Translating Chemical Names to Structures with Transformers
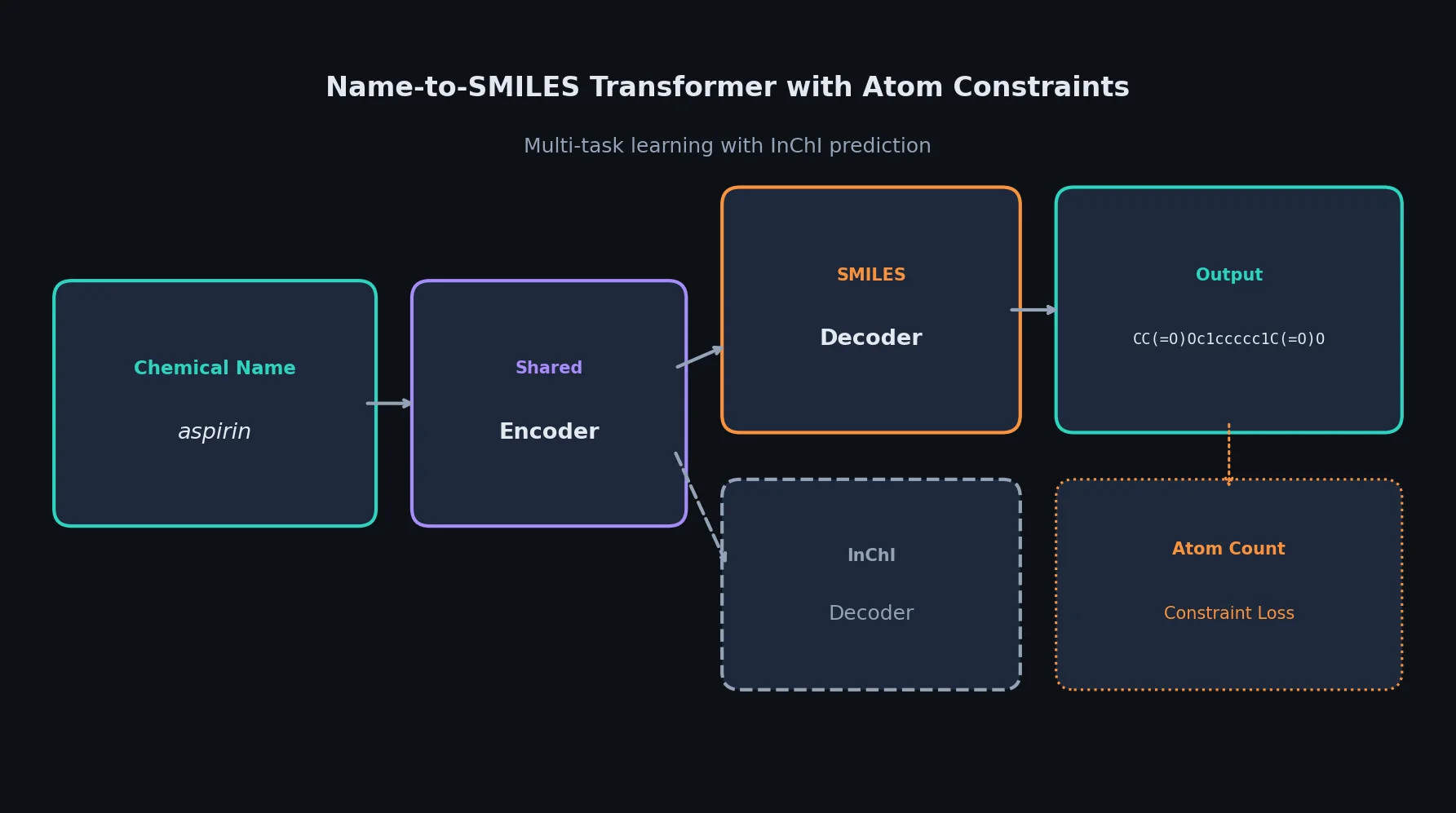
This is a Method paper that proposes using Transformer-based sequence-to-sequence models to predict chemical compound structures (represented as SMILES strings) from chemical compound names. The primary contribution is the application of neural machine translation techniques to the name-to-structure problem, along with two domain-specific improvements: an atom-count constraint loss function and a multi-task learning approach that jointly predicts SMILES and InChI strings.
Why Rule-Based Name-to-Structure Fails for Synonyms
Chemical compound names come in several varieties. IUPAC names follow systematic nomenclature and are well-handled by rule-based parsers like OPSIN. Database IDs (e.g., CAS registry numbers) can be resolved by dictionary lookup. The third category, Synonyms (which includes abbreviations, common names, and other informal designations), is problematic because naming patterns are complex and widely variable.
