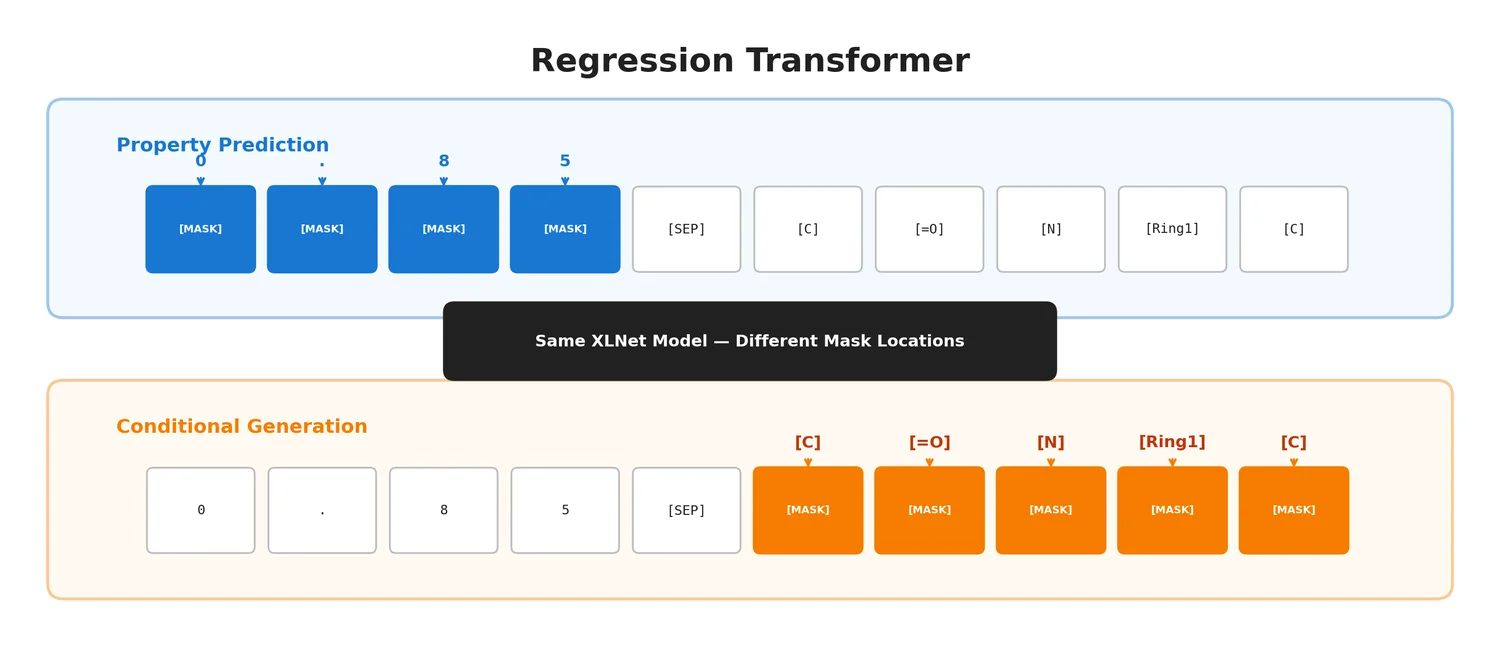
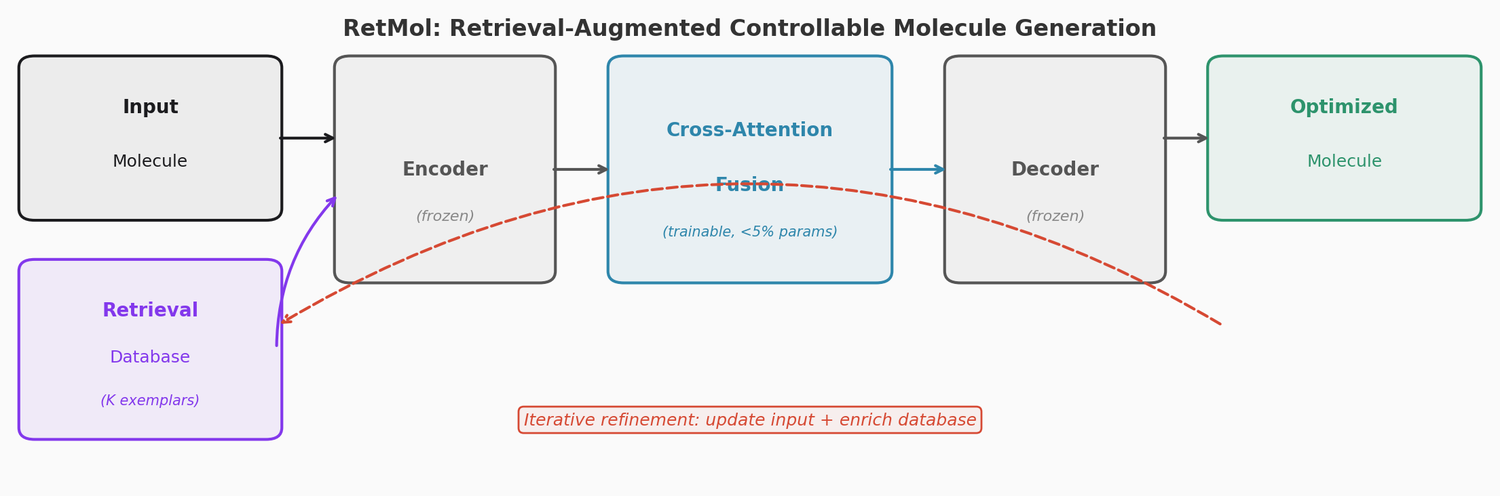
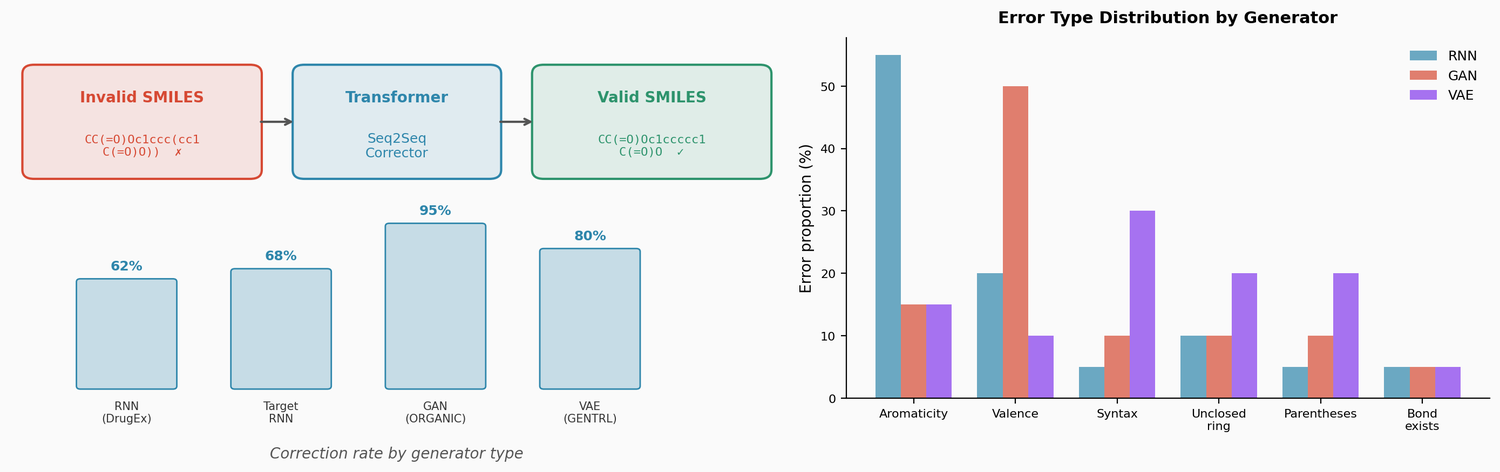
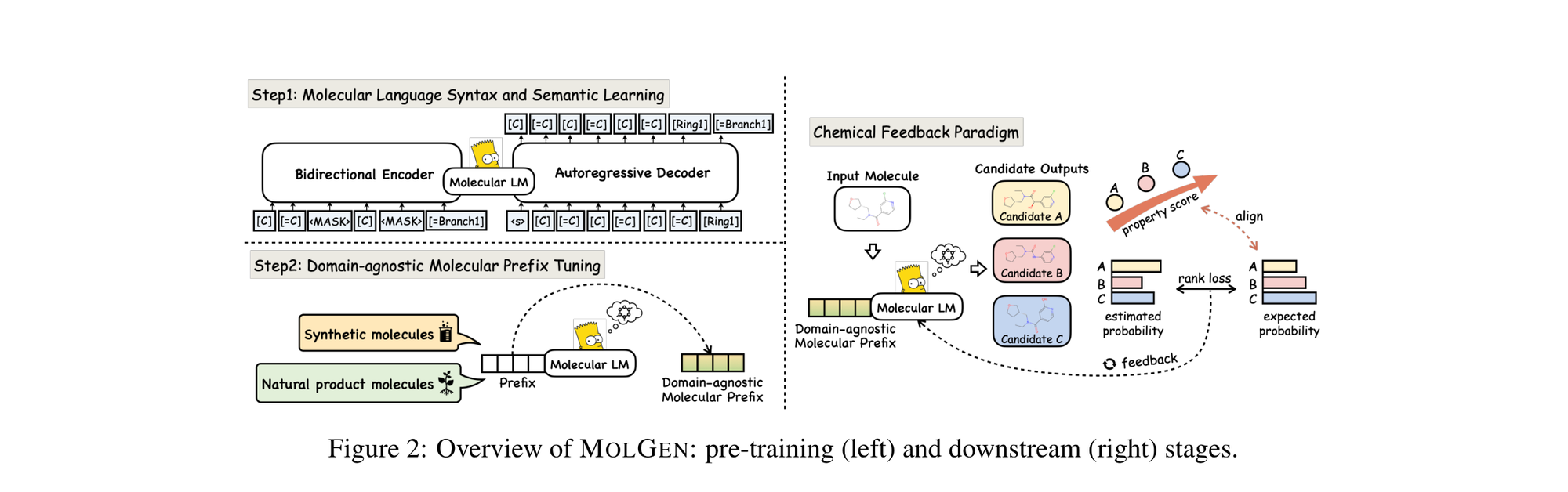
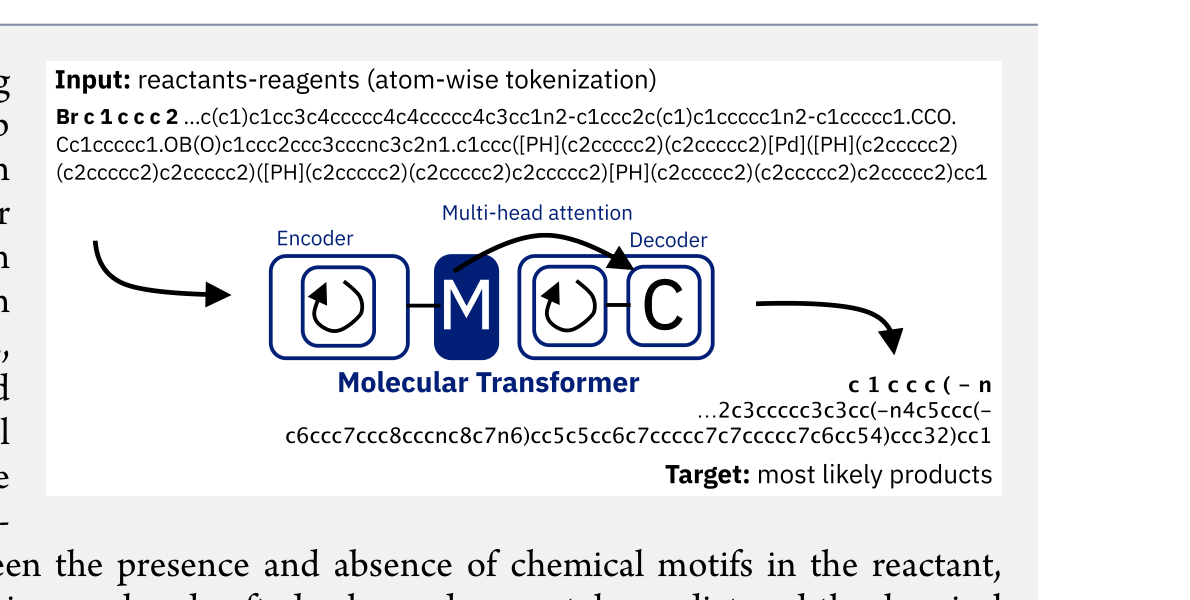
