Paper Information
Citation: Chilingaryan, G., Tamoyan, H., Tevosyan, A., Babayan, N., Khondkaryan, L., Hambardzumyan, K., Navoyan, Z., Khachatrian, H., & Aghajanyan, A. (2024). BARTSmiles: Generative Masked Language Models for Molecular Representations. Journal of Chemical Information and Modeling. https://doi.org/10.1021/acs.jcim.4c00512
Publication: Journal of Chemical Information and Modeling, 2024 (preprint: arXiv 2022)
Additional Resources:
A BART-Based Method for Molecular Self-Supervised Learning
BARTSmiles is a Method paper. It introduces a self-supervised pre-training approach for molecular representations based on the BART (Bidirectional and Auto-Regressive Transformers) architecture from Lewis et al. (2019). The primary contribution is a pre-training strategy, discovered through systematic ablations, that trains a BART-large model on 1.7 billion deduplicated SMILES strings from the ZINC20 dataset. BARTSmiles achieves the best reported results on 11 tasks spanning molecular property classification, regression, and chemical reaction generation.
Scaling Self-Supervised Molecular Representations Beyond Prior Work
At the time of publication, large-scale self-supervised representation learning had produced significant improvements in NLP, computer vision, and speech, but molecular representation learning had not benefited from comparable scale. Previous SMILES-based pre-trained models such as ChemBERTa (Chithrananda et al., 2020) and ChemFormer (Irwin et al., 2022) used encoder-only or encoder-decoder architectures with substantially less compute. ChemFormer, the most closely related prior work, also trained a BART-like model but with a fraction of the compute and data.
The paper argues that three gaps needed to be addressed:
- Scale: Prior molecular pre-training used orders of magnitude less compute than NLP pre-training.
- Architecture choice: Encoder-only models like ChemBERTa cannot perform generative fine-tuning (retrosynthesis, reaction prediction), limiting their applicability.
- Pre-training recipe: Standard BART hyperparameters (e.g., 30% mask token budget) were tuned for natural language and had not been validated for molecular SMILES strings.
Core Innovation: Ablation-Driven Pre-Training Recipe for SMILES
