Methodological and Resource Contributions
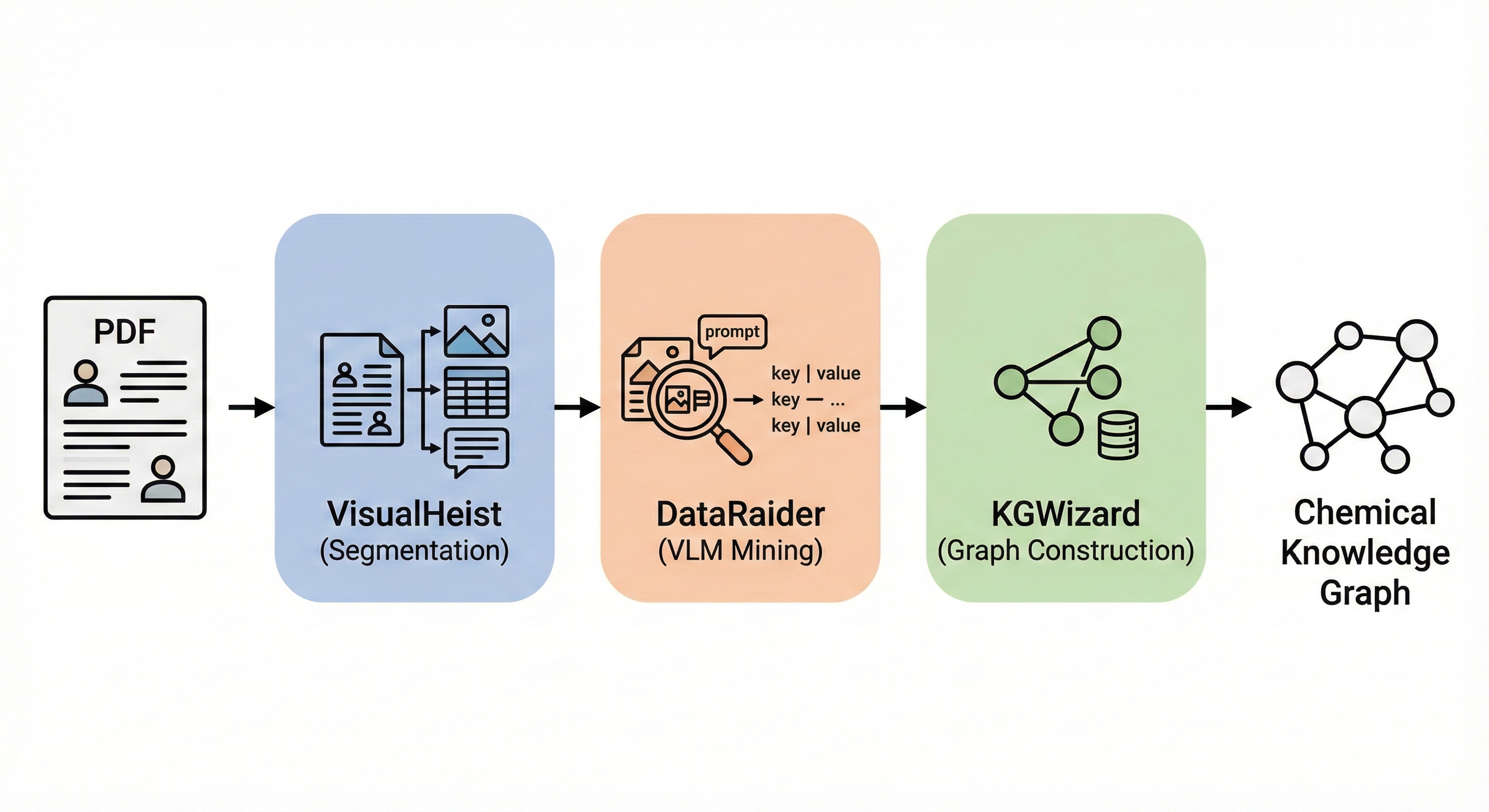
This is primarily a Methodological paper ($\Psi_{\text{Method}}$) that introduces a novel pipeline (MERMaid) for extracting structured chemical data from unstructured PDF documents. It proposes a specific architecture combining fine-tuned vision models (VisualHeist) with vision-language models (DataRaider) and a retrieval-augmented generation system (KGWizard) to solve the problem of multimodal data ingestion.
