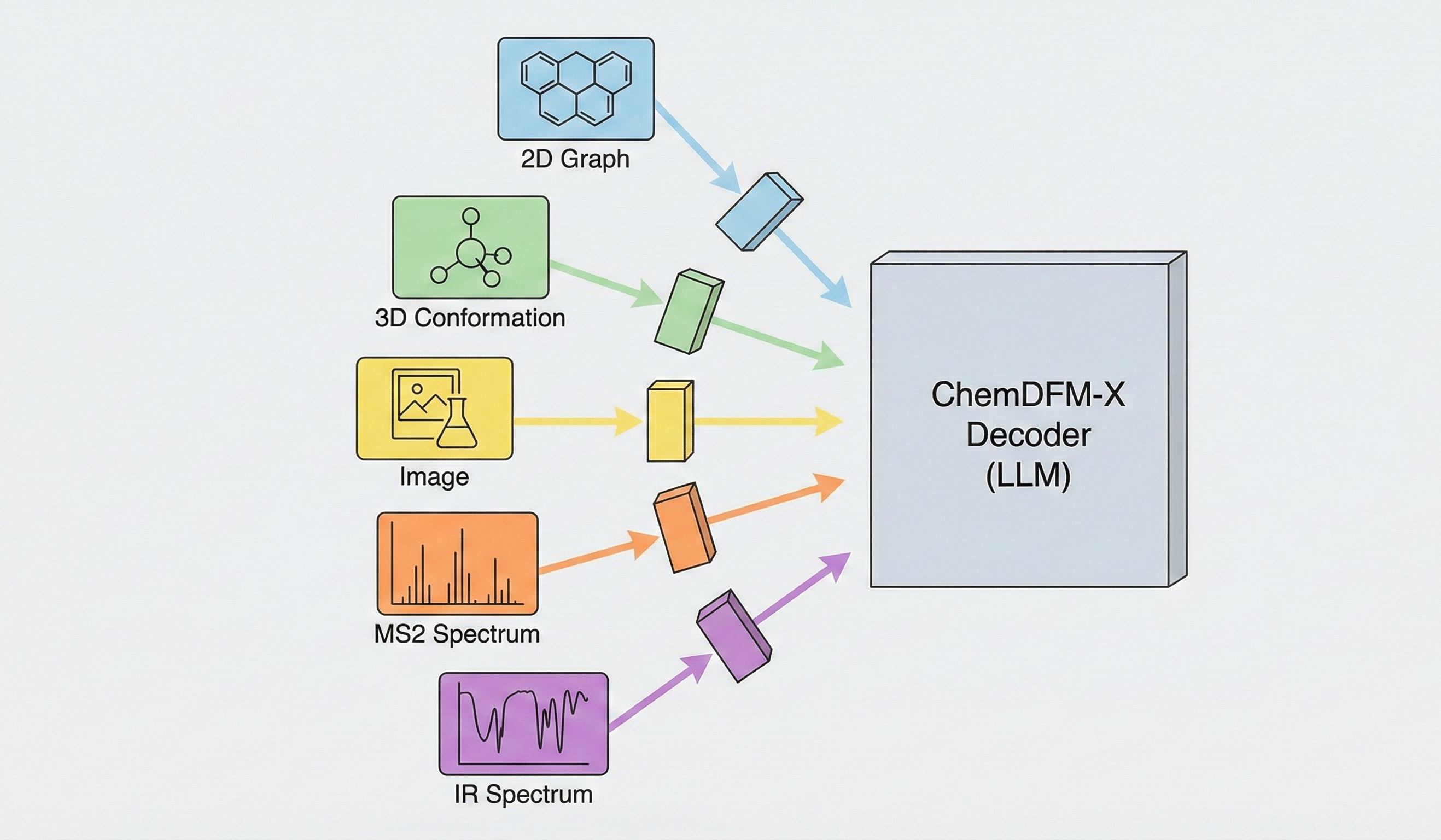
ChemDFM-X Contribution and Architecture
This is primarily a Method paper with a significant Resource contribution.