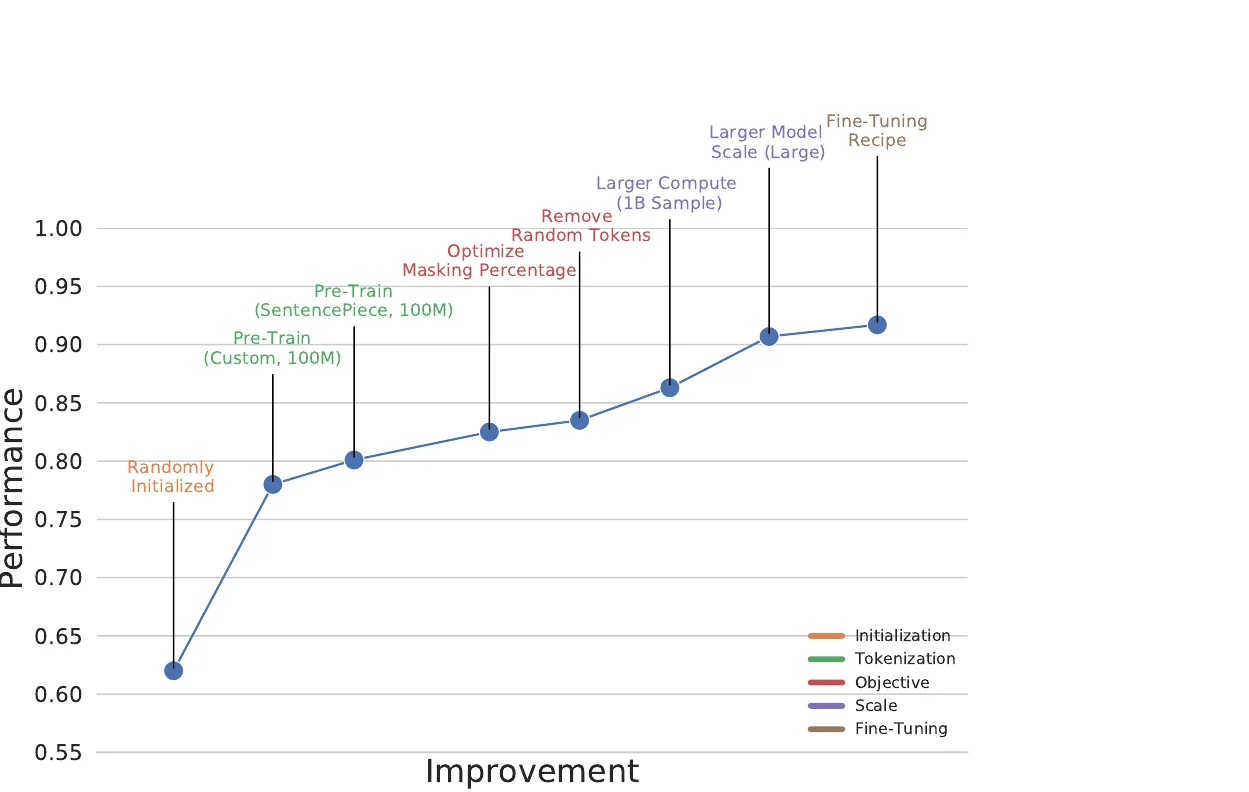
The key insight of BARTSmiles is that the BART denoising objective, when carefully tuned for the molecular domain, learns representations that implicitly encode downstream task information. The authors discover this through a systematic three-stage ablation:
Tokenization
Rather than using hand-crafted tokenization rules that separate individual atoms (C, N, H) and bond symbols (#, =), BARTSmiles uses a learned SentencePiece unigram tokenizer trained on 10 million random SMILES with a vocabulary size of 1,021. On matched compute budgets, learned tokenization achieves 0.801 average AUC-ROC vs. 0.779 for hand-crafted tokenization on the ablation benchmark (HIV, BBBP, ClinTox).
Masking Strategy
The BART denoising objective has three main hyperparameters: the mask token budget (fraction of tokens masked), random mask probability, and the Poisson $\lambda$ controlling mask span length. The ablation results show:
- Mask token budget: The standard BART value of 0.30 is suboptimal for molecules. A budget of 0.20 performs best (0.821 AUC-ROC), with performance degrading at both lower (0.10: 0.753) and higher (0.40: 0.701) budgets.
- Span masking: The choice of random mask probability and $\lambda$ has a minor effect once the budget is set to 0.20. Values of random mask = 0.10 and $\lambda$ = 2.5 or 3.5 all yield 0.821.
- Token randomization: Disabling the randomize-tokens noise (where some tokens are replaced with random tokens rather than masked) improves performance from 0.821 to 0.835.
Scale
Training on the full 1.7 billion molecule ZINC20 dataset (20 hours on 1,024 A100 GPUs, totaling 20,480 A100 GPU-hours) improves performance by 5 absolute AUC-ROC points over the same model trained on 100 million samples. The previous most compute-intensive molecular pre-training used 3,330 V100-hours (Ross et al., 2021).
Implicit Task Encoding
The paper provides a quantitative demonstration that frozen BARTSmiles representations encode task-specific information. Using L1-regularized logistic regression on frozen 1,024-dimensional mean-pooled representations, just 7 neurons are sufficient to achieve 0.987 AUC-ROC on ClinTox (within 2 percentage points of full fine-tuning). Even a single neuron achieves 0.77 AUC-ROC on ClinTox subtask 1.