Secondarily, it is a Resource paper ($\Psi_{\text{Resource}}$) as it releases the source code, prompts, and a new benchmark dataset (MERMaid-100) consisting of annotated reaction data across three chemical domains.
The Inaccessibility of Diagrammatic Reaction Data
- Data Inaccessibility: A significant volume of chemical knowledge currently resides in “print-optimized” PDF formats, specifically within graphical elements like figures, schemes, and tables, which resist standard text mining.
- Limitations of Prior Work: Existing tools (e.g., ChemDataExtractor, OpenChemIE) focus primarily on text, struggle with multimodal parsing, or lack the “contextual awareness” needed to interpret implicit information (e.g., “standard conditions” with modifications in optimization tables).
- Need for Structured Data: To enable self-driving laboratories and data-driven discovery, this unstructured literature must be converted into machine-actionable formats like knowledge graphs.
The MERMaid Pipeline: Vision Models and LLM RAG
- VisualHeist (Fine-tuned Segmentation): A custom fine-tuned model based on Microsoft’s Florence-2 that accurately segments figures, captions, and footnotes, even in messy supplementary materials.
- DataRaider (Context-Aware Extraction): A VLM-powered module (using GPT-4o) with a two-step prompt framework that performs “self-directed context completion.” It can infer missing reaction parameters from context and resolve footnote labels (e.g., linking “condition a” in a table to its footnote description).
- KGWizard (Schema-Adaptive Graph Construction): A text-to-graph engine that uses LLMs as higher-order functions to synthesize parsers dynamically. It employs Retrieval-Augmented Generation (RAG) to check for existing nodes during creation, implicitly resolving coreferences (e.g., unifying “MeCN” and “Acetonitrile”).
- Topic-Agnostic Design: MERMaid features a flexible design that works across three distinct domains: organic electrosynthesis, photocatalysis, and organic synthesis.
Benchmarking Segmentation and Extraction Accuracy
- Segmentation Benchmarking: The authors compared VisualHeist against OpenChemIE (LayoutParser) and PDFigCapX using a dataset of 121 PDFs from 5 publishers.
- End-to-End Extraction: Evaluated the full pipeline on MERMaid-100, a curated dataset of 100 articles across three domains (organic electrosynthesis, photocatalysis, organic synthesis).
- Validating extraction of specific parameters (e.g., catalysts, solvents, yields) using “hard-match” accuracy.
- Knowledge Graph Construction: Automatically generated knowledge graphs for the three domains and assessed the structural integrity and coreference resolution accuracy.
End-to-End Extraction Performance
- Segmentation Results: VisualHeist achieved >93% F1 score across all document types (including pre-2000 papers and supplementary materials), outperforming OpenChemIE by 15-75% and PDFigCapX by 28-75% across all metrics.
- Extraction Accuracy: DataRaider achieved >92% accuracy for VLM-based parameter extraction and near-unity accuracy for domain-specific reaction parameters (e.g., anode, cathode, photocatalyst).
- Graph Building: KGWizard achieved 96% accuracy in node creation and coreference resolution.
- Overall Performance: The pipeline demonstrated an 87% end-to-end overall accuracy.
- Limitations: The architecture relies heavily on closed-weight models (GPT-4o) for reasoning and graph construction, which risks future reproducibility if API snapshots are deprecated. Additionally, the system remains vulnerable to cumulative error propagation from upstream OCR/OCSR tools like RxnScribe.
- Availability: The authors provide a modular, extensible framework that can be adapted to other scientific domains.
Reproducibility Details
Data
- Training Data (VisualHeist):
- Dataset of 3,435 figures and 1,716 tables annotated from 3,518 PDF pages.
- Includes main text, supplementary materials, and unformatted archive papers.
- Evaluation Data (MERMaid-100):
- 100 PDF articles curated from three domains: organic electrosynthesis, photocatalysis, and organic synthesis.