Method: The paper proposes a novel “Cross-modal Dialogue Foundation Model” architecture that aligns five distinct chemical modalities (2D graphs, 3D conformations, images, MS2 spectra, IR spectra) to a single LLM decoder using separate encoders and projection modules. It establishes strong baseline performance across multiple modalities compared against current generalist models.
Resource: The paper addresses the scarcity of multimodal chemical data by constructing a 7.6M instruction-tuning dataset. This dataset is largely synthesized from seed SMILES strings using approximate calculations (MMFF94, CFM-ID, Chemprop-IR) and specialist model predictions.
Bridging Experimental Data and LLMs
Existing chemical AI models generally fall into two distinct categories. Task-specific specialist models achieve high accuracy on singular objectives, such as property prediction or molecular generation, but require strict formatting and lack conversational flexibility. Conversely, early chemical large language models provide natural language interaction but are restricted to text and SMILES strings. ChemDFM-X addresses this gap by enabling large multimodal models to process the experimental characterization data (MS2 and IR spectra) and visual data routinely used in practical chemistry workflows.
Synthetic Data Scaling for Modality Alignment
The core novelty lies in the “Any-to-Text” alignment strategy via synthetic data scaling:
Comprehensive Modality Support: ChemDFM-X incorporates experimental characterization data (MS2 and IR spectra) alongside 2D graphs, 3D conformations, and images. The data representations are formally defined mathematically rather than as raw pixels:
- Molecular Graph: An undirected graph $G = (\textbf{V}, \textbf{E})$ with atom set $\textbf{V}$ and bond set $\textbf{E}$.
- Molecular Conformation: An undirected graph $G = (\textbf{V}’, \textbf{E})$ storing spatial coordinates: $\textbf{v}_i = (x_i, y_i, z_i, a_i)$.
- MS2 Spectrum: Treated as a point sequence of discrete mass-to-charge ratios and intensities, tokenized via a discrete codebook: $\textbf{M} = ((r_1, I_1), (r_2, I_2), \dots, (r_n, I_n))$.
- IR Spectrum: Treated as a dense sequence of continuous wave lengths and absorption intensities, directly reshaped for feature extraction: $\textbf{R} = ((w_1, t_1), (w_2, t_2), \dots, (w_l, t_l))$.
The authors trained new Sequence Transformer encoders from scratch for the MS2 and IR modalities since suitable pre-trained models did not exist.
Synthetic Data Generation Pipeline: The authors generated a 7.6M sample dataset by starting with 1.3M seed SMILES and using “approximate calculations” to generate missing modalities:
- 3D conformations via MMFF94 force field optimization
- MS2 spectra via CFM-ID 4.0 (Competitive Fragmentation Modeling)
- IR spectra via Chemprop-IR (Message Passing Neural Network)
Cross-Modal Synergy: The model demonstrates that training on reaction images improves recognition performance by leveraging semantic chemical knowledge (reaction rules) to correct visual recognition errors, an emergent capability from multimodal training.
Multimodal Benchmarking with ChemLLMBench
The model was evaluated using a customized version of ChemLLMBench and MoleculeNet across three modality categories:
Structural Modalities (2D Graphs & 3D Conformations):
- Molecule recognition and captioning
- Property prediction (MoleculeNet: BACE, BBBP, ClinTox, HIV, Tox21)
- Compared against specialist models (Mole-BERT, Uni-Mol, MolXPT, MolCA) and generalist models (3D-MoLM, ChemDFM, ChemLLM)
Visual Modalities (Images):
- Single molecule image recognition
- Reaction image recognition
- Compared against GPT-4O, Gemini 1.5 Pro, Qwen-VL, LLaVA, and specialist models MolNextr and MolScribe
Characterization Modalities (MS2 & IR Spectra):
- Spectral analysis tasks (identifying molecules from spectra)
- Contextualized spectral interpretation (combining spectra with reaction context)
- Novel evaluation requiring integration of spectroscopic data with reaction knowledge
Cross-Modal Synergy and Generalist Performance
Key Findings:
Leading Generalist Performance: ChemDFM-X establishes a new benchmark among existing generalist models (such as 3D-MOLM and ChemLLM), achieving performance metrics that match dedicated specialist models across several multimodal tasks.