In preliminary experiments, rule-based tools achieved F-measures of 0.878 to 0.960 on IUPAC names but only 0.719 to 0.758 on Synonyms. This performance gap motivates a data-driven approach. The authors frame name-to-SMILES prediction as a machine translation problem: the source language is the chemical compound name and the target language is the SMILES string. A neural model trained on millions of name-SMILES pairs can learn patterns that rule-based systems miss, particularly for non-systematic nomenclature.
Atom-Count Constraints and Multi-Task Learning
The paper introduces two improvements over a vanilla Transformer seq2seq model.
Atom-Count Constraint Loss
A correct structure prediction must contain the right number of atoms of each element. The authors add an auxiliary loss that penalizes the squared difference between the predicted and true atom counts for each element. The predicted atom counts are obtained by summing Gumbel-softmax outputs across all decoded positions.
For the $i$-th output token, the Gumbel-softmax probability vector is:
$$ y_{ij} = \frac{\exp\left((\log(\pi_{ij}) + g_{ij}) / \tau\right)}{\sum_{k=1}^{|\mathcal{V}|} \exp\left((\log(\pi_{ik}) + g_{ik}) / \tau\right)} $$
where $\pi_{ij}$ is the model’s softmax output, $g_{ij}$ is a Gumbel noise sample, and $\tau = 0.1$ is the temperature. The predicted token frequency vector is $\mathbf{y}^{pred} = \sum_{i=1}^{m} \mathbf{y}_i$, and the atom-count loss is:
$$ \mathcal{L}_{atom} = \frac{1}{|A|} \sum_{a \in A} \left(N_a(T) - y_{idx(a)}^{pred}\right)^2 $$
where $A$ is the set of chemical elements in the vocabulary, $N_a(T)$ returns the number of atoms of element $a$ in the correct SMILES string $T$, and $idx(a)$ returns the vocabulary index of element $a$. Only element tokens (e.g., “C”, “O”) are counted; bond symbols (e.g., “=”, “#”) are excluded.
The combined objective is:
$$ \mathcal{L}_{smiles} + \lambda_{atom} \mathcal{L}_{atom} $$
with $\lambda_{atom} = 0.7$.
Multi-Task SMILES/InChI Prediction
SMILES and InChI strings encode the same chemical structure in different formats. The authors hypothesize that jointly predicting both representations can improve the shared encoder. The multi-task model shares the encoder between a SMILES decoder and an InChI decoder, minimizing:
$$ \mathcal{L}_{smiles} + \lambda_{inchi} \mathcal{L}_{inchi} $$
where $\mathcal{L}_{inchi} = -\log P(I | X; \boldsymbol{\theta}_{enc}, \boldsymbol{\theta}_{inchi})$ and $\lambda_{inchi} = 0.3$.
Experimental Setup and Evaluation
Dataset
The dataset was constructed from PubChem dump data (97M compound records). Chemical compound names categorized as Synonyms were paired with canonical SMILES strings (converted via RDKit). Database-like IDs were filtered out using regular expressions. Duplicate names mapping to different CIDs were removed.
| Split | Size |
|---|---|
| Training | 5,000,000 |
| Development | 1,113 |
| Test | 11,194 |
Model Configuration
The Transformer uses 6 encoder/decoder layers, 8 attention heads, 512-dimensional embeddings, and 0.1 dropout. Training used label-smoothing cross-entropy ($\epsilon = 0.1$), Adam optimizer ($\beta_1 = 0.9$, $\beta_2 = 0.98$), and a warmup schedule with peak learning rate 0.0005 over 4,000 steps followed by inverse square root decay. Models were trained for 300,000 update steps. Final predictions averaged the last 10 checkpoints and used beam search (beam size 4, length penalty $\alpha = 0.6$, max output length 200).
Tokenization
Three tokenization strategies were compared:
- BPE: Byte pair encoding learned on chemical compound names (500 merge operations) via fastBPE
- OPSIN-TK: The OPSIN rule-based tokenizer
- OPSIN-TK+BPE: A hybrid where OPSIN handles tokenizable names and BPE handles the rest
SMILES tokens were identified by regular expressions (elements as single tokens, remaining symbols as characters). InChI strings were tokenized by SentencePiece (vocabulary size 1,000).
Baselines
- OPSIN: Open-source rule-based parser
- Tool A and Tool B: Two commercially available name-to-structure tools
Results
| Method | Tokenizer | Recall | Precision | F-measure |
|---|---|---|---|---|
| OPSIN | Rule-based | 0.693 | 0.836 | 0.758 |
| Tool A | Rule-based | 0.711 | 0.797 | 0.752 |
| Tool B | Rule-based | 0.653 | 0.800 | 0.719 |
| Transformer | BPE | 0.793 | 0.806 | 0.799 |
| + atomnum | BPE | 0.798 | 0.808 | 0.803 |
| + inchigen | BPE | 0.810 | 0.819 | 0.814 |
| Transformer | OPSIN-TK+BPE | 0.763 | 0.873 | 0.814 |
| + atomnum | OPSIN-TK+BPE | 0.768 | 0.876 | 0.818 |
| + inchigen | OPSIN-TK+BPE | 0.779 | 0.886 | 0.829 |
| Transformer | OPSIN-TK | 0.755 | 0.868 | 0.808 |
| + atomnum | OPSIN-TK | 0.757 | 0.867 | 0.808 |
| + inchigen | OPSIN-TK | 0.754 | 0.869 | 0.807 |
The best configuration (inchigen with OPSIN-TK+BPE) achieved an F-measure of 0.829, surpassing OPSIN by 0.071 points. The multi-task learning approach (inchigen) consistently outperformed the atom-count constraint alone (atomnum) across all tokenizer settings.