Experimental Setup: MoleculeNet, Toxicology, and Generative Benchmarks
Classification Tasks
BARTSmiles is evaluated on 7 classification datasets from MoleculeNet (SIDER, ClinTox, Tox21, ToxCast, HIV, BACE, BBBP) plus 2 toxicology datasets (Ames, Micronucleus Assay). All classification tasks use AUC-ROC. Baselines include both supervised graph models (D-MPNN, Attentive FP, 3D InfoMax) and self-supervised methods (ChemBERTa, MolFormer-XL, GROVER-large, MolCLR, iMolCLR).
Selected classification results (AUC-ROC):
| Dataset | BARTSmiles | Previous Best | Previous Best Model |
|---|---|---|---|
| ClinTox | 0.997 | 0.954 | iMolCLR |
| ToxCast | 0.825 | 0.805 | Attentive FP |
| SIDER | 0.705 | 0.699 | iMolCLR |
| Tox21 | 0.851 | 0.858 | Attentive FP |
The authors note that three scaffold-split datasets (HIV, BACE, BBBP) are highly sensitive to the specific split used, and they suspect some baseline results use different or random splits. These results are marked with caveats in the paper.
Regression Tasks
All three MoleculeNet regression tasks (ESOL, FreeSolv, Lipophilicity) are evaluated using RMSE:
| Dataset | BARTSmiles | Previous Best | Previous Best Model |
|---|---|---|---|
| ESOL | 0.095 | 0.279 | MoLFormer-XL |
| FreeSolv | 0.114 | 0.231 | MoLFormer-XL |
| Lipophilicity | 0.292 | 0.529 | MoLFormer-XL |
BARTSmiles achieves substantial improvements on all three regression tasks.
Generative Tasks
Retrosynthesis (USPTO-50k): BARTSmiles achieves 55.6% Top-1 accuracy using a sample-128 + perplexity re-ranking strategy, compared to 55.3% for Dual-TF and 54.3% for ChemFormer. Top-5 and Top-10 results are 74.2% and 80.9% respectively.
Chemical Reaction Prediction (USPTO MIT/LEF/STEREO): BARTSmiles with beam search outperforms the Molecular Transformer baseline across all six evaluation settings. On USPTO-MIT (split), BARTSmiles achieves 91.8% vs. 90.4% for the Transformer baseline.
Fine-Tuning Recipe
The fine-tuning approach is designed to minimize hyperparameter tuning:
- Batch size 16, 10 epochs, polynomial decay learning rate schedule with warmup at 16% of training
- Grid search over dropout (0.1, 0.2, 0.3) and learning rate ($5 \times 10^{-6}$, $1 \times 10^{-5}$, $3 \times 10^{-5}$)
- Stochastic Weight Averaging (SWA) over three sets of four checkpoints
- For generative tasks: R3F regularization (Aghajanyan et al., 2020a) and full fp32 precision
- For generation: beam search (beam size 10) or sample 128 sequences with perplexity re-ranking
Key Findings and Limitations
Key Findings
- Scale matters for molecular pre-training: Training on 1.7B molecules with 20,480 A100 GPU-hours yields 5 absolute points of AUC-ROC improvement over training on 100M molecules.
- Domain-specific ablation is necessary: The optimal BART masking configuration for molecules (20% budget, no token randomization) differs from the standard NLP configuration (30% budget, with randomization).
- Frozen representations capture task structure: A small number of neurons from the frozen model can nearly match full fine-tuning performance on certain tasks, suggesting the pre-training objective implicitly encodes molecular properties.
- Interpretability aligns with domain knowledge: Integrated Gradients attribution on fine-tuned BARTSmiles highlights known structural alerts (e.g., nitro groups in mutagenic compounds, hydroxyl groups in soluble compounds).
Limitations
- Scaffold split sensitivity: Results on HIV, BACE, and BBBP are sensitive to the specific scaffold split, making direct comparison with baselines difficult.