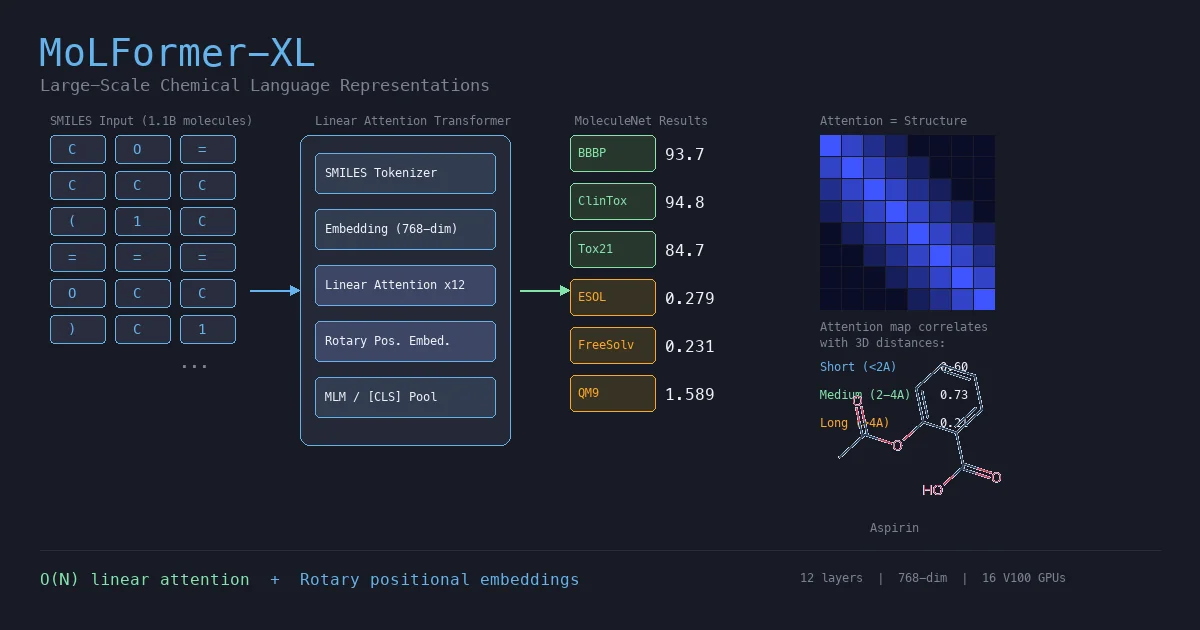
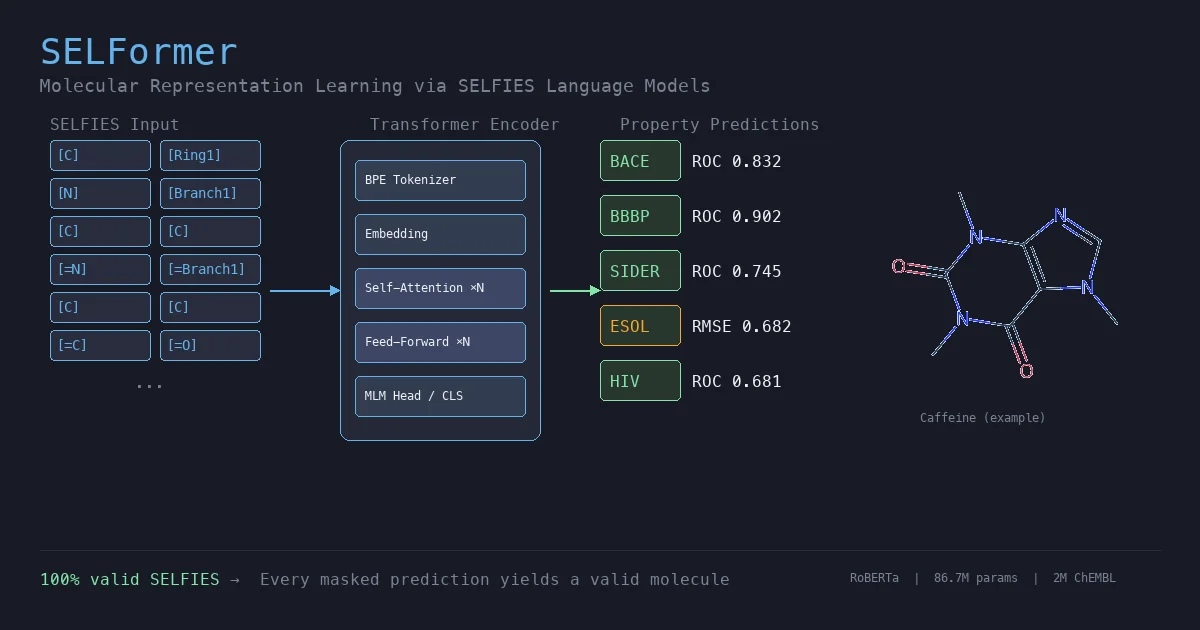
This section covers chemical language models: architectures that learn molecular representations directly from chemical string notations (SMILES, SELFIES, InChI). Notes here include encoder-only transformers like ChemBERTa and MoLFormer for property prediction, sequence-to-sequence models like Chemformer for reaction prediction and GP-MoLFormer for molecular generation, and translation models like STOUT for SMILES-to-IUPAC conversion. For multimodal and reasoning LLMs applied to chemistry, see LLMs for Chemistry.