Key Findings and Error Analysis
The Transformer-based approach produced grammatically correct SMILES strings (parseable by RDKit) for 99% of test examples, compared to 81.6-88.4% for the rule-based tools. Even when predictions were incorrect, they tended to be structurally similar to the correct answer. Using MACCS fingerprints and Jaccard (Tanimoto) similarity, the average similarity between incorrectly predicted and correct structures was 0.753.
The OPSIN-TK tokenizer yielded higher precision than BPE because approximately 11.5% (1,293 of 11,194) of test compounds could not be tokenized by OPSIN, reducing the number of outputs. BPE-based tokenizers achieved higher recall by covering all inputs. The hybrid OPSIN-TK+BPE approach balanced both, achieving the highest overall F-measure.
Limitations: The paper does not evaluate on IUPAC names separately with the Transformer models (only comparing rule-based tools on IUPAC). The atom-count constraint and multi-task learning are not combined in a single model. The dataset is released but the training code is not. Hardware details and training times are not reported. The evaluation uses only exact-match F-measure and Jaccard similarity, without measuring partial credit for nearly-correct structures.
Future work: The authors plan to explore additional tokenization methods, combine the atom-count constraint with multi-task learning, and apply the constraint loss to other chemistry problems including chemical reaction prediction.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Training | PubChem Synonyms (custom split) | 5,000,000 pairs | Chemical compound names to canonical SMILES |
| Development | PubChem Synonyms (custom split) | 1,113 pairs | Filtered for duplicates |
| Test | PubChem Synonyms (custom split) | 11,194 pairs | Filtered for duplicates; released as benchmark |
The authors state the dataset is released for future research. The data was constructed from the PubChem dump (97M compound records) using RDKit for SMILES canonicalization. Database-like IDs were removed with regular expressions and duplicate names across CIDs were filtered.
Algorithms
- Transformer seq2seq (6 layers, 8 heads, 512-dim embeddings)
- BPE tokenization via fastBPE (500 merge operations)
- SentencePiece for InChI tokenization (vocabulary size 1,000)
- Gumbel-softmax atom-count constraint ($\tau = 0.1$, $\lambda_{atom} = 0.7$)
- Multi-task SMILES/InChI loss ($\lambda_{inchi} = 0.3$)
- Adam optimizer ($\beta_1 = 0.9$, $\beta_2 = 0.98$, $\epsilon = 10^{-8}$)
- Label smoothing ($\epsilon = 0.1$), 300K training steps
- Beam search (beam size 4, length penalty $\alpha = 0.6$)
Models
Standard Transformer architecture following Vaswani et al. (2017). No pre-trained weights or model checkpoints are released.
Evaluation
| Metric | Best Value | Model | Notes |
|---|---|---|---|
| F-measure | 0.829 | inchigen (OPSIN-TK+BPE) | Highest overall |
| Precision | 0.886 | inchigen (OPSIN-TK+BPE) | Highest overall |
| Recall | 0.810 | inchigen (BPE) | Highest overall |
| Grammatical correctness | 99% | inchigen (BPE) | SMILES parseable by RDKit |
| Avg. Jaccard similarity (errors) | 0.753 | inchigen (BPE) | On incorrect predictions only |
Hardware
Not reported.
Paper Information
Citation: Omote, Y., Matsushita, K., Iwakura, T., Tamura, A., & Ninomiya, T. (2020). Transformer-based Approach for Predicting Chemical Compound Structures. Proceedings of the 1st Conference of the Asia-Pacific Chapter of the Association for Computational Linguistics and the 10th International Joint Conference on Natural Language Processing, 154-162. https://doi.org/10.18653/v1/2020.aacl-main.19
@inproceedings{omote2020transformer,
title={Transformer-based Approach for Predicting Chemical Compound Structures},
author={Omote, Yutaro and Matsushita, Kyoumoto and Iwakura, Tomoya and Tamura, Akihiro and Ninomiya, Takashi},
booktitle={Proceedings of the 1st Conference of the Asia-Pacific Chapter of the Association for Computational Linguistics and the 10th International Joint Conference on Natural Language Processing},
pages={154--162},
year={2020},
publisher={Association for Computational Linguistics},
doi={10.18653/v1/2020.aacl-main.19}
}