- Pre-training data distribution: The Frechet distance analysis shows that some downstream datasets (BBBP, SIDER) are far from ZINC20 in representation space, which may explain weaker performance on those tasks.
- Fingerprints carry complementary information: On the Ames and Micronucleus Assay datasets, BARTSmiles alone does not beat fingerprint-based baselines. Combining BARTSmiles with ECFP4 fingerprints closes the gap, implying that SMILES-based pre-training does not fully capture all structural information.
- Compute requirements: Pre-training requires 1,024 A100 GPUs, which limits accessibility.
Future Directions
The authors suggest investigating the impact of pre-training data composition, noting that ZINC20 contains over a billion molecules but its distribution may be irrelevant for many downstream tasks. They also propose further collaboration between ML and chemistry experts to discover new molecular substructure-property relationships.
Reproducibility Details
Artifacts
| Artifact | Type | License | Notes |
|---|---|---|---|
| BARTSmiles (GitHub) | Code + Model | MIT | Pre-training, fine-tuning, and evaluation scripts with pre-trained weights |
Data
| Purpose | Dataset | Size | Notes |
|---|---|---|---|
| Pre-training | ZINC20 (deduplicated) | ~1.7B molecules | Canonicalized SMILES, 10K validation holdout |
| Classification | MoleculeNet (7 datasets) | 1,427-41,127 compounds | AUC-ROC metric |
| Regression | MoleculeNet (3 datasets) | 642-4,200 compounds | RMSE metric |
| Toxicology | Ames, MN Assay | 6,512 / 641 compounds | Cross-validation for Ames; external test for MN |
| Retrosynthesis | USPTO-50k | Standard split | Top-K accuracy |
| Reaction prediction | USPTO (MIT/LEF/STEREO) | Standard splits | Top-1 accuracy |
Algorithms
- Architecture: BART-Large (pre-layer norm Transformer encoder-decoder)
- Tokenizer: SentencePiece unigram, vocabulary size 1,021, max sequence length 128
- Pre-training objective: BART denoising (mask token budget 0.20, Poisson span masking with $\lambda$ = 2.5, no token randomization)
- Fine-tuning: polynomial decay LR, SWA, grid search over dropout and LR
- Generative fine-tuning: R3F regularization, fp32 precision, Adam initialized from pre-training moving averages
Models
- BART-Large architecture (exact parameter count not specified in paper)
- Pre-trained checkpoint released on GitHub
- Maximum sequence length: 128 tokens
Evaluation
| Task | Metric | BARTSmiles | Notes |
|---|---|---|---|
| ClinTox | AUC-ROC | 0.997 | New SOTA |
| ToxCast | AUC-ROC | 0.825 | New SOTA |
| ESOL | RMSE | 0.095 | New SOTA |
| FreeSolv | RMSE | 0.114 | New SOTA |
| Lipophilicity | RMSE | 0.292 | New SOTA |
| USPTO-50k Retro (Top-1) | Accuracy | 55.6% | New SOTA (sample + re-rank) |
| USPTO-MIT Rxn (Split) | Accuracy | 91.8% | New SOTA (beam-10) |
Hardware
- Pre-training: 1,024 NVIDIA A100 GPUs for 20 hours (20,480 A100 GPU-hours)
- Ablation runs: 128 A100 GPUs per run
- Framework: FairSeq with FairScale (fully sharded data parallel), automatic mixed precision
- Experiment tracking: Aim
Citation
@article{chilingaryan2024bartsmiles,
title={BARTSmiles: Generative Masked Language Models for Molecular Representations},
author={Chilingaryan, Gayane and Tamoyan, Hovhannes and Tevosyan, Ani and Babayan, Nelly and Khondkaryan, Lusine and Hambardzumyan, Karen and Navoyan, Zaven and Khachatrian, Hrant and Aghajanyan, Armen},
journal={Journal of Chemical Information and Modeling},
doi={10.1021/acs.jcim.4c00512},
year={2024}
}