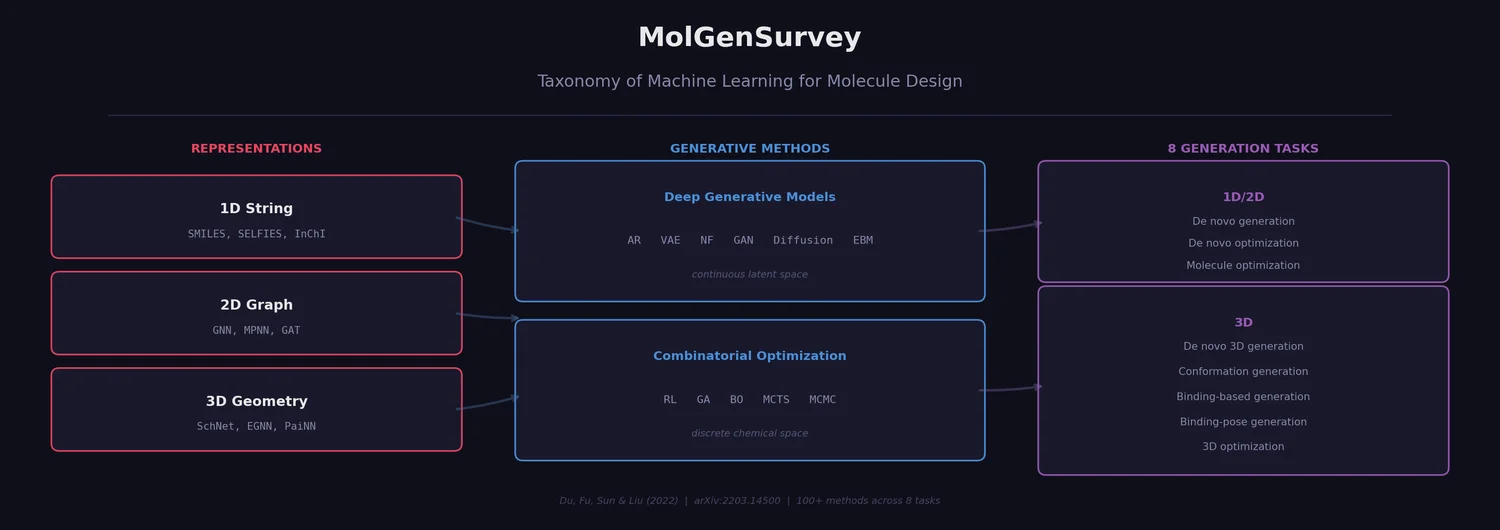
MolGenSurvey: Systematic Survey of ML for Molecule Design
MolGenSurvey systematically reviews ML models for molecule design, organizing the field by molecular representation (1D/2D/3D), generative method (deep generative models vs. combinatorial optimization), and task type (8 distinct generation/optimization tasks). It catalogs over 100 methods, unifies task definitions via input/output/goal taxonomy, and identifies key challenges including out-of-distribution generation, oracle costs, and lack of unified benchmarks.