- Includes 104 image-caption/table-heading pairs relevant to reaction optimization.
- Available for download at Zenodo (DOI: 10.5281/zenodo.14917752).
Algorithms
- Two-Step Prompt Framework (DataRaider):
- Step 1: Generic base prompt + domain keys to extract “reaction dictionaries” and “footnote dictionaries”. Uses “fill-in-the-blank” inference for missing details.
- Step 2: Safety check prompt where the VLM updates the reaction dictionary using the footnote dictionary to resolve entry-specific modifications.
- LLM-Synthesized Parsers (KGWizard):
- Uses LLM as a function $g_{A,B}: A \times B \rightarrow (X \rightarrow Y)$ to generate Python code (parsers) dynamically based on input schema instructions.
- RAG for Coreference:
- During graph construction, the system queries the existing database for matching values (e.g., “MeCN”) before creating new nodes to prevent duplication.
- Batching:
- Articles processed in dynamic batch sizes (starting at 1, increasing to 30) to balance speed and redundancy checks.
Models
- VisualHeist: Fine-tuned Florence-2-large (Microsoft vision foundation model).
- Hyperparameters: 12 epochs, learning rate $5 \times 10^{-6}$, batch size 4.
- DataRaider & KGWizard: GPT-4o (version
gpt-4o-2024-08-06). Note: Requires an active OpenAI API key. The pipeline’s long-term reproducibility is currently tied to the continued availability of this specific closed-source endpoint. - RxnScribe: Used for Optical Chemical Structure Recognition (OCSR) to convert reactant/product images to SMILES.
Evaluation
- Metrics:
- Segmentation: Precision, Recall, F1, Accuracy.
- Caption Extraction: Evaluated via Jaccard similarity, mapping predicted token sets $A$ and true token sets $B$ to a threshold condition: $$J(A, B) = \frac{|A \cap B|}{|A \cup B|} \ge 0.70$$
- Data Extraction: Evaluated via Hard-Match accuracy, requiring exact correspondence between predicted sets ($\hat{Y}$) and ground-truth parameters ($Y$) for specific roles (e.g., anode vs. cathode): $$\text{HMA} = \frac{1}{|N|} \sum_{i=1}^{N} \mathbb{1}[y_i = \hat{y}_i]$$
- Baselines: OpenChemIE (LayoutParser + EasyOCR) and PDFigCapX.
Hardware
- Training (VisualHeist): 2x NVLINK Nvidia RTX A6000 GPUs (48GB VRAM) + Intel Xeon w7-2495X CPU (48 cores).
- DataRaider Evaluation: 13th Gen Intel Core i7-1360P CPU (12 cores).
- Inference Costs:
- DataRaider: ~$0.051 per image.
- KGWizard: ~$0.40 per JSON.
- Timing:
- VisualHeist inference: ~4.5 seconds/image.
- DataRaider inference: ~41.3 seconds/image.
- KGWizard processing: ~110.6 seconds/file.
Paper Information
Citation: Leong, S. X., Pablo-García, S., Wong, B., & Aspuru-Guzik, A. (2025). MERMaid: Universal multimodal mining of chemical reactions from PDFs using vision-language models. Matter, 8(12), 102331. https://doi.org/10.1016/j.matt.2025.102331
Publication: Matter, 2025
Artifacts:
| Artifact | Type | License | Notes |
|---|---|---|---|
| GitHub Repository | Code | MIT | Official implementation (VisualHeist, DataRaider, KGWizard) |
| Zenodo Data/Prompts | Dataset | Unknown | MERMaid-100 benchmark, prompts, and raw VLM responses |
@article{leong2025mermaid,
title={MERMaid: Universal multimodal mining of chemical reactions from PDFs using vision-language models},
author={Leong, Shi Xuan and Pablo-Garc{\'i}a, Sergio and Wong, Brandon and Aspuru-Guzik, Al{\'a}n},
journal={Matter},
volume={8},
number={12},
pages={102331},
year={2025},
doi={10.1016/j.matt.2025.102331}
}