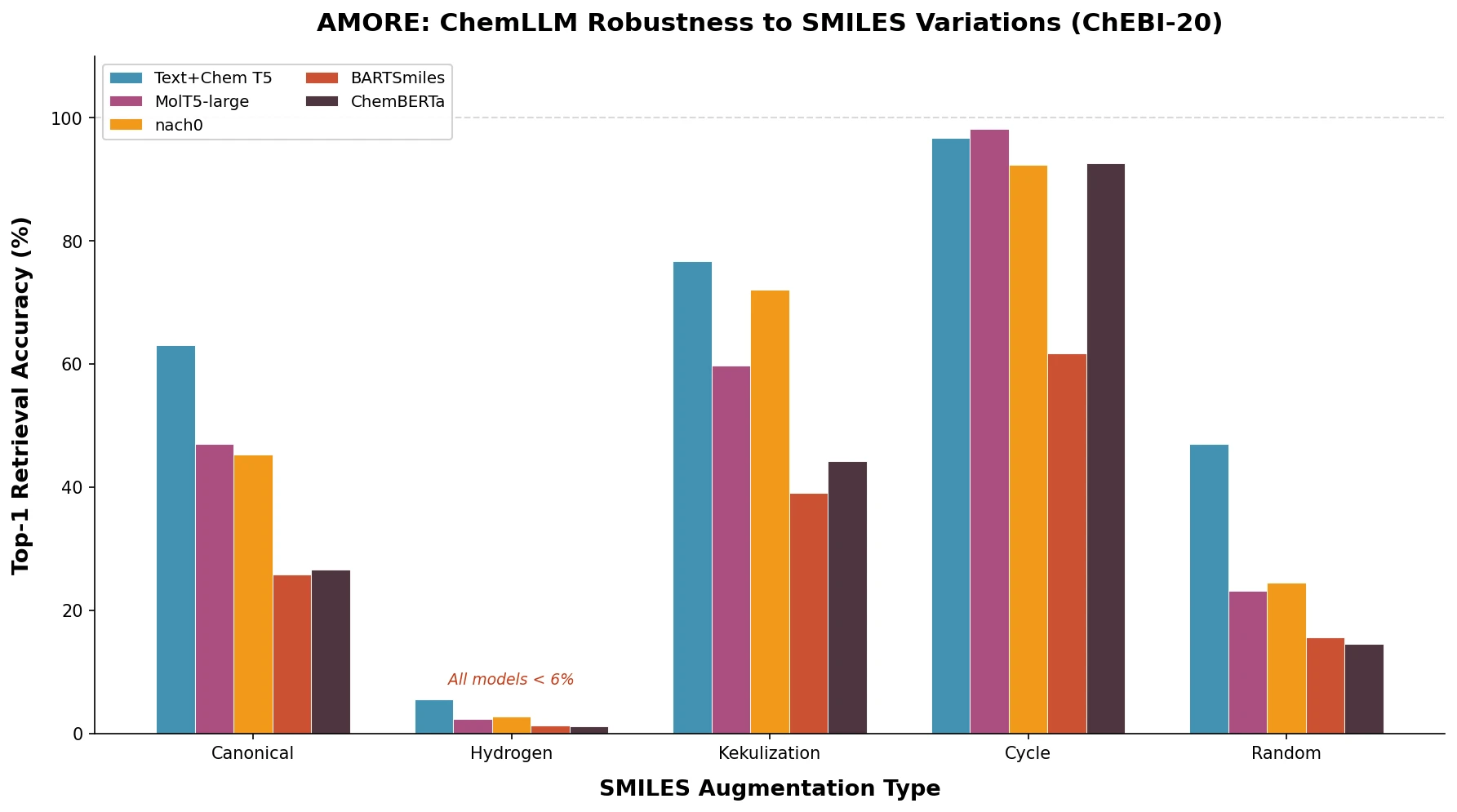
| AMORE | 2025 | Evaluation framework | Testing ChemLLM robustness to SMILES variants |
| ChemBERTa-3 | 2026 | RoBERTa | Open-source scalable chemical foundation models |