A Method for Neural Translation of Chemical Names
This is a Method paper that introduces deep learning approaches for translating chemical nomenclature between English and Chinese. The primary contribution is demonstrating that character-level sequence-to-sequence neural networks (both CNN-based and LSTM-based) can serve as viable alternatives to hand-crafted rule-based translation systems for chemical names. The work compares two neural architectures against an existing rule-based tool on bilingual chemical name datasets.
Bridging the English-Chinese Chemical Nomenclature Gap
English and Chinese are the two most widely used languages for chemical nomenclature worldwide. Translation between them is important for chemical data processing, especially for converting Chinese chemical names extracted via named entity recognition into English names that existing name-to-structure tools can parse. Rule-based translation between these languages faces considerable challenges:
- Chinese chemical names lack word boundaries (no spaces), making segmentation difficult.
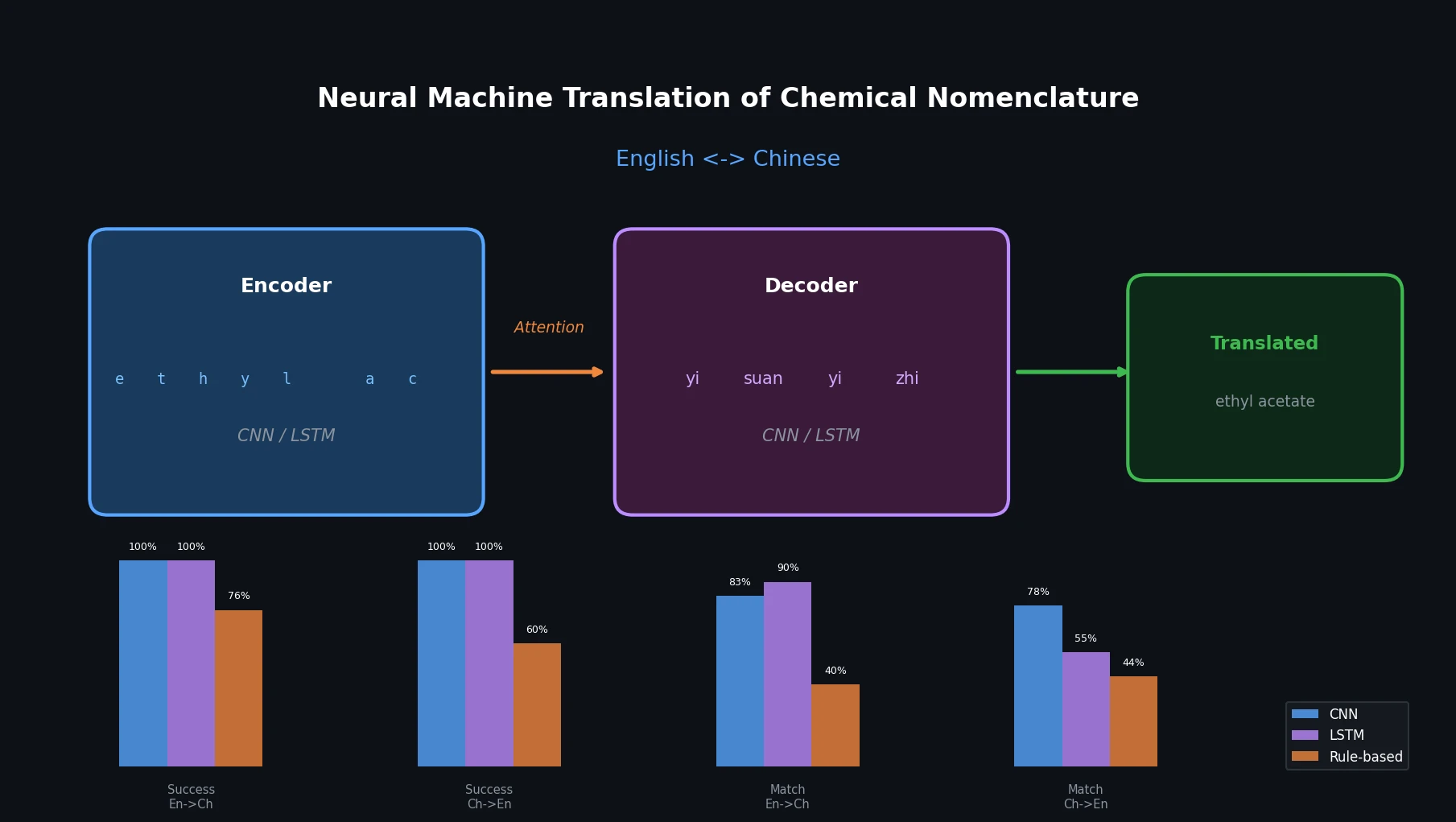
- Word order is often reversed between English and Chinese chemical names (e.g., “ethyl acetate” maps to characters meaning “acetate-ethyl” in Chinese).