Failure of General LMMs: General vision models (GPT-4O, Gemini 1.5 Pro, Qwen-VL, LLaVA, InternLM-XComposer2, DocOwl) failed significantly on chemical image recognition tasks (0% accuracy for most models on molecule and reaction recognition, Table 9), demonstrating that chemical domain knowledge cannot be assumed from general pre-training.
Cross-Modal Error Correction: In reaction image recognition, ChemDFM-X achieved higher accuracy (53.0%) than on single molecules (46.0%) (Table 9). The authors conclude the model uses its internal knowledge of chemical reaction rules to correct recognition errors in the visual modality, an emergent capability from multimodal training.
Reliance on Reaction Context for Spectra: In zero-shot scenarios, ChemDFM-X essentially fails at pure spectral recognition (achieving 0% and 1% top-1 accuracy on MS2 and IR spectra alone, Table 11). However, when SMILES-based reaction context is included, performance rises to 45% (MS2) and 64% (IR) on the reaction prediction task, and 29% (MS2) and 60% (IR) on retrosynthesis (Table 11). This indicates the model uses spectral data as a soft prior to constrain textual deductions. Furthermore, the paper compares ChemDFM-X’s spectral identification performance exclusively against text-only LLMs that cannot process spectra, omitting comparisons against established specialist tools.
Surrogate Distillation Trade-offs: Because the spectral training data relies entirely on outputs from CFM-ID 4.0 and Chemprop-IR, ChemDFM-X effectively distills these surrogate models. Any inherent predictive biases or inaccuracies from these underlying tools are permanently embedded in the new ChemDFM-X encoders.
Main Conclusion: The “separate encoders + unified decoder” architecture with synthetic data generation enables effective multimodal chemical understanding, bridging the gap between specialist and generalist AI systems for chemistry.
Reproducibility Details
Data
The authors constructed a 7.6M sample instruction-tuning dataset derived from 1.3M seed SMILES (sourced from PubChem and USPTO). Note: The final 7.6M multimodal tuning dataset itself isn’t publicly available.
Generation Pipeline:
| Modality | Generation Method | Tool/Model | Sample Count |
|---|---|---|---|
| 2D Graphs | Direct extraction from SMILES | RDKit | 1.1M |
| 3D Conformations | Force field optimization | RDKit + MMFF94 | 1.3M (pseudo-optimal) |
| Molecule Images | Rendering with augmentation | RDKit, Indigo, ChemPix | ~1M (including handwritten style) |
| Reaction Images | Rendering from reaction SMILES | RDKit | 300K |
| MS2 Spectra | Computational prediction | CFM-ID 4.0 | ~700K |
| IR Spectra | Computational prediction | Chemprop-IR | ~1M |
Data Augmentation:
- Molecule images augmented with “handwritten” style using the ChemPix pipeline
- Multiple rendering styles (RDKit default, Indigo clean)
- Spectra generated at multiple energy levels (10eV, 20eV, 40eV for MS2)
Algorithms
Architecture: “Separate Encoders + Unified Decoder”
Code Availability: The authors have only released inference logic. The cross-modal projection training and synthetic data-generation scripts are closed.