- The same English morpheme can map to different Chinese characters depending on chemical context (e.g., “ethyl” translates differently in “ethyl acetate” vs. “ethyl alcohol”).
- Trivial names, especially for natural products, follow irregular translation patterns or are transliterations.
Building comprehensive rule sets requires a formally trained chemist fluent in both languages, making rule-based approaches expensive and fragile.
Character-Level Sequence-to-Sequence Translation
The core idea is to treat chemical name translation as a character-level machine translation task, applying encoder-decoder architectures with attention mechanisms. Two architectures are proposed:
CNN-based architecture: Three 1D convolutional layers encode the input character sequence. A decoder with three 1D convolutional layers processes the target sequence offset by one timestep, combined with attention mechanism layers that connect encoder and decoder outputs. Two additional 1D convolutional layers produce the final decoded output sequence.
LSTM-based architecture: An LSTM encoder converts the input sequence into two state vectors. An LSTM decoder is trained with teacher forcing, using the encoder’s state vectors as its initial state, and generating the target sequence offset by one timestep.
Both models operate at the character level. Input chemical name strings are transformed into embedding vectors, with the vocabulary size equal to the number of unique characters in the respective language (100 unique characters for English names, 2,056 unique characters for Chinese names).
Experimental Setup and Comparison with Rule-Based Tool
Datasets
The authors built two directional datasets from a manually curated corpus of scientific literature maintained at their institution:
- En2Ch (English to Chinese): 30,394 name pairs after deduplication
- Ch2En (Chinese to English): 37,207 name pairs after deduplication
The datasets cover systematic compound names through trivial names. For names with multiple valid translations, the most commonly used translation was selected. Each dataset was split 80/20 for training and validation.
Model Configuration
Both neural network models used the following hyperparameters:
- Batch size: 64
- Epochs: 100
- Latent dimensionality: 256 (encoding and decoding space)
- Implementation: Python 3.7 with Keras 2.3 and TensorFlow backend
Evaluation Metrics
The models were evaluated on five metrics across both translation directions:
- Success Rate: Percentage of inputs that produced any output
- String Matching Accuracy: Exact match with the single target name
- Data Matching Accuracy: Exact match allowing any valid translation from the corpus
- Manual Spot Check: Blind evaluation of 100 random samples per approach
- Running Time: Wall-clock time on the same hardware
Baseline
The rule-based comparison system operates in three steps: disassemble the input name into word fragments, translate each fragment, and reassemble into the target language. This tool had been deployed as an online service with over one million uses at the time of publication.
Key Findings and Limitations
Main Results
| Metric | CNN | LSTM | Rule-based |
|---|---|---|---|
| Success Rate En2Ch | 100% | 100% | 75.97% |
| Success Rate Ch2En | 100% | 100% | 59.90% |
| String Match En2Ch | 82.92% | 89.64% | 39.81% |
| String Match Ch2En | 78.11% | 55.44% | 43.77% |
| Data Match En2Ch | 84.44% | 90.82% | 45.15% |
| Data Match Ch2En | 80.22% | 57.40% | 44.91% |
| Manual Check En2Ch | 90.00% | 89.00% | 80.00% |
| Manual Check Ch2En | 82.00% | 61.00% | 78.00% |
| Time En2Ch (s) | 1423 | 190 | 288 |
| Time Ch2En (s) | 1876 | 303 | 322 |
Both neural approaches achieved 100% success rate (always producing output), while the rule-based tool failed on 24% and 40% of inputs for En2Ch and Ch2En respectively. The rule-based tool’s failures were concentrated on Chinese names lacking word boundaries and on trivial names of natural products.