Modality Alignment:
- Each modality has a dedicated encoder (frozen pre-trained models where available)
- For graph, conformation, MS2, and IR modalities: 2-layer MLP projector (Linear, GELU, Linear) maps encoder features to LLM input space
- For images: H-Reducer module compresses image tokens by factor of $n=8$ to handle high-resolution chemical images, then projects to LLM input space
- All projected features are concatenated and fed to the unified LLM decoder
Models
Base LLM:
- ChemDFM (13B): LLaMA-based model pre-trained on chemical text and SMILES
Modality Encoders:
| Modality | Encoder | Pre-training Data | Parameter Count | Status |
|---|---|---|---|---|
| 2D Graph | Mole-BERT | 2M molecules | - | Frozen |
| 3D Conformation | Uni-Mol | 209M conformations | - | Frozen |
| Image | CLIP (ViT) | General domain | - | Frozen |
| MS2 Spectrum | Transformer (SeqT) | Trained from scratch | - | Trainable |
| IR Spectrum | Transformer (SeqT) | Trained from scratch | - | Trainable |
Design Rationale: MS2 and IR encoders trained from scratch as Sequence Transformers treating spectral peaks as token sequences, since no suitable pre-trained models exist for chemical spectra.
Evaluation
Metrics:
- Accuracy (Acc) for recognition tasks
- BLEU-2/4 and METEOR for captioning tasks
- AUC-ROC for property prediction (classification)
Code Availability: The adapted code for evaluating on ChemLLMBench and their custom spectral recognition tasks is closed-source.
Benchmarks:
- ChemLLMBench: Adapted for multimodal inputs across molecule captioning, property prediction, and reaction understanding
- MoleculeNet: Standard molecular property prediction tasks (BACE, BBBP, ClinTox, HIV, Tox21)
- USPTO: Reaction prediction and retrosynthesis tasks
- Custom Spectral Tasks: Novel evaluations requiring spectral interpretation
Hardware
Note: The type and quantity of GPUs used, along with the total training wall-time, were not published.
Training Configuration:
- Total Batch Size: 256
- Epochs: 3
- Optimizer: AdamW
Modality-Specific Learning Rates (Peak):
| Modality | Learning Rate | Feature Dimension |
|---|---|---|
| Graph | 1e-5 | 300 |
| Conformation | 2e-4 | 512 |
| Image | 2e-3 | 1024 |
| MS2 / IR | 2e-4 | 768 |
Note: Different learning rates reflect the varying degrees of domain adaptation required. Images (general CLIP) need more adaptation than graphs (chemical Mole-BERT).
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| ChemDFM-X (GitHub) | Code | Apache-2.0 | Inference code only; training and data generation scripts are closed |
| ChemDFM-X-v1.0-13B (HuggingFace) | Model | AGPL-3.0 | 13B parameter multimodal model weights |
Paper Information
Citation: Zhao, Z., Chen, B., Li, J., Chen, L., Wen, L., Wang, P., Zhu, Z., Zhang, D., Wan, Z., Li, Y., Dai, Z., Chen, X., & Yu, K. (2024). ChemDFM-X: Towards Large Multimodal Model for Chemistry. Science China Information Sciences, 67(12), 220109. https://doi.org/10.1007/s11432-024-4243-0
Publication: Science China Information Sciences, December 2024
Additional Resources:
@article{zhaoChemDFMXLargeMultimodal2024,
title = {{{ChemDFM-X}}: {{Towards Large Multimodal Model}} for {{Chemistry}}},
author = {Zhao, Zihan and Chen, Bo and Li, Jingpiao and Chen, Lu and Wen, Liyang and Wang, Pengyu and Zhu, Zichen and Zhang, Danyang and Wan, Ziping and Li, Yansi and Dai, Zhongyang and Chen, Xin and Yu, Kai},
year = {2024},
month = dec,
journal = {Science China Information Sciences},
volume = {67},
number = {12},
pages = {220109},
doi = {10.1007/s11432-024-4243-0},
archiveprefix = {arXiv},
eprint = {2409.13194},
primaryclass = {cs.LG}
}