For English-to-Chinese translation, LSTM performed best at 89.64% string matching accuracy (90.82% data matching), followed by CNN at 82.92%. For Chinese-to-English, CNN substantially outperformed LSTM (78.11% vs. 55.44% string matching), suggesting that LSTM had difficulty with long-term dependencies in Chinese character sequences. The authors observed that many LSTM errors appeared at the ends of chemical names.
Analysis by Name Type
The CNN-based approach outperformed LSTM on CAS names (80% vs. 52% in manual checks) and was more robust for longer names. The rule-based tool showed consistent performance regardless of name length, suggesting it was more suited to regular systematic names but struggled with the diversity of real-world chemical nomenclature.
Limitations
- Performance depends heavily on training data quality and quantity.
- Neither neural approach was validated on an external test set outside the institution’s corpus.
- The CNN model was considerably slower (5-6x) than the other two approaches.
- No comparison against modern transformer-based NMT architectures (the study predates widespread adoption of transformers for this task).
- The dataset is relatively small by modern NMT standards (30-37K pairs).
- The authors noted that some neural translations were actually better than the target labels, suggesting the evaluation metrics understate true performance.
Future Directions
The authors suggest that combining CNN and LSTM architectures could yield further improvements, and that the approach has practical applications in scientific publishing (Chinese journals requiring English abstracts) and chemical database interoperability.
Reproducibility Details
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Training/Validation (En2Ch) | Curated bilingual corpus | 30,394 pairs | 80/20 split, from SIOC chemical data system |
| Training/Validation (Ch2En) | Curated bilingual corpus | 37,207 pairs | 80/20 split, from SIOC chemical data system |
| Testing (En2Ch) | Held-out validation split | 6,079 records | Same source |
| Testing (Ch2En) | Held-out validation split | 7,441 records | Same source |
Training data, Python code for both models, and result data are provided as supplementary files with the paper.
Algorithms
- Character-level CNN encoder-decoder with attention (3+3+2 conv layers)
- Character-level LSTM encoder-decoder with teacher forcing
- Batch size: 64, epochs: 100, latent dim: 256
Models
Both models implemented in Python 3.7 with Keras 2.3 / TensorFlow. No pre-trained weights are released separately, but the training code is provided as supplementary material.
Evaluation
| Metric | Best Value (En2Ch) | Best Value (Ch2En) | Notes |
|---|---|---|---|
| Success Rate | 100% (both DL) | 100% (both DL) | Rule-based: 75.97% / 59.90% |
| String Matching | 89.64% (LSTM) | 78.11% (CNN) | Best neural model per direction |
| Data Matching | 90.82% (LSTM) | 80.22% (CNN) | Allows multiple valid translations |
| Manual Spot Check | 90.00% (CNN) | 82.00% (CNN) | Blind evaluation of 100 samples |
Hardware
Not specified in the paper. Running times reported but hardware details not provided.
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| Supplementary files | Code + Data | CC-BY 4.0 | Training data, CNN/LSTM code, results (Additional files 1-6) |
| SIOC Translation Tool | Other | Not specified | Rule-based baseline tool, online service |
Paper Information
Citation: Xu, T., Chen, W., Zhou, J., Dai, J., Li, Y., & Zhao, Y. (2020). Neural machine translation of chemical nomenclature between English and Chinese. Journal of Cheminformatics, 12, 50. https://doi.org/10.1186/s13321-020-00457-0
@article{xu2020neural,
title={Neural machine translation of chemical nomenclature between English and Chinese},
author={Xu, Tingjun and Chen, Weiming and Zhou, Junhong and Dai, Jingfang and Li, Yingyong and Zhao, Yingli},
journal={Journal of Cheminformatics},
volume={12},
pages={50},
year={2020},
doi={10.1186/s13321-020-00457-0},
publisher={Springer}
}